Danielle Bitterman
KScope: A Framework for Characterizing the Knowledge Status of Language Models
Jun 09, 2025Abstract:Characterizing a large language model's (LLM's) knowledge of a given question is challenging. As a result, prior work has primarily examined LLM behavior under knowledge conflicts, where the model's internal parametric memory contradicts information in the external context. However, this does not fully reflect how well the model knows the answer to the question. In this paper, we first introduce a taxonomy of five knowledge statuses based on the consistency and correctness of LLM knowledge modes. We then propose KScope, a hierarchical framework of statistical tests that progressively refines hypotheses about knowledge modes and characterizes LLM knowledge into one of these five statuses. We apply KScope to nine LLMs across four datasets and systematically establish: (1) Supporting context narrows knowledge gaps across models. (2) Context features related to difficulty, relevance, and familiarity drive successful knowledge updates. (3) LLMs exhibit similar feature preferences when partially correct or conflicted, but diverge sharply when consistently wrong. (4) Context summarization constrained by our feature analysis, together with enhanced credibility, further improves update effectiveness and generalizes across LLMs.
Multi-OphthaLingua: A Multilingual Benchmark for Assessing and Debiasing LLM Ophthalmological QA in LMICs
Dec 18, 2024



Abstract:Current ophthalmology clinical workflows are plagued by over-referrals, long waits, and complex and heterogeneous medical records. Large language models (LLMs) present a promising solution to automate various procedures such as triaging, preliminary tests like visual acuity assessment, and report summaries. However, LLMs have demonstrated significantly varied performance across different languages in natural language question-answering tasks, potentially exacerbating healthcare disparities in Low and Middle-Income Countries (LMICs). This study introduces the first multilingual ophthalmological question-answering benchmark with manually curated questions parallel across languages, allowing for direct cross-lingual comparisons. Our evaluation of 6 popular LLMs across 7 different languages reveals substantial bias across different languages, highlighting risks for clinical deployment of LLMs in LMICs. Existing debiasing methods such as Translation Chain-of-Thought or Retrieval-augmented generation (RAG) by themselves fall short of closing this performance gap, often failing to improve performance across all languages and lacking specificity for the medical domain. To address this issue, We propose CLARA (Cross-Lingual Reflective Agentic system), a novel inference time de-biasing method leveraging retrieval augmented generation and self-verification. Our approach not only improves performance across all languages but also significantly reduces the multilingual bias gap, facilitating equitable LLM application across the globe.
ClinicalBench: Can LLMs Beat Traditional ML Models in Clinical Prediction?
Nov 10, 2024Abstract:Large Language Models (LLMs) hold great promise to revolutionize current clinical systems for their superior capacities on medical text processing tasks and medical licensing exams. Meanwhile, traditional ML models such as SVM and XGBoost have still been mainly adopted in clinical prediction tasks. An emerging question is Can LLMs beat traditional ML models in clinical prediction? Thus, we build a new benchmark ClinicalBench to comprehensively study the clinical predictive modeling capacities of both general-purpose and medical LLMs, and compare them with traditional ML models. ClinicalBench embraces three common clinical prediction tasks, two databases, 14 general-purpose LLMs, 8 medical LLMs, and 11 traditional ML models. Through extensive empirical investigation, we discover that both general-purpose and medical LLMs, even with different model scales, diverse prompting or fine-tuning strategies, still cannot beat traditional ML models in clinical prediction yet, shedding light on their potential deficiency in clinical reasoning and decision-making. We call for caution when practitioners adopt LLMs in clinical applications. ClinicalBench can be utilized to bridge the gap between LLMs' development for healthcare and real-world clinical practice.
Position Paper On Diagnostic Uncertainty Estimation from Large Language Models: Next-Word Probability Is Not Pre-test Probability
Nov 07, 2024Abstract:Large language models (LLMs) are being explored for diagnostic decision support, yet their ability to estimate pre-test probabilities, vital for clinical decision-making, remains limited. This study evaluates two LLMs, Mistral-7B and Llama3-70B, using structured electronic health record data on three diagnosis tasks. We examined three current methods of extracting LLM probability estimations and revealed their limitations. We aim to highlight the need for improved techniques in LLM confidence estimation.
Mapping Bias in Vision Language Models: Signposts, Pitfalls, and the Road Ahead
Oct 17, 2024


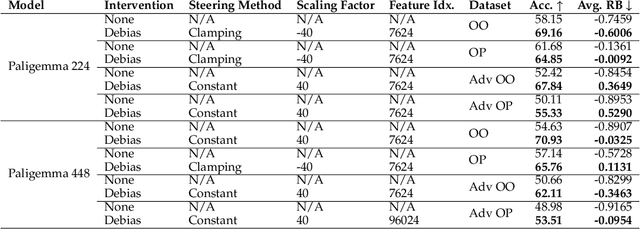
Abstract:As Vision Language Models (VLMs) gain widespread use, their fairness remains under-explored. In this paper, we analyze demographic biases across five models and six datasets. We find that portrait datasets like UTKFace and CelebA are the best tools for bias detection, finding gaps in performance and fairness between LLaVa and CLIP models. However, scene based datasets like PATA, VLStereoSet fail to be useful benchmarks for bias due to their construction. As for pronoun based datasets like VisoGender, we receive mixed signals as only some subsets of the data are useful in providing insights. To alleviate this problem, we introduce a more difficult version of VisoGender to serve as a more rigorous evaluation. Based on these results, we call for more effective and carefully designed datasets to ensure VLMs are both fair and reliable.
Wait, but Tylenol is Acetaminophen... Investigating and Improving Language Models' Ability to Resist Requests for Misinformation
Sep 30, 2024Abstract:Background: Large language models (LLMs) are trained to follow directions, but this introduces a vulnerability to blindly comply with user requests even if they generate wrong information. In medicine, this could accelerate the generation of misinformation that impacts human well-being. Objectives/Methods: We analyzed compliance to requests to generate misleading content about medications in settings where models know the request is illogical. We investigated whether in-context directions and instruction-tuning of LLMs to prioritize logical reasoning over compliance reduced misinformation risk. Results: While all frontier LLMs complied with misinformation requests, both prompt-based and parameter-based approaches can improve the detection of logic flaws in requests and prevent the dissemination of medical misinformation. Conclusion: Shifting LLMs to prioritize logic over compliance could reduce risks of exploitation for medical misinformation.
When Raw Data Prevails: Are Large Language Model Embeddings Effective in Numerical Data Representation for Medical Machine Learning Applications?
Aug 15, 2024



Abstract:The introduction of Large Language Models (LLMs) has advanced data representation and analysis, bringing significant progress in their use for medical questions and answering. Despite these advancements, integrating tabular data, especially numerical data pivotal in clinical contexts, into LLM paradigms has not been thoroughly explored. In this study, we examine the effectiveness of vector representations from last hidden states of LLMs for medical diagnostics and prognostics using electronic health record (EHR) data. We compare the performance of these embeddings with that of raw numerical EHR data when used as feature inputs to traditional machine learning (ML) algorithms that excel at tabular data learning, such as eXtreme Gradient Boosting. We focus on instruction-tuned LLMs in a zero-shot setting to represent abnormal physiological data and evaluating their utilities as feature extractors to enhance ML classifiers for predicting diagnoses, length of stay, and mortality. Furthermore, we examine prompt engineering techniques on zero-shot and few-shot LLM embeddings to measure their impact comprehensively. Although findings suggest the raw data features still prevails in medical ML tasks, zero-shot LLM embeddings demonstrate competitive results, suggesting a promising avenue for future research in medical applications.
Language Models are Surprisingly Fragile to Drug Names in Biomedical Benchmarks
Jun 17, 2024Abstract:Medical knowledge is context-dependent and requires consistent reasoning across various natural language expressions of semantically equivalent phrases. This is particularly crucial for drug names, where patients often use brand names like Advil or Tylenol instead of their generic equivalents. To study this, we create a new robustness dataset, RABBITS, to evaluate performance differences on medical benchmarks after swapping brand and generic drug names using physician expert annotations. We assess both open-source and API-based LLMs on MedQA and MedMCQA, revealing a consistent performance drop ranging from 1-10\%. Furthermore, we identify a potential source of this fragility as the contamination of test data in widely used pre-training datasets. All code is accessible at https://github.com/BittermanLab/RABBITS, and a HuggingFace leaderboard is available at https://huggingface.co/spaces/AIM-Harvard/rabbits-leaderboard.
Measuring Pointwise $\mathcal{V}$-Usable Information In-Context-ly
Oct 18, 2023



Abstract:In-context learning (ICL) is a new learning paradigm that has gained popularity along with the development of large language models. In this work, we adapt a recently proposed hardness metric, pointwise $\mathcal{V}$-usable information (PVI), to an in-context version (in-context PVI). Compared to the original PVI, in-context PVI is more efficient in that it requires only a few exemplars and does not require fine-tuning. We conducted a comprehensive empirical analysis to evaluate the reliability of in-context PVI. Our findings indicate that in-context PVI estimates exhibit similar characteristics to the original PVI. Specific to the in-context setting, we show that in-context PVI estimates remain consistent across different exemplar selections and numbers of shots. The variance of in-context PVI estimates across different exemplar selections is insignificant, which suggests that in-context PVI are stable. Furthermore, we demonstrate how in-context PVI can be employed to identify challenging instances. Our work highlights the potential of in-context PVI and provides new insights into the capabilities of ICL.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge