Jia Qu
ViSA: Visited-State Augmentation for Generalized Goal-Space Contrastive Reinforcement Learning
Mar 16, 2026Abstract:Goal-Conditioned Reinforcement Learning (GCRL) is a framework for learning a policy that can reach arbitrarily given goals. In particular, Contrastive Reinforcement Learning (CRL) provides a framework for policy updates using an approximation of the value function estimated via contrastive learning, achieving higher sample efficiency compared to conventional methods. However, since CRL treats the visited state as a pseudo-goal during learning, it can accurately estimate the value function only for limited goals. To address this issue, we propose a novel data augmentation approach for CRL called ViSA (Visited-State Augmentation). ViSA consists of two components: 1) generating augmented state samples, with the aim of augmenting hard-to-visit state samples during on-policy exploration, and 2) learning consistent embedding space, which uses an augmented state as auxiliary information to regularize the embedding space by reformulating the objective function of the embedding space based on mutual information. We evaluate ViSA in simulation and real-world robotic tasks and show improved goal-space generalization, which permits accurate value estimation for hard-to-visit goals. Further details can be found on the project page: \href{https://issa-n.github.io/projectPage_ViSA/}{\texttt{https://issa-n.github.io/projectPage\_ViSA/}}
DePS: An improved deep learning model for de novo peptide sequencing
Mar 16, 2022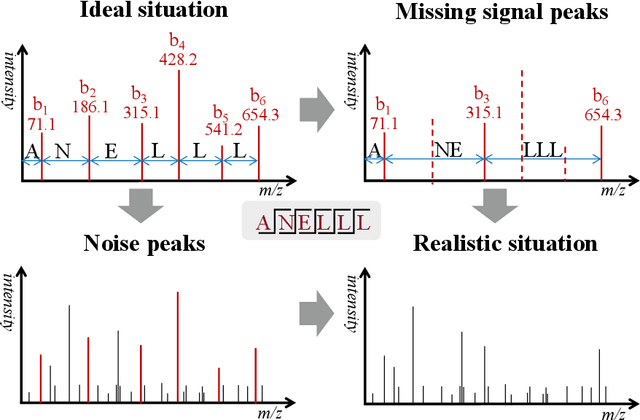
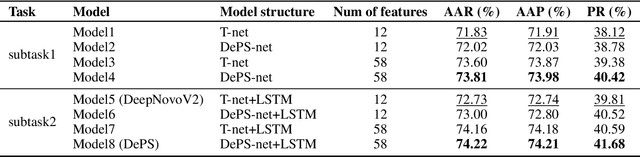
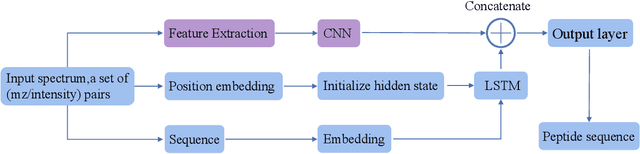
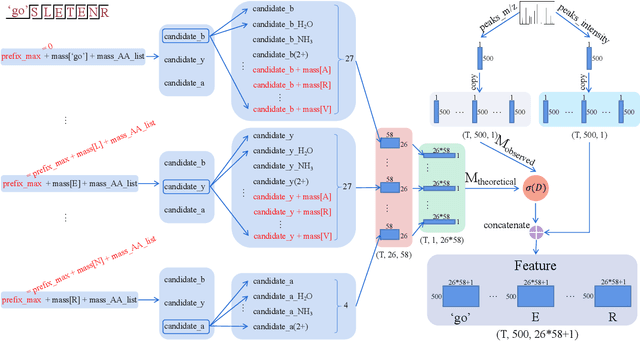
Abstract:De novo peptide sequencing from mass spectrometry data is an important method for protein identification. Recently, various deep learning approaches were applied for de novo peptide sequencing and DeepNovoV2 is one of the represetative models. In this study, we proposed an enhanced model, DePS, which can improve the accuracy of de novo peptide sequencing even with missing signal peaks or large number of noisy peaks in tandem mass spectrometry data. It is showed that, for the same test set of DeepNovoV2, the DePS model achieved excellent results of 74.22%, 74.21% and 41.68% for amino acid recall, amino acid precision and peptide recall respectively. Furthermore, the results suggested that DePS outperforms DeepNovoV2 on the cross species dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge