Cheng Ge
Why GRPO Needs Normalization: A Local-Curvature Perspective on Adaptive Gradients
Jan 30, 2026Abstract:Reinforcement learning (RL) has become a key driver of language model reasoning. Among RL algorithms, Group Relative Policy Optimization (GRPO) is the de facto standard, avoiding the need for a critic by using per-prompt baselines and variance normalization. Yet why and when this normalization helps remains unclear. In this work, we provide an explanation through the lens of local curvature of the sequence-level policy gradient: standard deviation normalization implements an adaptive gradient. Theoretically, under mild conditions, GRPO enjoys a strictly improved convergence rate over unnormalized REINFORCE, with gains characterized by the average within-prompt reward standard deviation across prompts and iterations. Empirically, our analysis on GSM8K and MATH benchmarks reveals three distinct training phases governed by the interplay between feature orthogonality and reward variance: (I) an early acceleration phase where high variance and orthogonality favor adaptive scaling; (II) a relatively stable transition phase; and (III) a late-stage regime where the loss of orthogonality limits further gains. Together, these results provide a principled account of when std normalization helps in GRPO, and offer broader insights into the design of critic-free RL algorithms.
CreoPep: A Universal Deep Learning Framework for Target-Specific Peptide Design and Optimization
May 05, 2025Abstract:Target-specific peptides, such as conotoxins, exhibit exceptional binding affinity and selectivity toward ion channels and receptors. However, their therapeutic potential remains underutilized due to the limited diversity of natural variants and the labor-intensive nature of traditional optimization strategies. Here, we present CreoPep, a deep learning-based conditional generative framework that integrates masked language modeling with a progressive masking scheme to design high-affinity peptide mutants while uncovering novel structural motifs. CreoPep employs an integrative augmentation pipeline, combining FoldX-based energy screening with temperature-controlled multinomial sampling, to generate structurally and functionally diverse peptides that retain key pharmacological properties. We validate this approach by designing conotoxin inhibitors targeting the $\alpha$7 nicotinic acetylcholine receptor, achieving submicromolar potency in electrophysiological assays. Structural analysis reveals that CreoPep-generated variants engage in both conserved and novel binding modes, including disulfide-deficient forms, thus expanding beyond conventional design paradigms. Overall, CreoPep offers a robust and generalizable platform that bridges computational peptide design with experimental validation, accelerating the discovery of next-generation peptide therapeutics.
Solving Minimum-Cost Reach Avoid using Reinforcement Learning
Oct 29, 2024Abstract:Current reinforcement-learning methods are unable to directly learn policies that solve the minimum cost reach-avoid problem to minimize cumulative costs subject to the constraints of reaching the goal and avoiding unsafe states, as the structure of this new optimization problem is incompatible with current methods. Instead, a surrogate problem is solved where all objectives are combined with a weighted sum. However, this surrogate objective results in suboptimal policies that do not directly minimize the cumulative cost. In this work, we propose RC-PPO, a reinforcement-learning-based method for solving the minimum-cost reach-avoid problem by using connections to Hamilton-Jacobi reachability. Empirical results demonstrate that RC-PPO learns policies with comparable goal-reaching rates to while achieving up to 57% lower cumulative costs compared to existing methods on a suite of minimum-cost reach-avoid benchmarks on the Mujoco simulator. The project page can be found at https://oswinso.xyz/rcppo.
Automatic Organ and Pan-cancer Segmentation in Abdomen CT: the FLARE 2023 Challenge
Aug 22, 2024Abstract:Organ and cancer segmentation in abdomen Computed Tomography (CT) scans is the prerequisite for precise cancer diagnosis and treatment. Most existing benchmarks and algorithms are tailored to specific cancer types, limiting their ability to provide comprehensive cancer analysis. This work presents the first international competition on abdominal organ and pan-cancer segmentation by providing a large-scale and diverse dataset, including 4650 CT scans with various cancer types from over 40 medical centers. The winning team established a new state-of-the-art with a deep learning-based cascaded framework, achieving average Dice Similarity Coefficient scores of 92.3% for organs and 64.9% for lesions on the hidden multi-national testing set. The dataset and code of top teams are publicly available, offering a benchmark platform to drive further innovations https://codalab.lisn.upsaclay.fr/competitions/12239.
Unleashing the Strengths of Unlabeled Data in Pan-cancer Abdominal Organ Quantification: the FLARE22 Challenge
Aug 10, 2023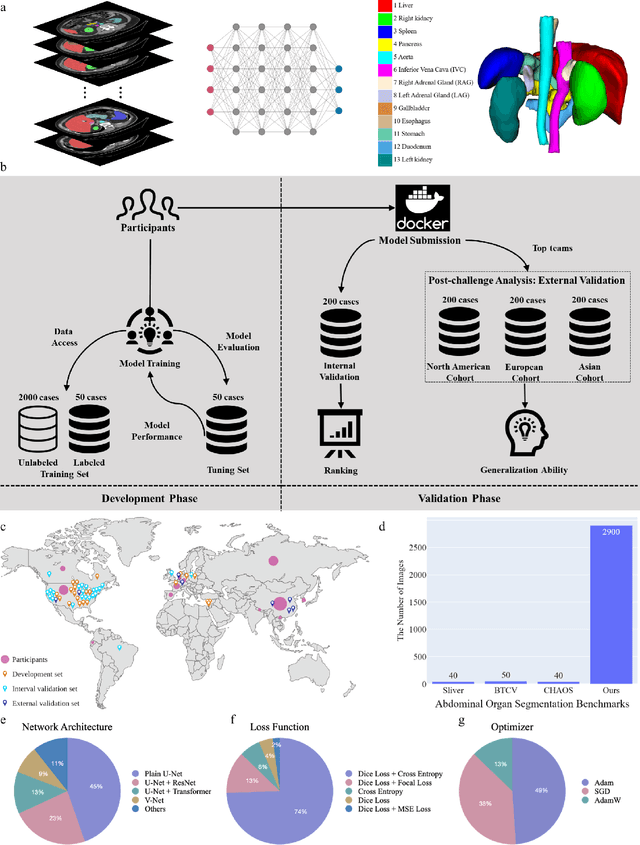
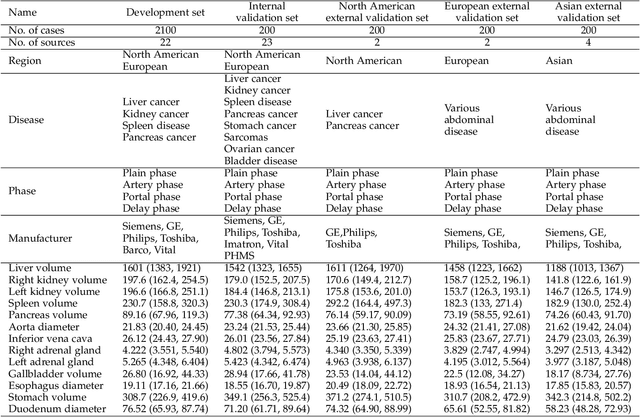
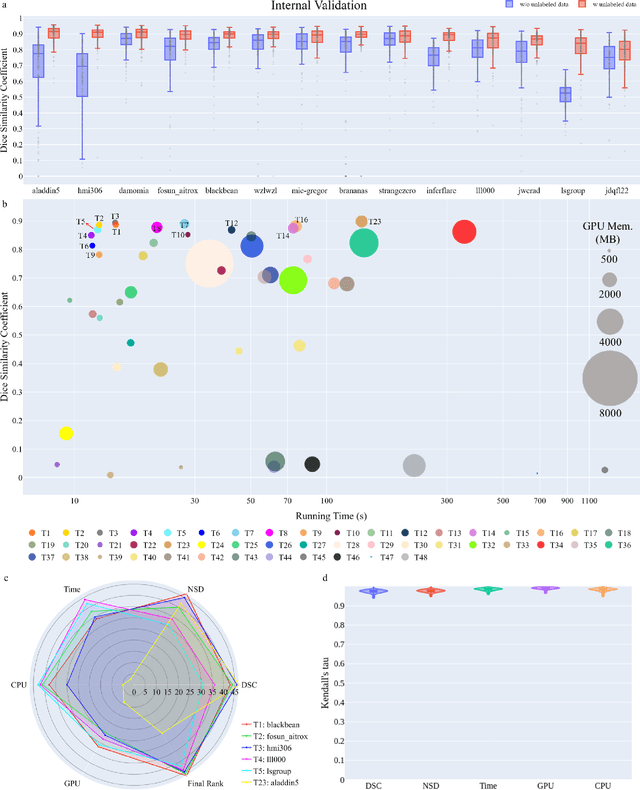
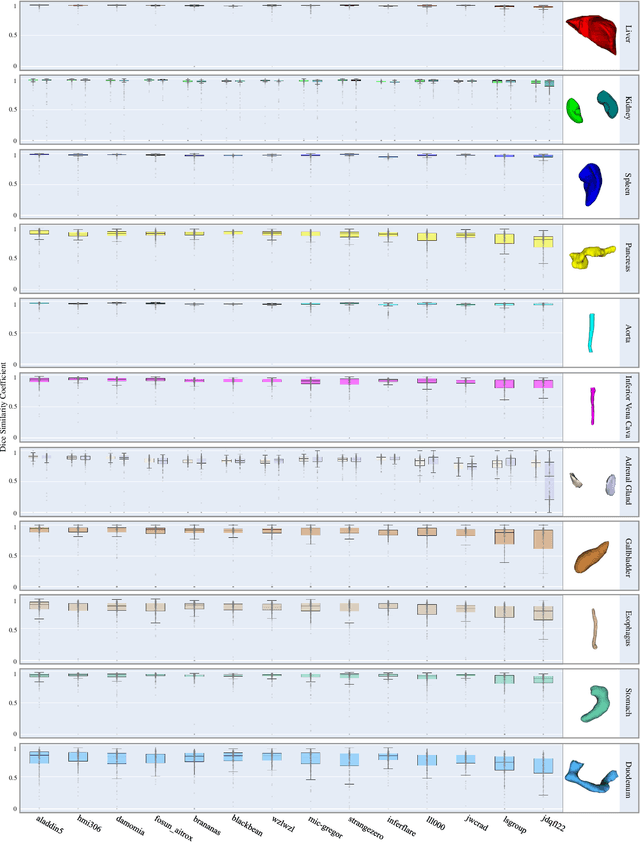
Abstract:Quantitative organ assessment is an essential step in automated abdominal disease diagnosis and treatment planning. Artificial intelligence (AI) has shown great potential to automatize this process. However, most existing AI algorithms rely on many expert annotations and lack a comprehensive evaluation of accuracy and efficiency in real-world multinational settings. To overcome these limitations, we organized the FLARE 2022 Challenge, the largest abdominal organ analysis challenge to date, to benchmark fast, low-resource, accurate, annotation-efficient, and generalized AI algorithms. We constructed an intercontinental and multinational dataset from more than 50 medical groups, including Computed Tomography (CT) scans with different races, diseases, phases, and manufacturers. We independently validated that a set of AI algorithms achieved a median Dice Similarity Coefficient (DSC) of 90.0\% by using 50 labeled scans and 2000 unlabeled scans, which can significantly reduce annotation requirements. The best-performing algorithms successfully generalized to holdout external validation sets, achieving a median DSC of 89.5\%, 90.9\%, and 88.3\% on North American, European, and Asian cohorts, respectively. They also enabled automatic extraction of key organ biology features, which was labor-intensive with traditional manual measurements. This opens the potential to use unlabeled data to boost performance and alleviate annotation shortages for modern AI models.
The Multi-modality Cell Segmentation Challenge: Towards Universal Solutions
Aug 10, 2023Abstract:Cell segmentation is a critical step for quantitative single-cell analysis in microscopy images. Existing cell segmentation methods are often tailored to specific modalities or require manual interventions to specify hyperparameters in different experimental settings. Here, we present a multi-modality cell segmentation benchmark, comprising over 1500 labeled images derived from more than 50 diverse biological experiments. The top participants developed a Transformer-based deep-learning algorithm that not only exceeds existing methods, but can also be applied to diverse microscopy images across imaging platforms and tissue types without manual parameter adjustments. This benchmark and the improved algorithm offer promising avenues for more accurate and versatile cell analysis in microscopy imaging.
Cost Splitting for Multi-Objective Conflict-Based Search
Nov 23, 2022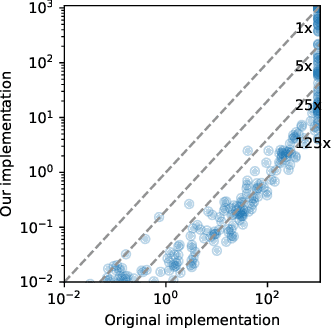
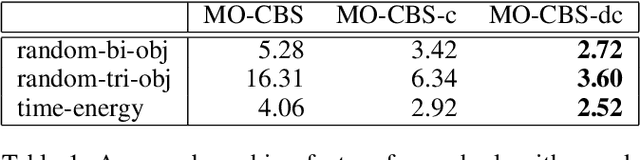

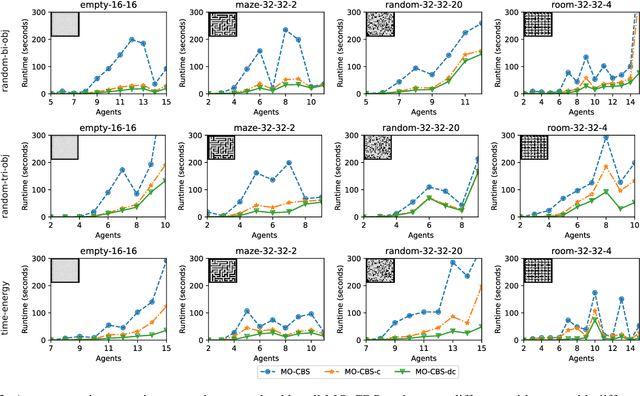
Abstract:The Multi-Objective Multi-Agent Path Finding (MO-MAPF) problem is the problem of finding the Pareto-optimal frontier of collision-free paths for a team of agents while minimizing multiple cost metrics. Examples of such cost metrics include arrival times, travel distances, and energy consumption.In this paper, we focus on the Multi-Objective Conflict-Based Search (MO-CBS) algorithm, a state-of-the-art MO-MAPF algorithm. We show that the standard splitting strategy used by MO-CBS can lead to duplicate search nodes and hence can duplicate the search effort that MO-CBS needs to make. To address this issue, we propose two new splitting strategies for MO-CBS, namely cost splitting and disjoint cost splitting. Our theoretical results show that, when combined with either of these two new splitting strategies, MO-CBS maintains its completeness and optimality guarantees. Our experimental results show that disjoint cost splitting, our best splitting strategy, speeds up MO-CBS by up to two orders of magnitude and substantially improves its success rates in various settings.
Deep Baseline Network for Time Series Modeling and Anomaly Detection
Sep 10, 2022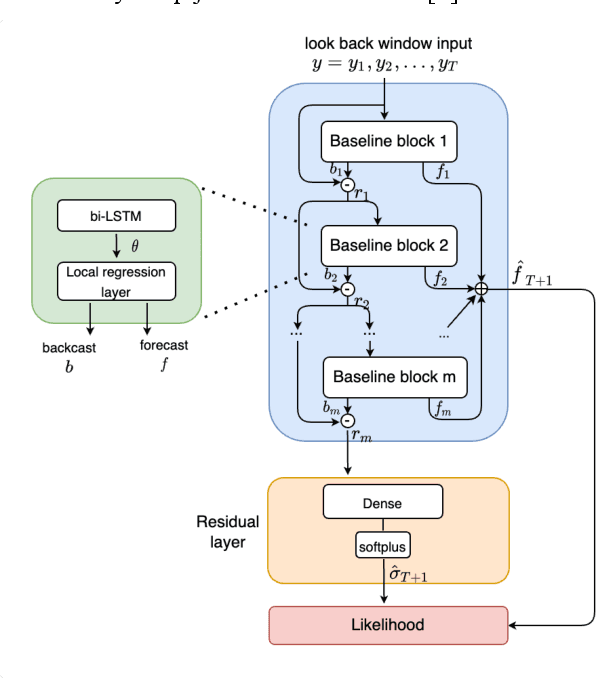
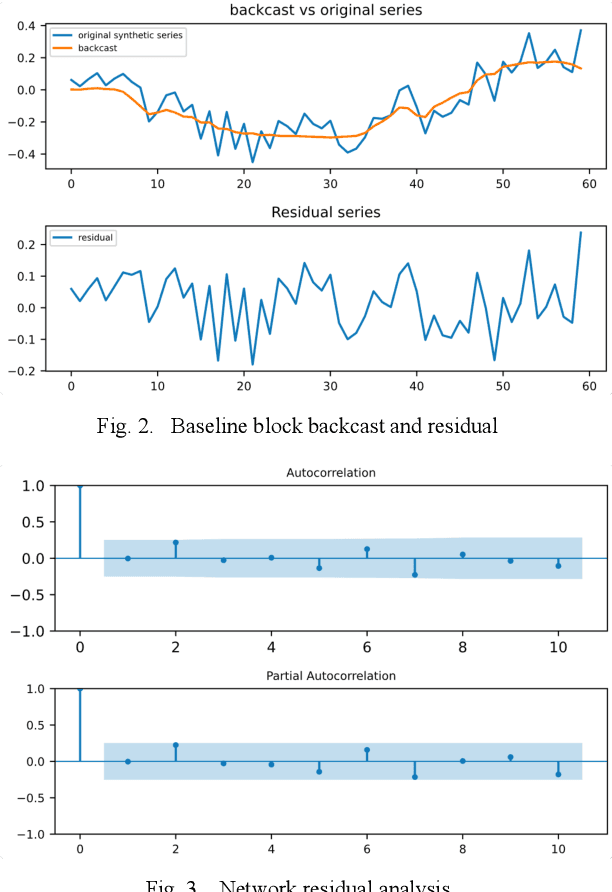
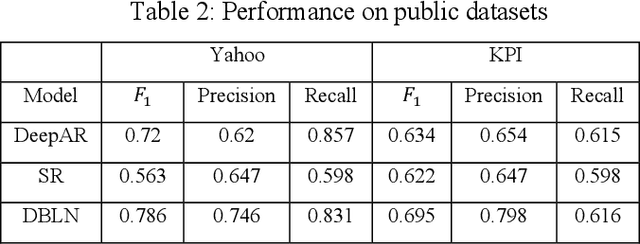
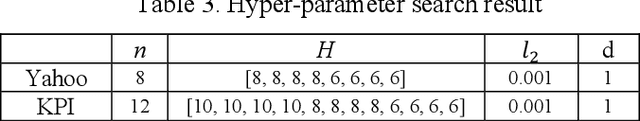
Abstract:Deep learning has seen increasing applications in time series in recent years. For time series anomaly detection scenarios, such as in finance, Internet of Things, data center operations, etc., time series usually show very flexible baselines depending on various external factors. Anomalies unveil themselves by lying far away from the baseline. However, the detection is not always easy due to some challenges including baseline shifting, lacking of labels, noise interference, real time detection in streaming data, result interpretability, etc. In this paper, we develop a novel deep architecture to properly extract the baseline from time series, namely Deep Baseline Network (DBLN). By using this deep network, we can easily locate the baseline position and then provide reliable and interpretable anomaly detection result. Empirical evaluation on both synthetic and public real-world datasets shows that our purely unsupervised algorithm achieves superior performance compared with state-of-art methods and has good practical applications.
DePS: An improved deep learning model for de novo peptide sequencing
Mar 16, 2022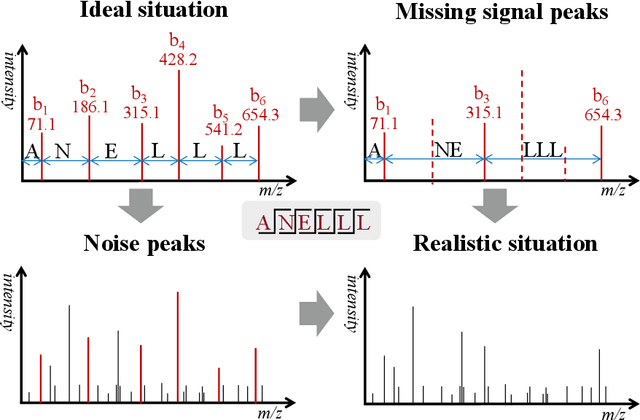
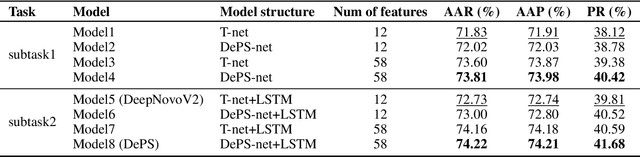
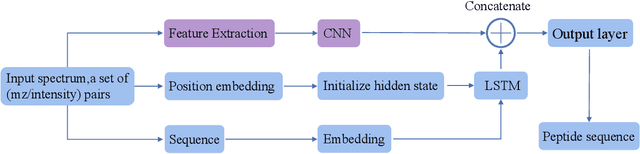
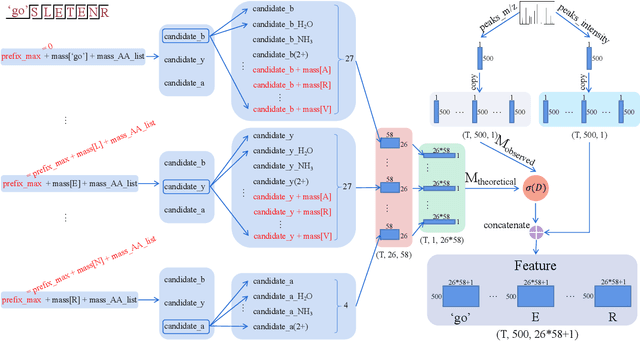
Abstract:De novo peptide sequencing from mass spectrometry data is an important method for protein identification. Recently, various deep learning approaches were applied for de novo peptide sequencing and DeepNovoV2 is one of the represetative models. In this study, we proposed an enhanced model, DePS, which can improve the accuracy of de novo peptide sequencing even with missing signal peaks or large number of noisy peaks in tandem mass spectrometry data. It is showed that, for the same test set of DeepNovoV2, the DePS model achieved excellent results of 74.22%, 74.21% and 41.68% for amino acid recall, amino acid precision and peptide recall respectively. Furthermore, the results suggested that DePS outperforms DeepNovoV2 on the cross species dataset.
AbdomenCT-1K: Is Abdominal Organ Segmentation A Solved Problem?
Oct 28, 2020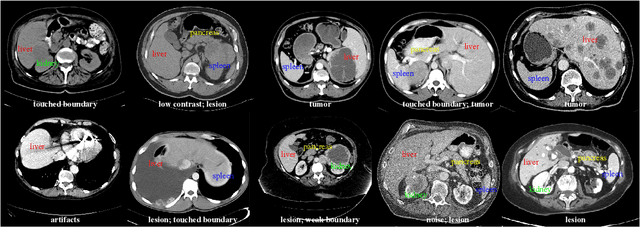
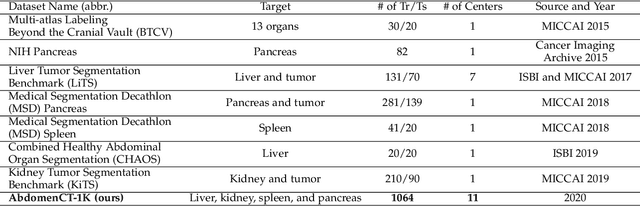
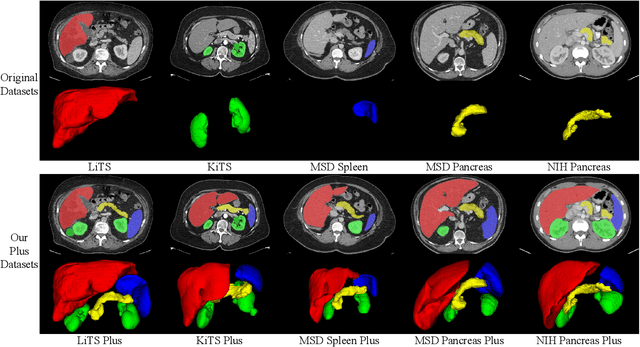
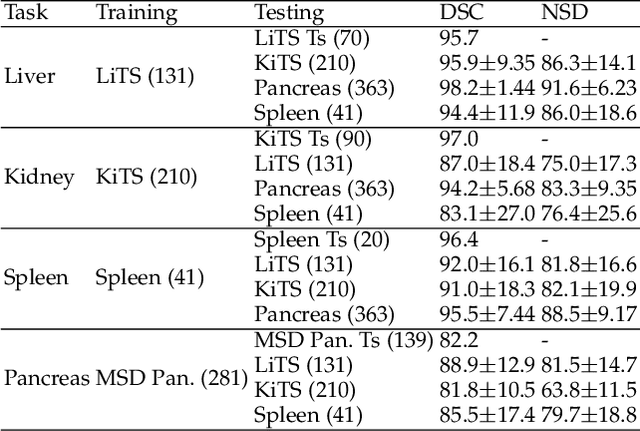
Abstract:With the unprecedented developments in deep learning, automatic segmentation of main abdominal organs (i.e., liver, kidney, and spleen) seems to be a solved problem as the state-of-the-art (SOTA) methods have achieved comparable results with inter-observer variability on existing benchmark datasets. However, most of the existing abdominal organ segmentation benchmark datasets only contain single-center, single-phase, single-vendor, or single-disease cases, thus, it is unclear whether the excellent performance can generalize on more diverse datasets. In this paper, we present a large and diverse abdominal CT organ segmentation dataset, termed as AbdomenCT-1K, with more than 1000 (1K) CT scans from 11 countries, including multi-center, multi-phase, multi-vendor, and multi-disease cases. Furthermore, we conduct a large-scale study for liver, kidney, spleen, and pancreas segmentation, as well as reveal the unsolved segmentation problems of the SOTA method, such as the limited generalization ability on distinct medical centers, phases, and unseen diseases. To advance the unsolved problems, we build four organ segmentation benchmarks for fully supervised, semi-supervised, weakly supervised, and continual learning, which are currently challenging and active research topics. Accordingly, we develop a simple and effective method for each benchmark, which can be used as out-of-the-box methods and strong baselines. We believe the introduction of the AbdomenCT-1K dataset will promote future in-depth research towards clinical applicable abdominal organ segmentation methods. Moreover, the datasets, codes, and trained models of baseline methods will be publicly available at https://github.com/JunMa11/AbdomenCT-1K.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge