David Restrepo
On the Cone Effect and Modality Gap in Medical Vision-Language Embeddings
Mar 18, 2026Abstract:Vision-Language Models (VLMs) exhibit a characteristic "cone effect" in which nonlinear encoders map embeddings into highly concentrated regions of the representation space, contributing to cross-modal separation known as the modality gap. While this phenomenon has been widely observed, its practical impact on supervised multimodal learning -particularly in medical domains- remains unclear. In this work, we introduce a lightweight post-hoc mechanism that keeps pretrained VLM encoders frozen while continuously controlling cross-modal separation through a single hyperparameter {λ}. This enables systematic analysis of how the modality gap affects downstream multimodal performance without expensive retraining. We evaluate generalist (CLIP, SigLIP) and medically specialized (BioMedCLIP, MedSigLIP) models across diverse medical and natural datasets in a supervised multimodal settings. Results consistently show that reducing excessive modality gap improves downstream performance, with medical datasets exhibiting stronger sensitivity to gap modulation; however, fully collapsing the gap is not always optimal, and intermediate, task-dependent separation yields the best results. These findings position the modality gap as a tunable property of multimodal representations rather than a quantity that should be universally minimized.
Learning Representations from Incomplete EHR Data with Dual-Masked Autoencoding
Feb 16, 2026Abstract:Learning from electronic health records (EHRs) time series is challenging due to irregular sam- pling, heterogeneous missingness, and the resulting sparsity of observations. Prior self-supervised meth- ods either impute before learning, represent missingness through a dedicated input signal, or optimize solely for imputation, reducing their capacity to efficiently learn representations that support clinical downstream tasks. We propose the Augmented-Intrinsic Dual-Masked Autoencoder (AID-MAE), which learns directly from incomplete time series by applying an intrinsic missing mask to represent naturally missing values and an augmented mask that hides a subset of observed values for reconstruction during training. AID-MAE processes only the unmasked subset of tokens and consistently outperforms strong baselines, including XGBoost and DuETT, across multiple clinical tasks on two datasets. In addition, the learned embeddings naturally stratify patient cohorts in the representation space.
CURENet: Combining Unified Representations for Efficient Chronic Disease Prediction
Nov 14, 2025Abstract:Electronic health records (EHRs) are designed to synthesize diverse data types, including unstructured clinical notes, structured lab tests, and time-series visit data. Physicians draw on these multimodal and temporal sources of EHR data to form a comprehensive view of a patient's health, which is crucial for informed therapeutic decision-making. Yet, most predictive models fail to fully capture the interactions, redundancies, and temporal patterns across multiple data modalities, often focusing on a single data type or overlooking these complexities. In this paper, we present CURENet, a multimodal model (Combining Unified Representations for Efficient chronic disease prediction) that integrates unstructured clinical notes, lab tests, and patients' time-series data by utilizing large language models (LLMs) for clinical text processing and textual lab tests, as well as transformer encoders for longitudinal sequential visits. CURENet has been capable of capturing the intricate interaction between different forms of clinical data and creating a more reliable predictive model for chronic illnesses. We evaluated CURENet using the public MIMIC-III and private FEMH datasets, where it achieved over 94\% accuracy in predicting the top 10 chronic conditions in a multi-label framework. Our findings highlight the potential of multimodal EHR integration to enhance clinical decision-making and improve patient outcomes.
On the Risk of Misleading Reports: Diagnosing Textual Biases in Multimodal Clinical AI
Jul 31, 2025



Abstract:Clinical decision-making relies on the integrated analysis of medical images and the associated clinical reports. While Vision-Language Models (VLMs) can offer a unified framework for such tasks, they can exhibit strong biases toward one modality, frequently overlooking critical visual cues in favor of textual information. In this work, we introduce Selective Modality Shifting (SMS), a perturbation-based approach to quantify a model's reliance on each modality in binary classification tasks. By systematically swapping images or text between samples with opposing labels, we expose modality-specific biases. We assess six open-source VLMs-four generalist models and two fine-tuned for medical data-on two medical imaging datasets with distinct modalities: MIMIC-CXR (chest X-ray) and FairVLMed (scanning laser ophthalmoscopy). By assessing model performance and the calibration of every model in both unperturbed and perturbed settings, we reveal a marked dependency on text input, which persists despite the presence of complementary visual information. We also perform a qualitative attention-based analysis which further confirms that image content is often overshadowed by text details. Our findings highlight the importance of designing and evaluating multimodal medical models that genuinely integrate visual and textual cues, rather than relying on single-modality signals.
Performance Gains of LLMs With Humans in a World of LLMs Versus Humans
May 13, 2025



Abstract:Currently, a considerable research effort is devoted to comparing LLMs to a group of human experts, where the term "expert" is often ill-defined or variable, at best, in a state of constantly updating LLM releases. Without proper safeguards in place, LLMs will threaten to cause harm to the established structure of safe delivery of patient care which has been carefully developed throughout history to keep the safety of the patient at the forefront. A key driver of LLM innovation is founded on community research efforts which, if continuing to operate under "humans versus LLMs" principles, will expedite this trend. Therefore, research efforts moving forward must focus on effectively characterizing the safe use of LLMs in clinical settings that persist across the rapid development of novel LLM models. In this communication, we demonstrate that rather than comparing LLMs to humans, there is a need to develop strategies enabling efficient work of humans with LLMs in an almost symbiotic manner.
Representation Learning of Lab Values via Masked AutoEncoder
Jan 05, 2025



Abstract:Accurate imputation of missing laboratory values in electronic health records (EHRs) is critical to enable robust clinical predictions and reduce biases in AI systems in healthcare. Existing methods, such as variational autoencoders (VAEs) and decision tree-based approaches such as XGBoost, struggle to model the complex temporal and contextual dependencies in EHR data, mainly in underrepresented groups. In this work, we propose Lab-MAE, a novel transformer-based masked autoencoder framework that leverages self-supervised learning for the imputation of continuous sequential lab values. Lab-MAE introduces a structured encoding scheme that jointly models laboratory test values and their corresponding timestamps, enabling explicit capturing temporal dependencies. Empirical evaluation on the MIMIC-IV dataset demonstrates that Lab-MAE significantly outperforms the state-of-the-art baselines such as XGBoost across multiple metrics, including root mean square error (RMSE), R-squared (R2), and Wasserstein distance (WD). Notably, Lab-MAE achieves equitable performance across demographic groups of patients, advancing fairness in clinical predictions. We further investigate the role of follow-up laboratory values as potential shortcut features, revealing Lab-MAE's robustness in scenarios where such data is unavailable. The findings suggest that our transformer-based architecture, adapted to the characteristics of the EHR data, offers a foundation model for more accurate and fair clinical imputation models. In addition, we measure and compare the carbon footprint of Lab-MAE with the baseline XGBoost model, highlighting its environmental requirements.
Multi-OphthaLingua: A Multilingual Benchmark for Assessing and Debiasing LLM Ophthalmological QA in LMICs
Dec 18, 2024



Abstract:Current ophthalmology clinical workflows are plagued by over-referrals, long waits, and complex and heterogeneous medical records. Large language models (LLMs) present a promising solution to automate various procedures such as triaging, preliminary tests like visual acuity assessment, and report summaries. However, LLMs have demonstrated significantly varied performance across different languages in natural language question-answering tasks, potentially exacerbating healthcare disparities in Low and Middle-Income Countries (LMICs). This study introduces the first multilingual ophthalmological question-answering benchmark with manually curated questions parallel across languages, allowing for direct cross-lingual comparisons. Our evaluation of 6 popular LLMs across 7 different languages reveals substantial bias across different languages, highlighting risks for clinical deployment of LLMs in LMICs. Existing debiasing methods such as Translation Chain-of-Thought or Retrieval-augmented generation (RAG) by themselves fall short of closing this performance gap, often failing to improve performance across all languages and lacking specificity for the medical domain. To address this issue, We propose CLARA (Cross-Lingual Reflective Agentic system), a novel inference time de-biasing method leveraging retrieval augmented generation and self-verification. Our approach not only improves performance across all languages but also significantly reduces the multilingual bias gap, facilitating equitable LLM application across the globe.
WorldMedQA-V: a multilingual, multimodal medical examination dataset for multimodal language models evaluation
Oct 16, 2024Abstract:Multimodal/vision language models (VLMs) are increasingly being deployed in healthcare settings worldwide, necessitating robust benchmarks to ensure their safety, efficacy, and fairness. Multiple-choice question and answer (QA) datasets derived from national medical examinations have long served as valuable evaluation tools, but existing datasets are largely text-only and available in a limited subset of languages and countries. To address these challenges, we present WorldMedQA-V, an updated multilingual, multimodal benchmarking dataset designed to evaluate VLMs in healthcare. WorldMedQA-V includes 568 labeled multiple-choice QAs paired with 568 medical images from four countries (Brazil, Israel, Japan, and Spain), covering original languages and validated English translations by native clinicians, respectively. Baseline performance for common open- and closed-source models are provided in the local language and English translations, and with and without images provided to the model. The WorldMedQA-V benchmark aims to better match AI systems to the diverse healthcare environments in which they are deployed, fostering more equitable, effective, and representative applications.
MEDFuse: Multimodal EHR Data Fusion with Masked Lab-Test Modeling and Large Language Models
Jul 17, 2024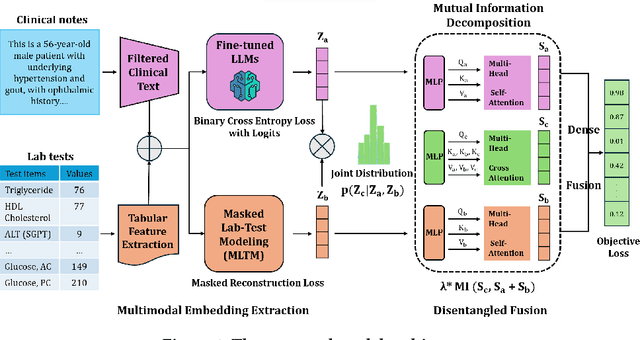
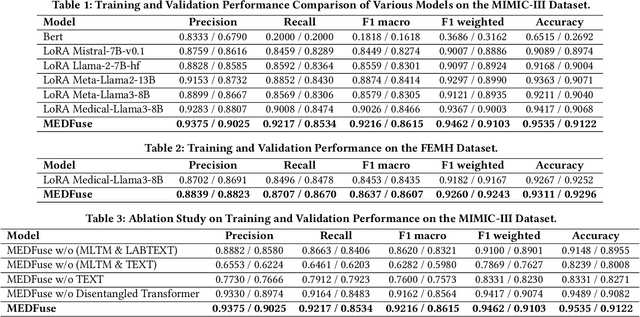
Abstract:Electronic health records (EHRs) are multimodal by nature, consisting of structured tabular features like lab tests and unstructured clinical notes. In real-life clinical practice, doctors use complementary multimodal EHR data sources to get a clearer picture of patients' health and support clinical decision-making. However, most EHR predictive models do not reflect these procedures, as they either focus on a single modality or overlook the inter-modality interactions/redundancy. In this work, we propose MEDFuse, a Multimodal EHR Data Fusion framework that incorporates masked lab-test modeling and large language models (LLMs) to effectively integrate structured and unstructured medical data. MEDFuse leverages multimodal embeddings extracted from two sources: LLMs fine-tuned on free clinical text and masked tabular transformers trained on structured lab test results. We design a disentangled transformer module, optimized by a mutual information loss to 1) decouple modality-specific and modality-shared information and 2) extract useful joint representation from the noise and redundancy present in clinical notes. Through comprehensive validation on the public MIMIC-III dataset and the in-house FEMH dataset, MEDFuse demonstrates great potential in advancing clinical predictions, achieving over 90% F1 score in the 10-disease multi-label classification task.
Efficient In-Context Medical Segmentation with Meta-driven Visual Prompt Selection
Jul 15, 2024



Abstract:In-context learning (ICL) with Large Vision Models (LVMs) presents a promising avenue in medical image segmentation by reducing the reliance on extensive labeling. However, the ICL performance of LVMs highly depends on the choices of visual prompts and suffers from domain shifts. While existing works leveraging LVMs for medical tasks have focused mainly on model-centric approaches like fine-tuning, we study an orthogonal data-centric perspective on how to select good visual prompts to facilitate generalization to medical domain. In this work, we propose a label-efficient in-context medical segmentation method by introducing a novel Meta-driven Visual Prompt Selection mechanism (MVPS), where a prompt retriever obtained from a meta-learning framework actively selects the optimal images as prompts to promote model performance and generalizability. Evaluated on 8 datasets and 4 tasks across 3 medical imaging modalities, our proposed approach demonstrates consistent gains over existing methods under different scenarios, improving both computational and label efficiency. Finally, we show that MVPS is a flexible, finetuning-free module that could be easily plugged into different backbones and combined with other model-centric approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge