Luis Filipe Nakayama
Benchmarking Ophthalmology Foundation Models for Clinically Significant Age Macular Degeneration Detection
May 08, 2025Abstract:Self-supervised learning (SSL) has enabled Vision Transformers (ViTs) to learn robust representations from large-scale natural image datasets, enhancing their generalization across domains. In retinal imaging, foundation models pretrained on either natural or ophthalmic data have shown promise, but the benefits of in-domain pretraining remain uncertain. To investigate this, we benchmark six SSL-pretrained ViTs on seven digital fundus image (DFI) datasets totaling 70,000 expert-annotated images for the task of moderate-to-late age-related macular degeneration (AMD) identification. Our results show that iBOT pretrained on natural images achieves the highest out-of-distribution generalization, with AUROCs of 0.80-0.97, outperforming domain-specific models, which achieved AUROCs of 0.78-0.96 and a baseline ViT-L with no pretraining, which achieved AUROCs of 0.68-0.91. These findings highlight the value of foundation models in improving AMD identification and challenge the assumption that in-domain pretraining is necessary. Furthermore, we release BRAMD, an open-access dataset (n=587) of DFIs with AMD labels from Brazil.
Enhancing Retinal Vessel Segmentation Generalization via Layout-Aware Generative Modelling
Mar 03, 2025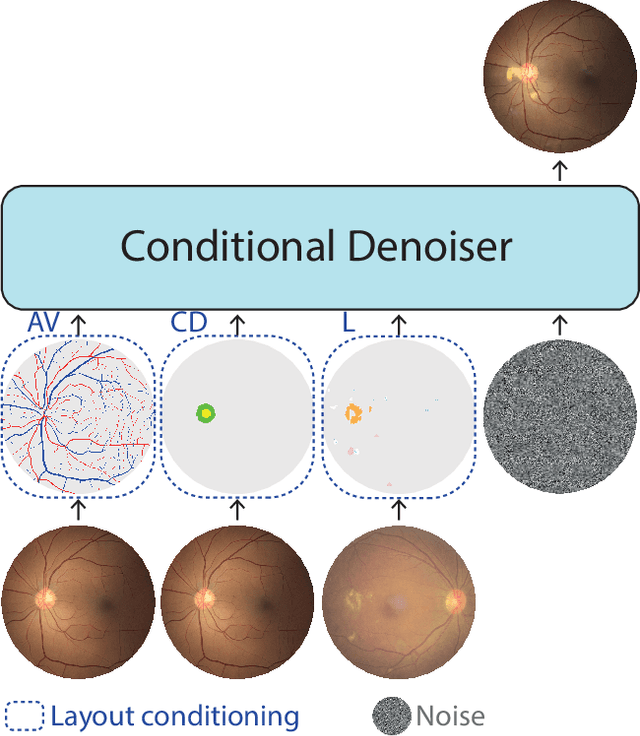
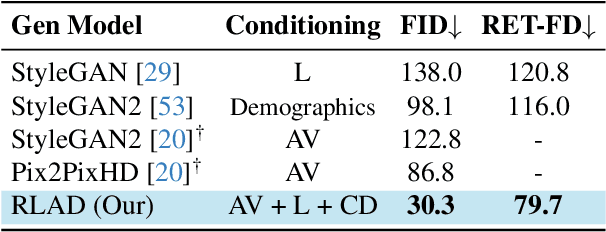
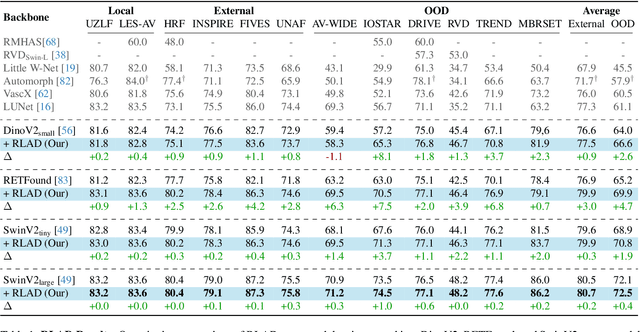
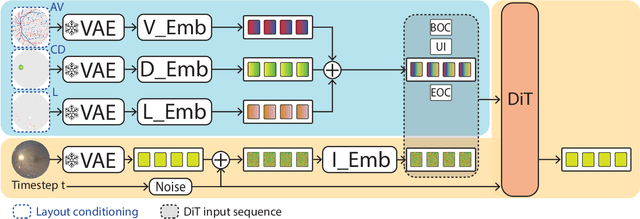

Abstract:Generalization in medical segmentation models is challenging due to limited annotated datasets and imaging variability. To address this, we propose Retinal Layout-Aware Diffusion (RLAD), a novel diffusion-based framework for generating controllable layout-aware images. RLAD conditions image generation on multiple key layout components extracted from real images, ensuring high structural fidelity while enabling diversity in other components. Applied to retinal fundus imaging, we augmented the training datasets by synthesizing paired retinal images and vessel segmentations conditioned on extracted blood vessels from real images, while varying other layout components such as lesions and the optic disc. Experiments demonstrated that RLAD-generated data improved generalization in retinal vessel segmentation by up to 8.1%. Furthermore, we present REYIA, a comprehensive dataset comprising 586 manually segmented retinal images. To foster reproducibility and drive innovation, both our code and dataset will be made publicly accessible.
Multi-OphthaLingua: A Multilingual Benchmark for Assessing and Debiasing LLM Ophthalmological QA in LMICs
Dec 18, 2024



Abstract:Current ophthalmology clinical workflows are plagued by over-referrals, long waits, and complex and heterogeneous medical records. Large language models (LLMs) present a promising solution to automate various procedures such as triaging, preliminary tests like visual acuity assessment, and report summaries. However, LLMs have demonstrated significantly varied performance across different languages in natural language question-answering tasks, potentially exacerbating healthcare disparities in Low and Middle-Income Countries (LMICs). This study introduces the first multilingual ophthalmological question-answering benchmark with manually curated questions parallel across languages, allowing for direct cross-lingual comparisons. Our evaluation of 6 popular LLMs across 7 different languages reveals substantial bias across different languages, highlighting risks for clinical deployment of LLMs in LMICs. Existing debiasing methods such as Translation Chain-of-Thought or Retrieval-augmented generation (RAG) by themselves fall short of closing this performance gap, often failing to improve performance across all languages and lacking specificity for the medical domain. To address this issue, We propose CLARA (Cross-Lingual Reflective Agentic system), a novel inference time de-biasing method leveraging retrieval augmented generation and self-verification. Our approach not only improves performance across all languages but also significantly reduces the multilingual bias gap, facilitating equitable LLM application across the globe.
Classification of Keratitis from Eye Corneal Photographs using Deep Learning
Nov 13, 2024



Abstract:Keratitis is an inflammatory corneal condition responsible for 10% of visual impairment in low- and middle-income countries (LMICs), with bacteria, fungi, or amoeba as the most common infection etiologies. While an accurate and timely diagnosis is crucial for the selected treatment and the patients' sight outcomes, due to the high cost and limited availability of laboratory diagnostics in LMICs, diagnosis is often made by clinical observation alone, despite its lower accuracy. In this study, we investigate and compare different deep learning approaches to diagnose the source of infection: 1) three separate binary models for infection type predictions; 2) a multitask model with a shared backbone and three parallel classification layers (Multitask V1); and, 3) a multitask model with a shared backbone and a multi-head classification layer (Multitask V2). We used a private Brazilian cornea dataset to conduct the empirical evaluation. We achieved the best results with Multitask V2, with an area under the receiver operating characteristic curve (AUROC) confidence intervals of 0.7413-0.7740 (bacteria), 0.8395-0.8725 (fungi), and 0.9448-0.9616 (amoeba). A statistical analysis of the impact of patient features on models' performance revealed that sex significantly affects amoeba infection prediction, and age seems to affect fungi and bacteria predictions.
Does Data-Efficient Generalization Exacerbate Bias in Foundation Models?
Sep 02, 2024


Abstract:Foundation models have emerged as robust models with label efficiency in diverse domains. In medical imaging, these models contribute to the advancement of medical diagnoses due to the difficulty in obtaining labeled data. However, it is unclear whether using a large amount of unlabeled data, biased by the presence of sensitive attributes during pre-training, influences the fairness of the model. This research examines the bias in the Foundation model (RetFound) when it is applied to fine-tune the Brazilian Multilabel Ophthalmological Dataset (BRSET), which has a different population than the pre-training dataset. The model evaluation, in comparison with supervised learning, shows that the Foundation Model has the potential to reduce the gap between the maximum AUC and minimum AUC evaluations across gender and age groups. However, in a data-efficient generalization, the model increases the bias when the data amount decreases. These findings suggest that when deploying a Foundation Model in real-life scenarios with limited data, the possibility of fairness issues should be considered.
Multimodal Deep Learning for Low-Resource Settings: A Vector Embedding Alignment Approach for Healthcare Applications
Jun 02, 2024Abstract:Large-scale multi-modal deep learning models have revolutionized domains such as healthcare, highlighting the importance of computational power. However, in resource-constrained regions like Low and Middle-Income Countries (LMICs), limited access to GPUs and data poses significant challenges, often leaving CPUs as the sole resource. To address this, we advocate for leveraging vector embeddings to enable flexible and efficient computational methodologies, democratizing multimodal deep learning across diverse contexts. Our paper investigates the efficiency and effectiveness of using vector embeddings from single-modal foundation models and multi-modal Vision-Language Models (VLMs) for multimodal deep learning in low-resource environments, particularly in healthcare. Additionally, we propose a simple yet effective inference-time method to enhance performance by aligning image-text embeddings. Comparing these approaches with traditional methods, we assess their impact on computational efficiency and model performance using metrics like accuracy, F1-score, inference time, training time, and memory usage across three medical modalities: BRSET (ophthalmology), HAM10000 (dermatology), and SatelliteBench (public health). Our findings show that embeddings reduce computational demands without compromising model performance. Furthermore, our alignment method improves performance in medical tasks. This research promotes sustainable AI practices by optimizing resources in constrained environments, highlighting the potential of embedding-based approaches for efficient multimodal learning. Vector embeddings democratize multimodal deep learning in LMICs, particularly in healthcare, enhancing AI adaptability in varied use cases.
DF-DM: A foundational process model for multimodal data fusion in the artificial intelligence era
Apr 18, 2024


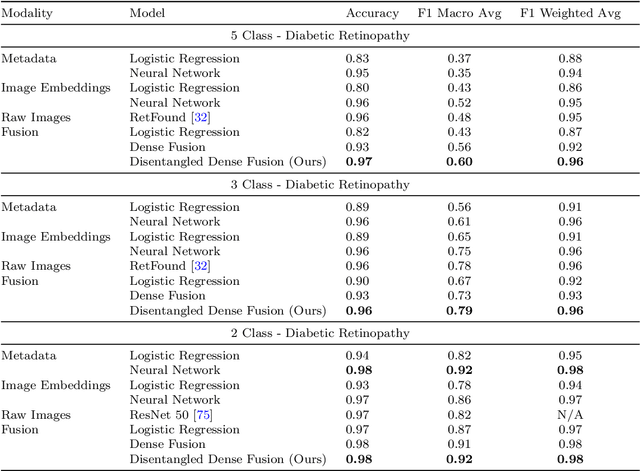
Abstract:In the big data era, integrating diverse data modalities poses significant challenges, particularly in complex fields like healthcare. This paper introduces a new process model for multimodal Data Fusion for Data Mining, integrating embeddings and the Cross-Industry Standard Process for Data Mining with the existing Data Fusion Information Group model. Our model aims to decrease computational costs, complexity, and bias while improving efficiency and reliability. We also propose "disentangled dense fusion", a novel embedding fusion method designed to optimize mutual information and facilitate dense inter-modality feature interaction, thereby minimizing redundant information. We demonstrate the model's efficacy through three use cases: predicting diabetic retinopathy using retinal images and patient metadata, domestic violence prediction employing satellite imagery, internet, and census data, and identifying clinical and demographic features from radiography images and clinical notes. The model achieved a Macro F1 score of 0.92 in diabetic retinopathy prediction, an R-squared of 0.854 and sMAPE of 24.868 in domestic violence prediction, and a macro AUC of 0.92 and 0.99 for disease prediction and sex classification, respectively, in radiological analysis. These results underscore the Data Fusion for Data Mining model's potential to significantly impact multimodal data processing, promoting its adoption in diverse, resource-constrained settings.
DRStageNet: Deep Learning for Diabetic Retinopathy Staging from Fundus Images
Dec 22, 2023



Abstract:Diabetic retinopathy (DR) is a prevalent complication of diabetes associated with a significant risk of vision loss. Timely identification is critical to curb vision impairment. Algorithms for DR staging from digital fundus images (DFIs) have been recently proposed. However, models often fail to generalize due to distribution shifts between the source domain on which the model was trained and the target domain where it is deployed. A common and particularly challenging shift is often encountered when the source- and target-domain supports do not fully overlap. In this research, we introduce DRStageNet, a deep learning model designed to mitigate this challenge. We used seven publicly available datasets, comprising a total of 93,534 DFIs that cover a variety of patient demographics, ethnicities, geographic origins and comorbidities. We fine-tune DINOv2, a pretrained model of self-supervised vision transformer, and implement a multi-source domain fine-tuning strategy to enhance generalization performance. We benchmark and demonstrate the superiority of our method to two state-of-the-art benchmarks, including a recently published foundation model. We adapted the grad-rollout method to our regression task in order to provide high-resolution explainability heatmaps. The error analysis showed that 59\% of the main errors had incorrect reference labels. DRStageNet is accessible at URL [upon acceptance of the manuscript].
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge