Fang-Ming Hung
Chain-of-Thought Reasoning with Large Language Models for Clinical Alzheimer's Disease Assessment and Diagnosis
Feb 15, 2026Abstract:Alzheimer's disease (AD) has become a prevalent neurodegenerative disease worldwide. Traditional diagnosis still relies heavily on medical imaging and clinical assessment by physicians, which is often time-consuming and resource-intensive in terms of both human expertise and healthcare resources. In recent years, large language models (LLMs) have been increasingly applied to the medical field using electronic health records (EHRs), yet their application in Alzheimer's disease assessment remains limited, particularly given that AD involves complex multifactorial etiologies that are difficult to observe directly through imaging modalities. In this work, we propose leveraging LLMs to perform Chain-of-Thought (CoT) reasoning on patients' clinical EHRs. Unlike direct fine-tuning of LLMs on EHR data for AD classification, our approach utilizes LLM-generated CoT reasoning paths to provide the model with explicit diagnostic rationale for AD assessment, followed by structured CoT-based predictions. This pipeline not only enhances the model's ability to diagnose intrinsically complex factors but also improves the interpretability of the prediction process across different stages of AD progression. Experimental results demonstrate that the proposed CoT-based diagnostic framework significantly enhances stability and diagnostic performance across multiple CDR grading tasks, achieving up to a 15% improvement in F1 score compared to the zero-shot baseline method.
CURENet: Combining Unified Representations for Efficient Chronic Disease Prediction
Nov 14, 2025Abstract:Electronic health records (EHRs) are designed to synthesize diverse data types, including unstructured clinical notes, structured lab tests, and time-series visit data. Physicians draw on these multimodal and temporal sources of EHR data to form a comprehensive view of a patient's health, which is crucial for informed therapeutic decision-making. Yet, most predictive models fail to fully capture the interactions, redundancies, and temporal patterns across multiple data modalities, often focusing on a single data type or overlooking these complexities. In this paper, we present CURENet, a multimodal model (Combining Unified Representations for Efficient chronic disease prediction) that integrates unstructured clinical notes, lab tests, and patients' time-series data by utilizing large language models (LLMs) for clinical text processing and textual lab tests, as well as transformer encoders for longitudinal sequential visits. CURENet has been capable of capturing the intricate interaction between different forms of clinical data and creating a more reliable predictive model for chronic illnesses. We evaluated CURENet using the public MIMIC-III and private FEMH datasets, where it achieved over 94\% accuracy in predicting the top 10 chronic conditions in a multi-label framework. Our findings highlight the potential of multimodal EHR integration to enhance clinical decision-making and improve patient outcomes.
MEDFuse: Multimodal EHR Data Fusion with Masked Lab-Test Modeling and Large Language Models
Jul 17, 2024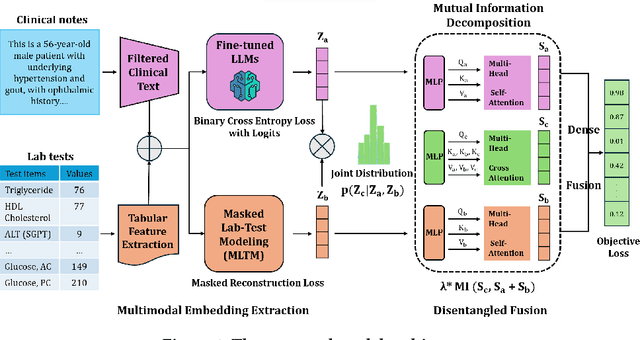
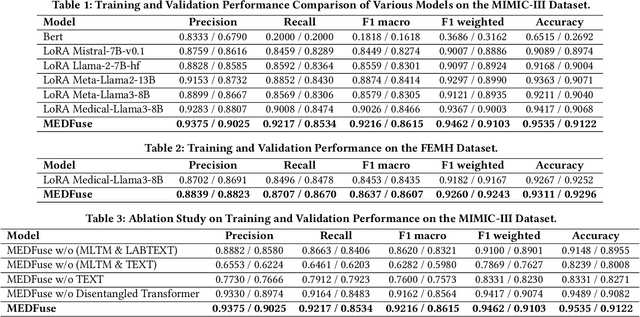
Abstract:Electronic health records (EHRs) are multimodal by nature, consisting of structured tabular features like lab tests and unstructured clinical notes. In real-life clinical practice, doctors use complementary multimodal EHR data sources to get a clearer picture of patients' health and support clinical decision-making. However, most EHR predictive models do not reflect these procedures, as they either focus on a single modality or overlook the inter-modality interactions/redundancy. In this work, we propose MEDFuse, a Multimodal EHR Data Fusion framework that incorporates masked lab-test modeling and large language models (LLMs) to effectively integrate structured and unstructured medical data. MEDFuse leverages multimodal embeddings extracted from two sources: LLMs fine-tuned on free clinical text and masked tabular transformers trained on structured lab test results. We design a disentangled transformer module, optimized by a mutual information loss to 1) decouple modality-specific and modality-shared information and 2) extract useful joint representation from the noise and redundancy present in clinical notes. Through comprehensive validation on the public MIMIC-III dataset and the in-house FEMH dataset, MEDFuse demonstrates great potential in advancing clinical predictions, achieving over 90% F1 score in the 10-disease multi-label classification task.
EHR-Based Mobile and Web Platform for Chronic Disease Risk Prediction Using Large Language Multimodal Models
Jun 26, 2024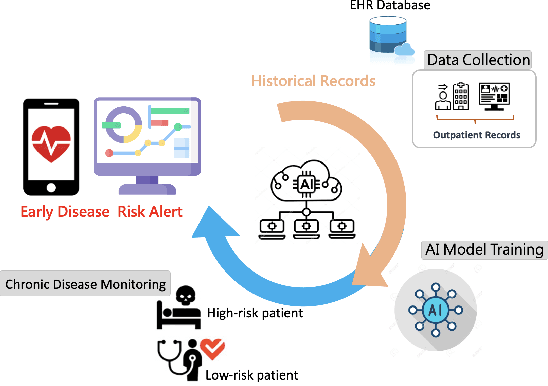
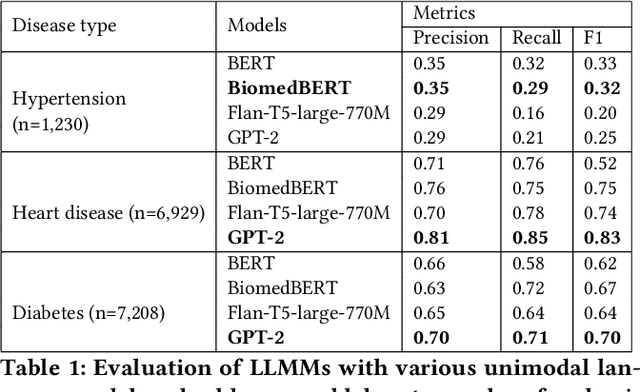
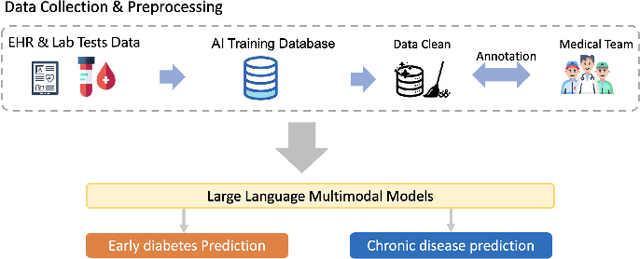
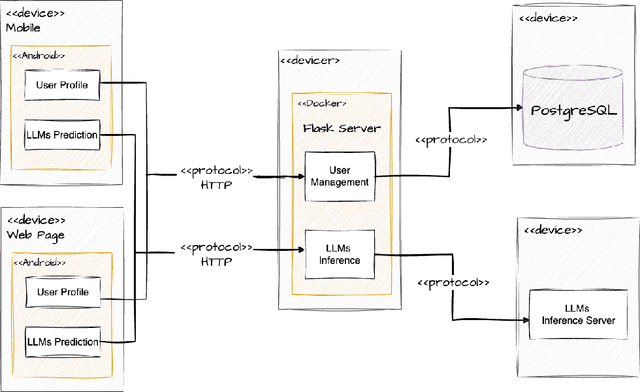
Abstract:Traditional diagnosis of chronic diseases involves in-person consultations with physicians to identify the disease. However, there is a lack of research focused on predicting and developing application systems using clinical notes and blood test values. We collected five years of Electronic Health Records (EHRs) from Taiwan's hospital database between 2017 and 2021 as an AI database. Furthermore, we developed an EHR-based chronic disease prediction platform utilizing Large Language Multimodal Models (LLMMs), successfully integrating with frontend web and mobile applications for prediction. This prediction platform can also connect to the hospital's backend database, providing physicians with real-time risk assessment diagnostics. The demonstration link can be found at https://www.youtube.com/watch?v=oqmL9DEDFgA.
Large Language Multimodal Models for 5-Year Chronic Disease Cohort Prediction Using EHR Data
Mar 02, 2024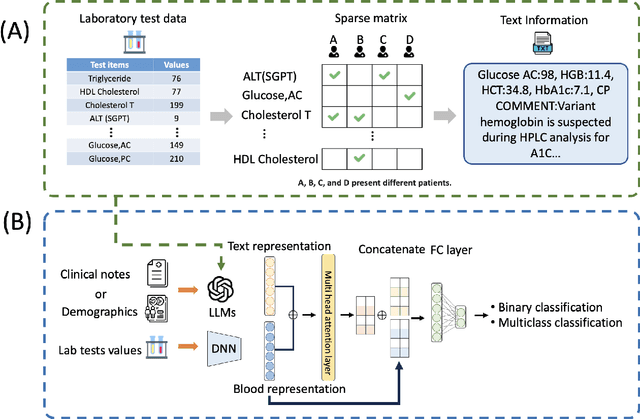
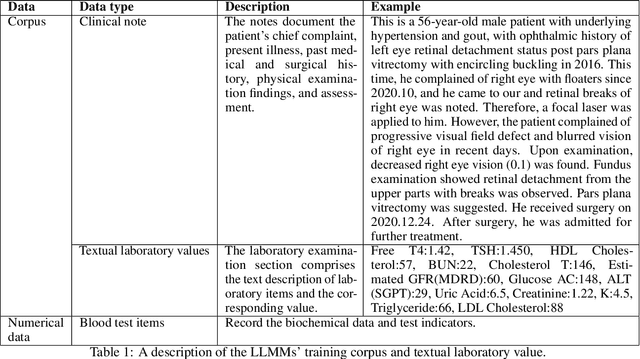

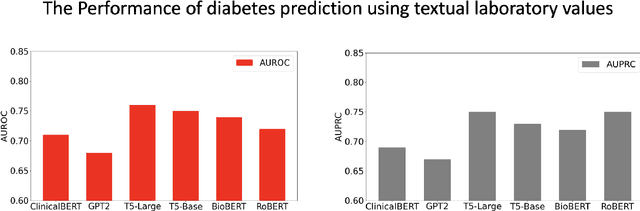
Abstract:Chronic diseases such as diabetes are the leading causes of morbidity and mortality worldwide. Numerous research studies have been attempted with various deep learning models in diagnosis. However, most previous studies had certain limitations, including using publicly available datasets (e.g. MIMIC), and imbalanced data. In this study, we collected five-year electronic health records (EHRs) from the Taiwan hospital database, including 1,420,596 clinical notes, 387,392 laboratory test results, and more than 1,505 laboratory test items, focusing on research pre-training large language models. We proposed a novel Large Language Multimodal Models (LLMMs) framework incorporating multimodal data from clinical notes and laboratory test results for the prediction of chronic disease risk. Our method combined a text embedding encoder and multi-head attention layer to learn laboratory test values, utilizing a deep neural network (DNN) module to merge blood features with chronic disease semantics into a latent space. In our experiments, we observe that clinicalBERT and PubMed-BERT, when combined with attention fusion, can achieve an accuracy of 73% in multiclass chronic diseases and diabetes prediction. By transforming laboratory test values into textual descriptions and employing the Flan T-5 model, we achieved a 76% Area Under the ROC Curve (AUROC), demonstrating the effectiveness of leveraging numerical text data for training and inference in language models. This approach significantly improves the accuracy of early-stage diabetes prediction.
Hospital transfer risk prediction for COVID-19 patients from a medicalized hotel based on Diffusion GraphSAGE
Dec 31, 2022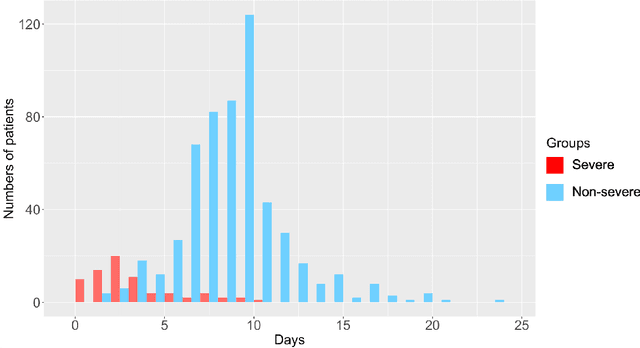
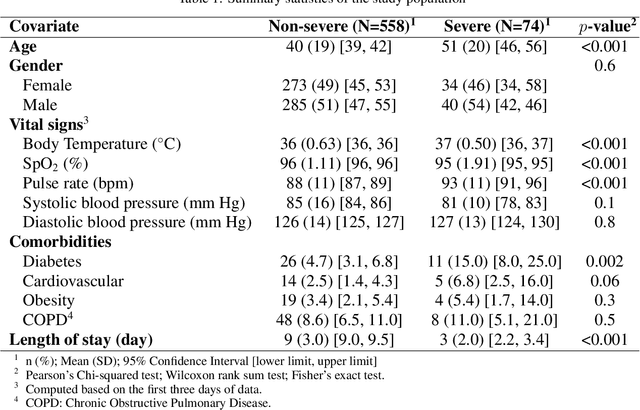
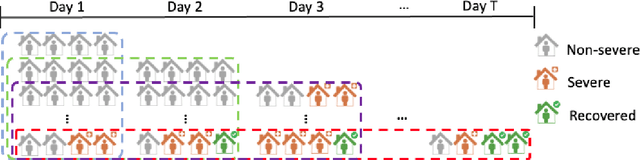
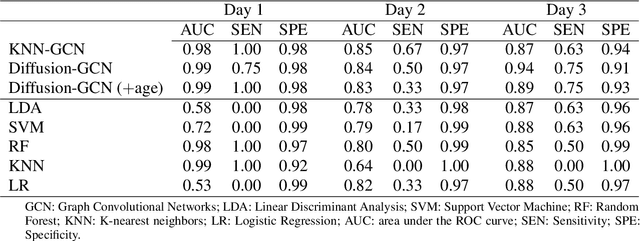
Abstract:The global COVID-19 pandemic has caused more than six million deaths worldwide. Medicalized hotels were established in Taiwan as quarantine facilities for COVID-19 patients with no or mild symptoms. Due to limited medical care available at these hotels, it is of paramount importance to identify patients at risk of clinical deterioration. This study aimed to develop and evaluate a graph-based deep learning approach for progressive hospital transfer risk prediction in a medicalized hotel setting. Vital sign measurements were obtained for 632 patients and daily patient similarity graphs were constructed. Inductive graph convolutional network models were trained on top of the temporally integrated graphs to predict hospital transfer risk. The proposed models achieved AUC scores above 0.83 for hospital transfer risk prediction based on the measurements of past 1, 2, and 3 days, outperforming baseline machine learning methods. A post-hoc analysis on the constructed diffusion-based graph using Local Clustering Coefficient discovered a high-risk cluster with significantly older mean age, higher body temperature, lower SpO2, and shorter length of stay. Further time-to-hospital-transfer survival analysis also revealed a significant decrease in survival probability in the discovered high-risk cluster. The obtained results demonstrated promising predictability and interpretability of the proposed graph-based approach. This technique may help preemptively detect high-risk patients at community-based medical facilities similar to a medicalized hotel.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge