Large Language Multimodal Models for 5-Year Chronic Disease Cohort Prediction Using EHR Data
Paper and Code
Mar 02, 2024
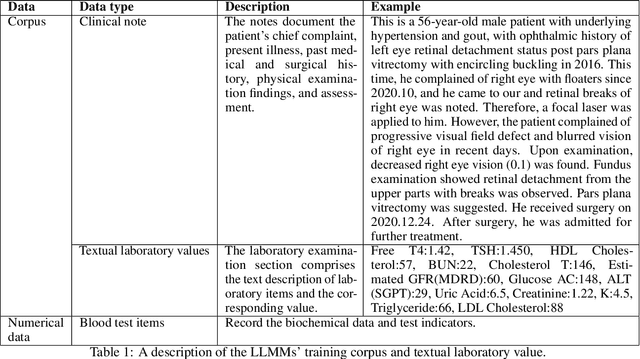

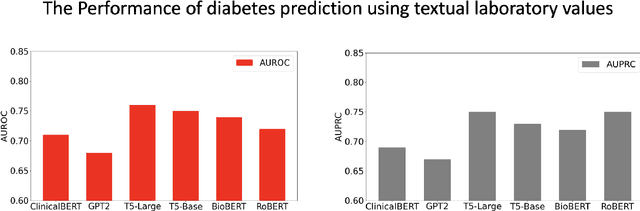
Chronic diseases such as diabetes are the leading causes of morbidity and mortality worldwide. Numerous research studies have been attempted with various deep learning models in diagnosis. However, most previous studies had certain limitations, including using publicly available datasets (e.g. MIMIC), and imbalanced data. In this study, we collected five-year electronic health records (EHRs) from the Taiwan hospital database, including 1,420,596 clinical notes, 387,392 laboratory test results, and more than 1,505 laboratory test items, focusing on research pre-training large language models. We proposed a novel Large Language Multimodal Models (LLMMs) framework incorporating multimodal data from clinical notes and laboratory test results for the prediction of chronic disease risk. Our method combined a text embedding encoder and multi-head attention layer to learn laboratory test values, utilizing a deep neural network (DNN) module to merge blood features with chronic disease semantics into a latent space. In our experiments, we observe that clinicalBERT and PubMed-BERT, when combined with attention fusion, can achieve an accuracy of 73% in multiclass chronic diseases and diabetes prediction. By transforming laboratory test values into textual descriptions and employing the Flan T-5 model, we achieved a 76% Area Under the ROC Curve (AUROC), demonstrating the effectiveness of leveraging numerical text data for training and inference in language models. This approach significantly improves the accuracy of early-stage diabetes prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge