Guoqiang Dong
Unleashing the Strengths of Unlabeled Data in Pan-cancer Abdominal Organ Quantification: the FLARE22 Challenge
Aug 10, 2023
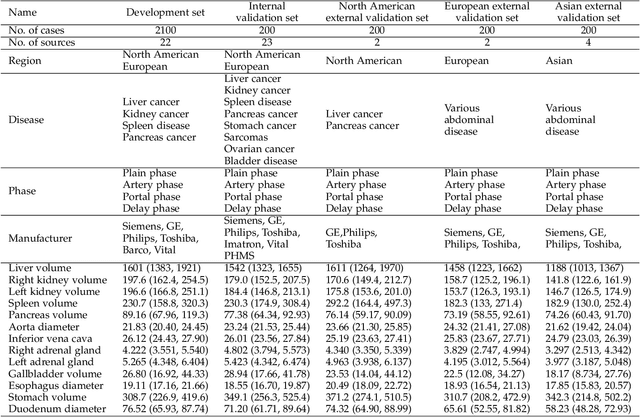
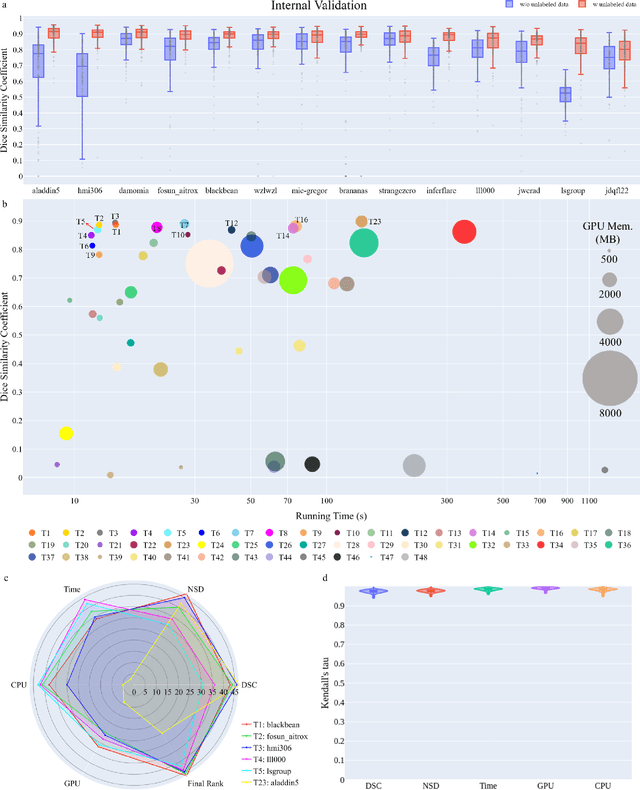
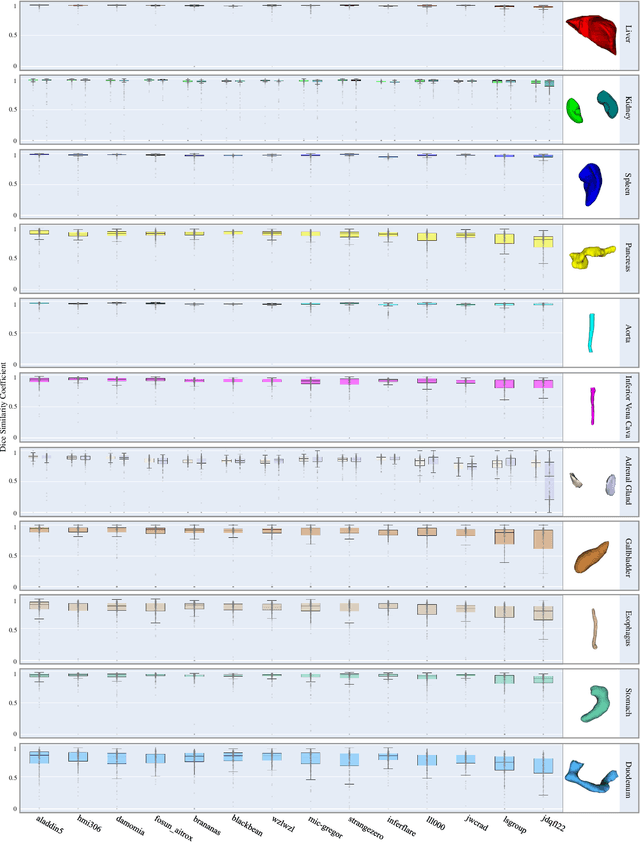
Abstract:Quantitative organ assessment is an essential step in automated abdominal disease diagnosis and treatment planning. Artificial intelligence (AI) has shown great potential to automatize this process. However, most existing AI algorithms rely on many expert annotations and lack a comprehensive evaluation of accuracy and efficiency in real-world multinational settings. To overcome these limitations, we organized the FLARE 2022 Challenge, the largest abdominal organ analysis challenge to date, to benchmark fast, low-resource, accurate, annotation-efficient, and generalized AI algorithms. We constructed an intercontinental and multinational dataset from more than 50 medical groups, including Computed Tomography (CT) scans with different races, diseases, phases, and manufacturers. We independently validated that a set of AI algorithms achieved a median Dice Similarity Coefficient (DSC) of 90.0\% by using 50 labeled scans and 2000 unlabeled scans, which can significantly reduce annotation requirements. The best-performing algorithms successfully generalized to holdout external validation sets, achieving a median DSC of 89.5\%, 90.9\%, and 88.3\% on North American, European, and Asian cohorts, respectively. They also enabled automatic extraction of key organ biology features, which was labor-intensive with traditional manual measurements. This opens the potential to use unlabeled data to boost performance and alleviate annotation shortages for modern AI models.
Towards Efficient COVID-19 CT Annotation: A Benchmark for Lung and Infection Segmentation
Apr 27, 2020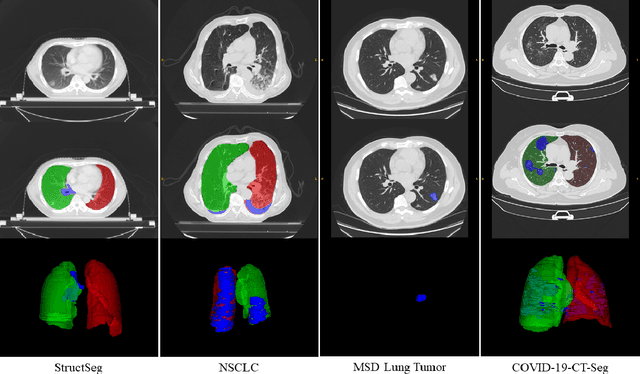

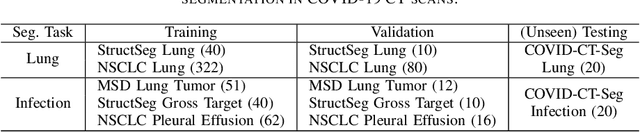
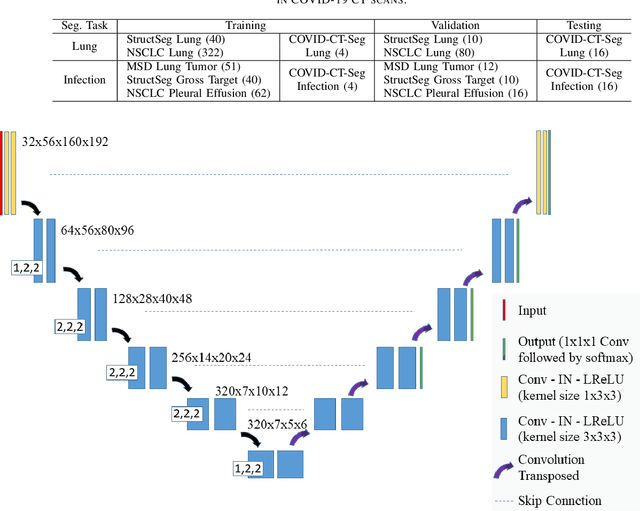
Abstract:Accurate segmentation of lung and infection in COVID-19 CT scans plays an important role in the quantitative management of patients. Most of the existing studies are based on large and private annotated datasets that are impractical to obtain from a single institution, especially when radiologists are busy fighting the coronavirus disease. Furthermore, it is hard to compare current COVID-19 CT segmentation methods as they are developed on different datasets, trained in different settings, and evaluated with different metrics. In this paper, we created a COVID-19 3D CT dataset with 20 cases that contains 1800+ annotated slices and made it publicly available. To promote the development of annotation-efficient deep learning methods, we built three benchmarks for lung and infection segmentation that contain current main research interests, e.g., few-shot learning, domain generalization, and knowledge transfer. For a fair comparison among different segmentation methods, we also provide unified training, validation and testing dataset splits, and evaluation metrics and corresponding code. In addition, we provided more than 40 pre-trained baseline models for the benchmarks, which not only serve as out-of-the-box segmentation tools but also save computational time for researchers who are interested in COVID-19 lung and infection segmentation. To the best of our knowledge, this work presents the largest public annotated COVID-19 CT volume dataset, the first segmentation benchmark, and the most pre-trained models up to now. We hope these resources (\url{https://gitee.com/junma11/COVID-19-CT-Seg-Benchmark}) could advance the development of deep learning methods for COVID-19 CT segmentation with limited data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge