Shubham Innani
KPIs 2024 Challenge: Advancing Glomerular Segmentation from Patch- to Slide-Level
Feb 11, 2025
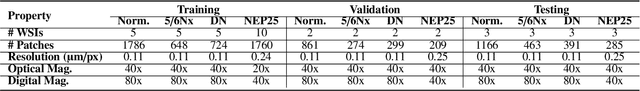
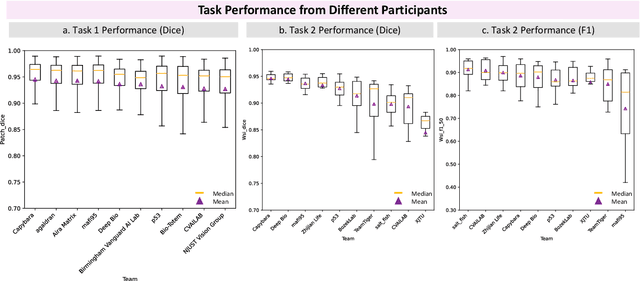

Abstract:Chronic kidney disease (CKD) is a major global health issue, affecting over 10% of the population and causing significant mortality. While kidney biopsy remains the gold standard for CKD diagnosis and treatment, the lack of comprehensive benchmarks for kidney pathology segmentation hinders progress in the field. To address this, we organized the Kidney Pathology Image Segmentation (KPIs) Challenge, introducing a dataset that incorporates preclinical rodent models of CKD with over 10,000 annotated glomeruli from 60+ Periodic Acid Schiff (PAS)-stained whole slide images. The challenge includes two tasks, patch-level segmentation and whole slide image segmentation and detection, evaluated using the Dice Similarity Coefficient (DSC) and F1-score. By encouraging innovative segmentation methods that adapt to diverse CKD models and tissue conditions, the KPIs Challenge aims to advance kidney pathology analysis, establish new benchmarks, and enable precise, large-scale quantification for disease research and diagnosis.
BraTS-Path Challenge: Assessing Heterogeneous Histopathologic Brain Tumor Sub-regions
May 17, 2024Abstract:Glioblastoma is the most common primary adult brain tumor, with a grim prognosis - median survival of 12-18 months following treatment, and 4 months otherwise. Glioblastoma is widely infiltrative in the cerebral hemispheres and well-defined by heterogeneous molecular and micro-environmental histopathologic profiles, which pose a major obstacle in treatment. Correctly diagnosing these tumors and assessing their heterogeneity is crucial for choosing the precise treatment and potentially enhancing patient survival rates. In the gold-standard histopathology-based approach to tumor diagnosis, detecting various morpho-pathological features of distinct histology throughout digitized tissue sections is crucial. Such "features" include the presence of cellular tumor, geographic necrosis, pseudopalisading necrosis, areas abundant in microvascular proliferation, infiltration into the cortex, wide extension in subcortical white matter, leptomeningeal infiltration, regions dense with macrophages, and the presence of perivascular or scattered lymphocytes. With these features in mind and building upon the main aim of the BraTS Cluster of Challenges https://www.synapse.org/brats2024, the goal of the BraTS-Path challenge is to provide a systematically prepared comprehensive dataset and a benchmarking environment to develop and fairly compare deep-learning models capable of identifying tumor sub-regions of distinct histologic profile. These models aim to further our understanding of the disease and assist in the diagnosis and grading of conditions in a consistent manner.
Generative Adversarial Networks based Skin Lesion Segmentation
May 29, 2023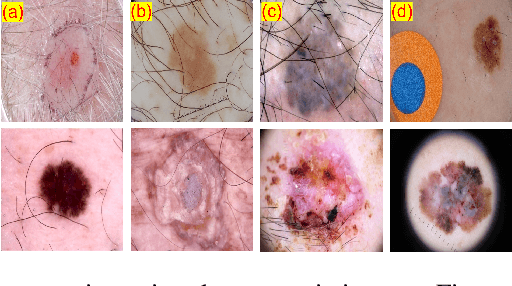
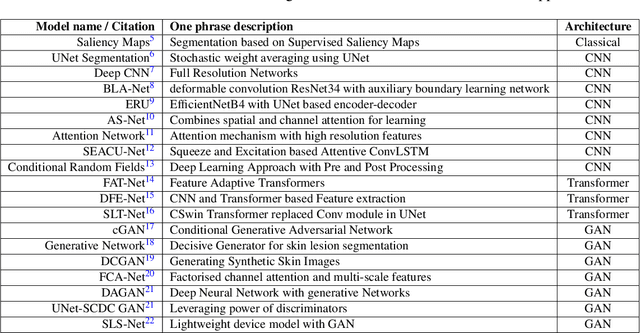
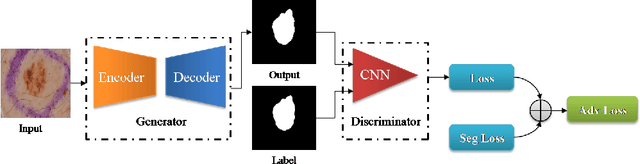
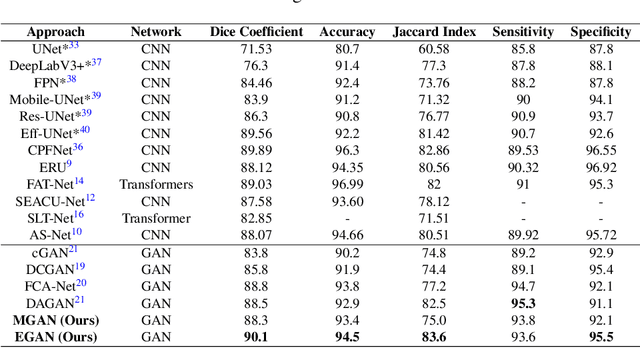
Abstract:Skin cancer is a serious condition that requires accurate identification and treatment. One way to assist clinicians in this task is by using computer-aided diagnosis (CAD) tools that can automatically segment skin lesions from dermoscopic images. To this end, a new adversarial learning-based framework called EGAN has been developed. This framework uses an unsupervised generative network to generate accurate lesion masks. It consists of a generator module with a top-down squeeze excitation-based compound scaled path and an asymmetric lateral connection-based bottom-up path, and a discriminator module that distinguishes between original and synthetic masks. Additionally, a morphology-based smoothing loss is implemented to encourage the network to create smooth semantic boundaries of lesions. The framework is evaluated on the International Skin Imaging Collaboration (ISIC) Lesion Dataset 2018 and outperforms the current state-of-the-art skin lesion segmentation approaches with a Dice coefficient, Jaccard similarity, and Accuracy of 90.1%, 83.6%, and 94.5%, respectively. This represents a 2% increase in Dice Coefficient, 1% increase in Jaccard Index, and 1% increase in Accuracy.
Detecting Histologic Glioblastoma Regions of Prognostic Relevance
Feb 01, 2023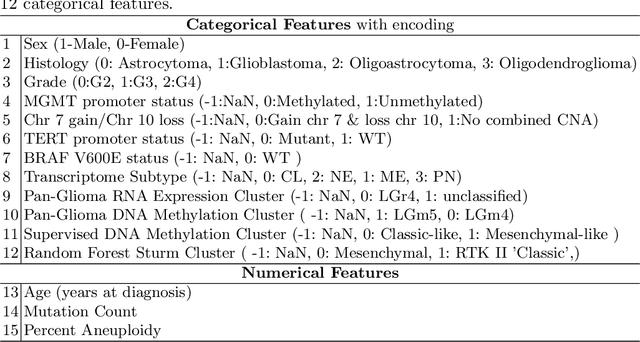
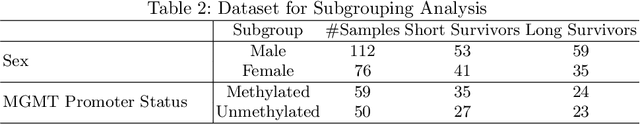
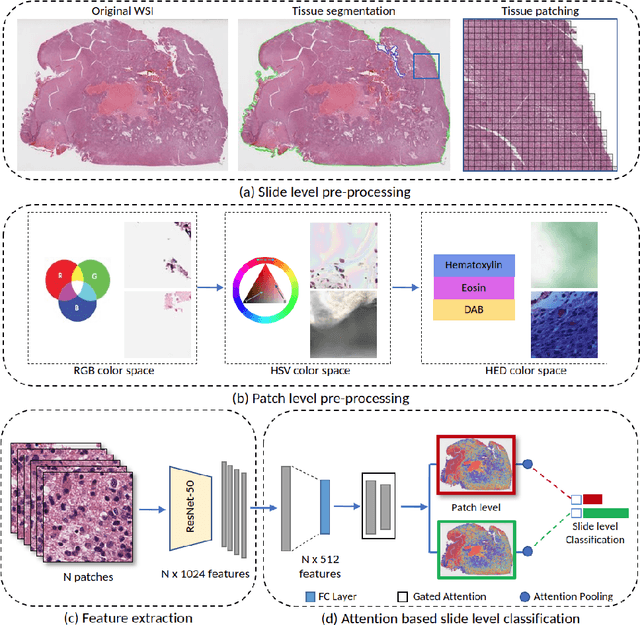

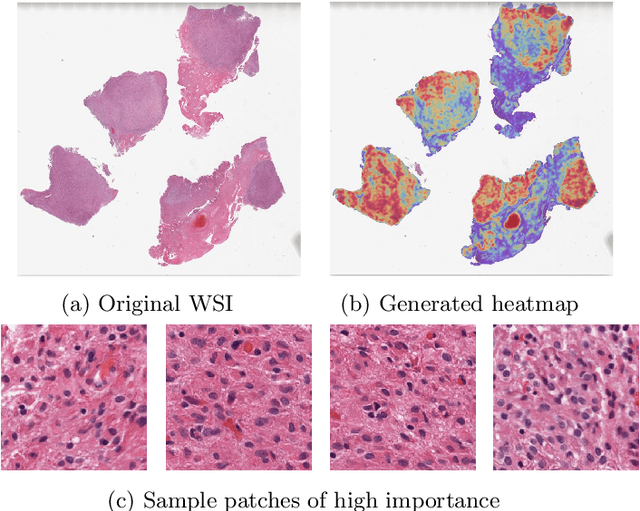
Abstract:Glioblastoma is the most common and aggressive malignant adult tumor of the central nervous system, with grim prognosis and heterogeneous morphologic and molecular profiles. Since the adoption of the current standard of care treatment, 18 years ago, there are no substantial prognostic improvements noticed. Accurate prediction of patient overall survival (OS) from clinical histopathology whole slide images (WSI) using advanced computational methods could contribute to optimization of clinical decision making and patient management. Here, we focus on identifying prognostically relevant glioblastoma morphologic patterns on H&E stained WSI. The exact approach capitalizes on the comprehensive WSI curation of apparent artifactual content and on an interpretability mechanism via a weakly supervised attention based multiple instance learning algorithm that further utilizes clustering to constrain the search space. The automatically identified patterns of high diagnostic value are used to classify the WSI as representative of a short or a long survivor. Identifying tumor morphologic patterns associated with short and long OS will allow the clinical neuropathologist to provide additional prognostic information gleaned during microscopic assessment to the treating team, as well as suggest avenues of biological investigation for understanding and potentially treating glioblastoma.
Deep Learning based Novel Cascaded Approach for Skin Lesion Analysis
Jan 16, 2023



Abstract:Automatic lesion analysis is critical in skin cancer diagnosis and ensures effective treatment. The computer aided diagnosis of such skin cancer in dermoscopic images can significantly reduce the clinicians workload and help improve diagnostic accuracy. Although researchers are working extensively to address this problem, early detection and accurate identification of skin lesions remain challenging. This research focuses on a two step framework for skin lesion segmentation followed by classification for lesion analysis. We explored the effectiveness of deep convolutional neural network based architectures by designing an encoder-decoder architecture for skin lesion segmentation and CNN based classification network. The proposed approaches are evaluated quantitatively in terms of the Accuracy, mean Intersection over Union and Dice Similarity Coefficient. Our cascaded end to end deep learning based approach is the first of its kind, where the classification accuracy of the lesion is significantly improved because of prior segmentation.
REFUGE2 Challenge: Treasure for Multi-Domain Learning in Glaucoma Assessment
Feb 24, 2022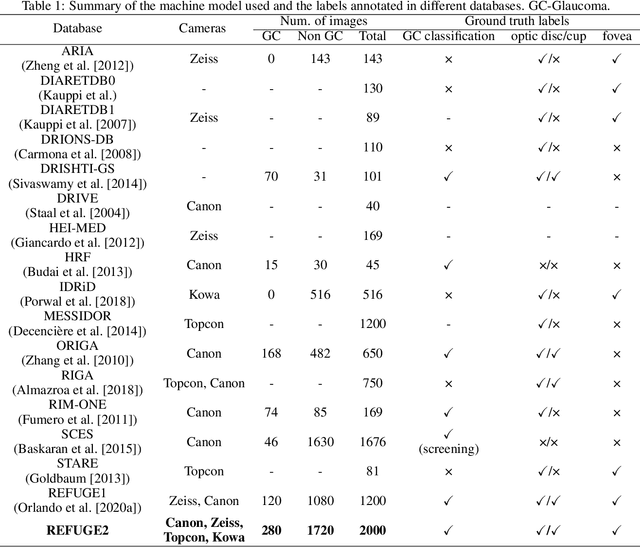
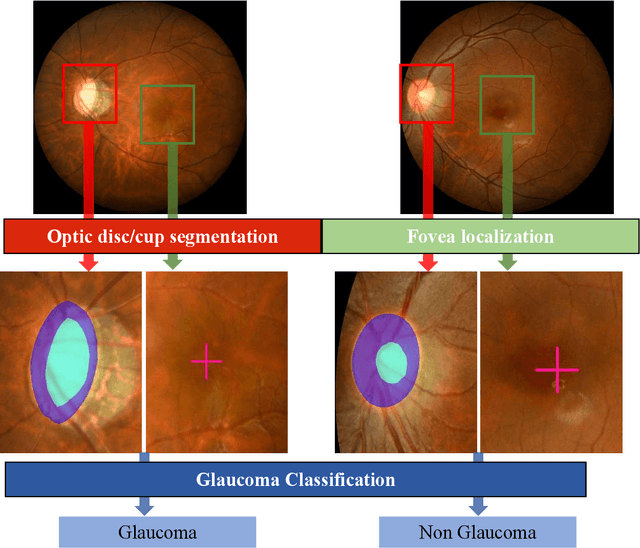
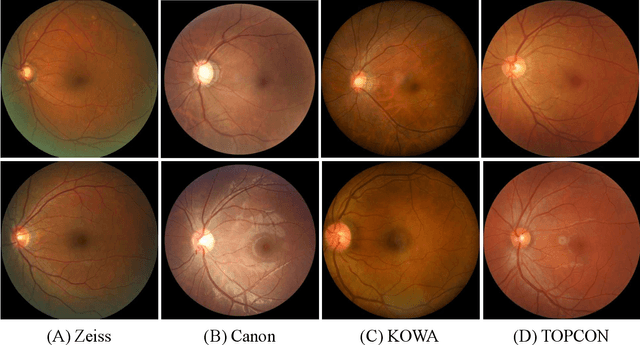
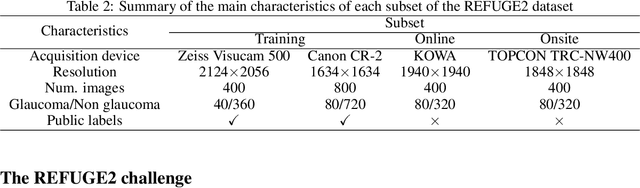
Abstract:Glaucoma is the second leading cause of blindness and is the leading cause of irreversible blindness disease in the world. Early screening for glaucoma in the population is significant. Color fundus photography is the most cost effective imaging modality to screen for ocular diseases. Deep learning network is often used in color fundus image analysis due to its powful feature extraction capability. However, the model training of deep learning method needs a large amount of data, and the distribution of data should be abundant for the robustness of model performance. To promote the research of deep learning in color fundus photography and help researchers further explore the clinical application signification of AI technology, we held a REFUGE2 challenge. This challenge released 2,000 color fundus images of four models, including Zeiss, Canon, Kowa and Topcon, which can validate the stabilization and generalization of algorithms on multi-domain. Moreover, three sub-tasks were designed in the challenge, including glaucoma classification, cup/optic disc segmentation, and macular fovea localization. These sub-tasks technically cover the three main problems of computer vision and clinicly cover the main researchs of glaucoma diagnosis. Over 1,300 international competitors joined the REFUGE2 challenge, 134 teams submitted more than 3,000 valid preliminary results, and 22 teams reached the final. This article summarizes the methods of some of the finalists and analyzes their results. In particular, we observed that the teams using domain adaptation strategies had high and robust performance on the dataset with multi-domain. This indicates that UDA and other multi-domain related researches will be the trend of deep learning field in the future, and our REFUGE2 datasets will play an important role in these researches.
ADAM Challenge: Detecting Age-related Macular Degeneration from Fundus Images
Feb 18, 2022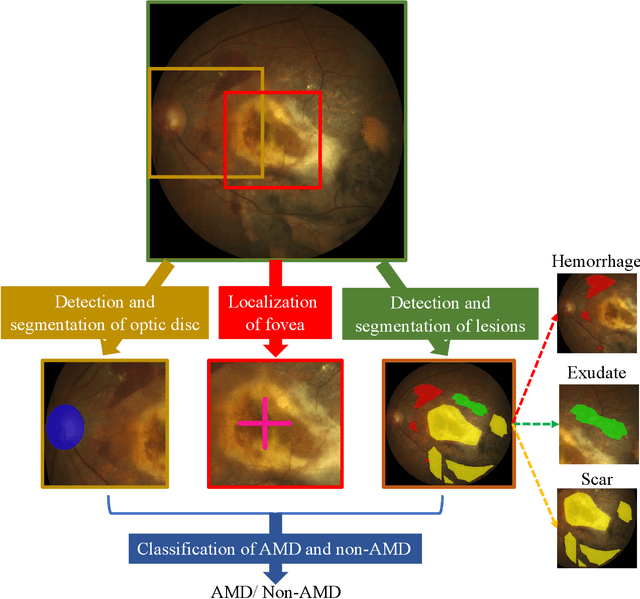
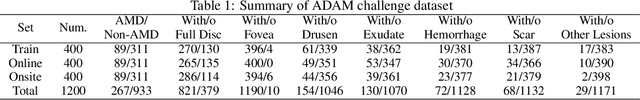
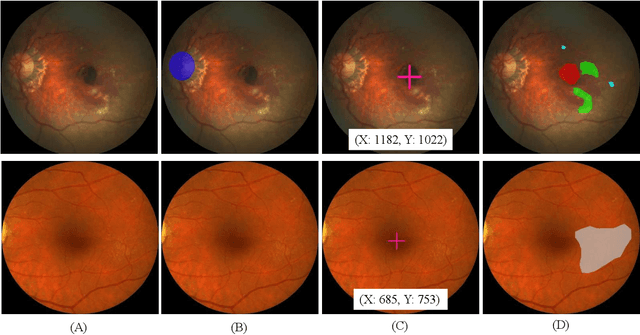
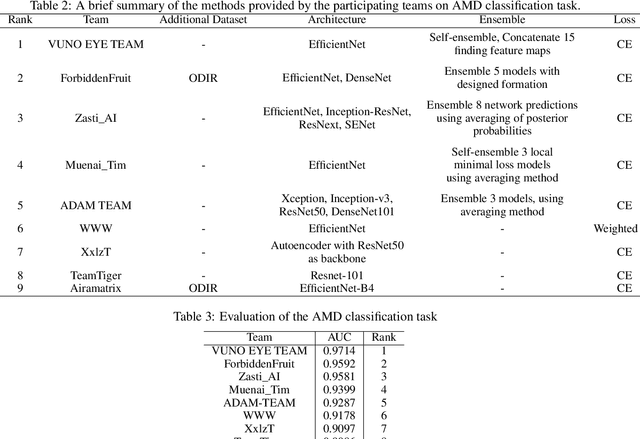
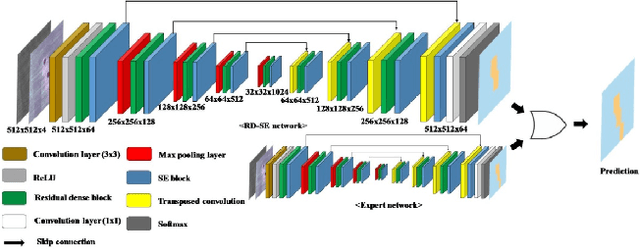
Abstract:Age-related macular degeneration (AMD) is the leading cause of visual impairment among elderly in the world. Early detection of AMD is of great importance as the vision loss caused by AMD is irreversible and permanent. Color fundus photography is the most cost-effective imaging modality to screen for retinal disorders. \textcolor{red}{Recently, some algorithms based on deep learning had been developed for fundus image analysis and automatic AMD detection. However, a comprehensive annotated dataset and a standard evaluation benchmark are still missing.} To deal with this issue, we set up the Automatic Detection challenge on Age-related Macular degeneration (ADAM) for the first time, held as a satellite event of the ISBI 2020 conference. The ADAM challenge consisted of four tasks which cover the main topics in detecting AMD from fundus images, including classification of AMD, detection and segmentation of optic disc, localization of fovea, and detection and segmentation of lesions. The ADAM challenge has released a comprehensive dataset of 1200 fundus images with the category labels of AMD, the pixel-wise segmentation masks of the full optic disc and lesions (drusen, exudate, hemorrhage, scar, and other), as well as the location coordinates of the macular fovea. A uniform evaluation framework has been built to make a fair comparison of different models. During the ADAM challenge, 610 results were submitted for online evaluation, and finally, 11 teams participated in the onsite challenge. This paper introduces the challenge, dataset, and evaluation methods, as well as summarizes the methods and analyzes the results of the participating teams of each task. In particular, we observed that ensembling strategy and clinical prior knowledge can better improve the performances of the deep learning models.
The 1st Agriculture-Vision Challenge: Methods and Results
Apr 23, 2020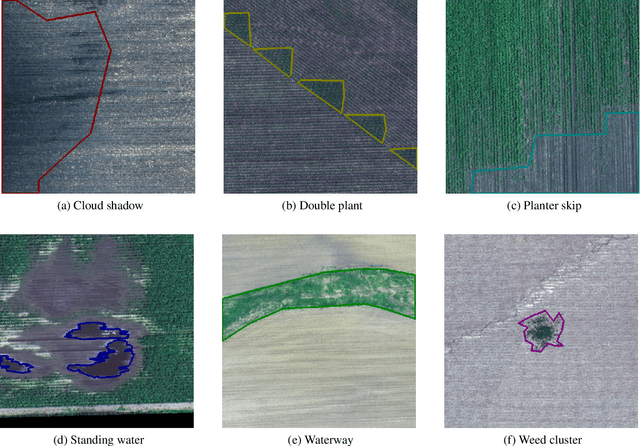
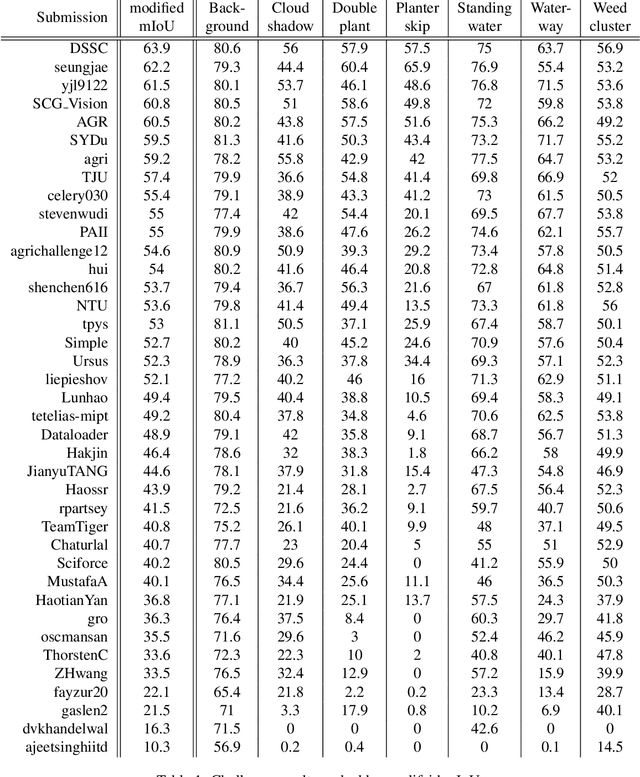
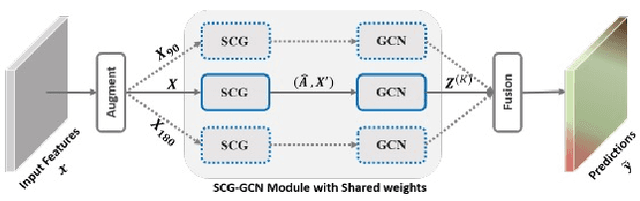
Abstract:The first Agriculture-Vision Challenge aims to encourage research in developing novel and effective algorithms for agricultural pattern recognition from aerial images, especially for the semantic segmentation task associated with our challenge dataset. Around 57 participating teams from various countries compete to achieve state-of-the-art in aerial agriculture semantic segmentation. The Agriculture-Vision Challenge Dataset was employed, which comprises of 21,061 aerial and multi-spectral farmland images. This paper provides a summary of notable methods and results in the challenge. Our submission server and leaderboard will continue to open for researchers that are interested in this challenge dataset and task; the link can be found here.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge