Cheng Bian
Constraint Latent Space Matters: An Anti-anomalous Waveform Transformation Solution from Photoplethysmography to Arterial Blood Pressure
Feb 23, 2024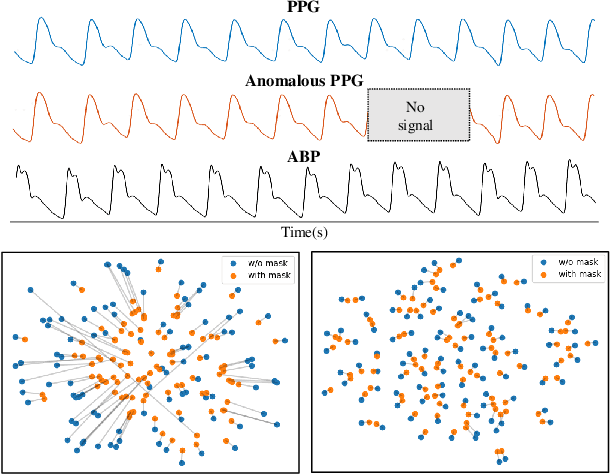
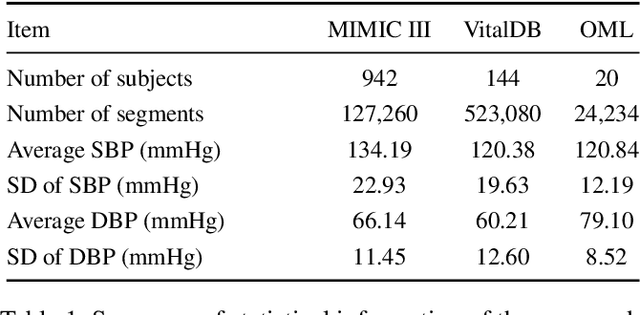
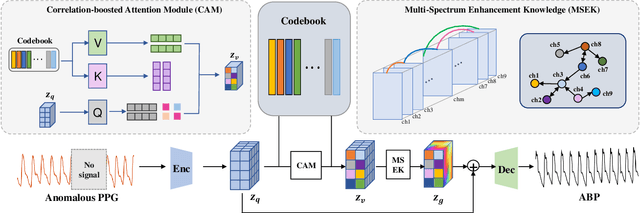
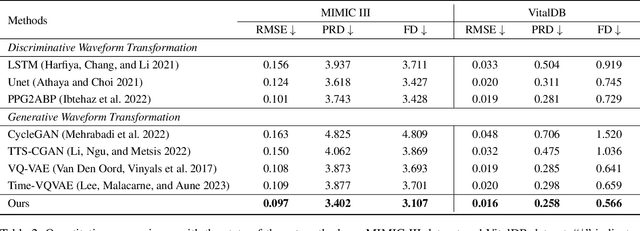
Abstract:Arterial blood pressure (ABP) holds substantial promise for proactive cardiovascular health management. Notwithstanding its potential, the invasive nature of ABP measurements confines their utility primarily to clinical environments, limiting their applicability for continuous monitoring beyond medical facilities. The conversion of photoplethysmography (PPG) signals into ABP equivalents has garnered significant attention due to its potential in revolutionizing cardiovascular disease management. Recent strides in PPG-to-ABP prediction encompass the integration of generative and discriminative models. Despite these advances, the efficacy of these models is curtailed by the latent space shift predicament, stemming from alterations in PPG data distribution across disparate hardware and individuals, potentially leading to distorted ABP waveforms. To tackle this problem, we present an innovative solution named the Latent Space Constraint Transformer (LSCT), leveraging a quantized codebook to yield robust latent spaces by employing multiple discretizing bases. To facilitate improved reconstruction, the Correlation-boosted Attention Module (CAM) is introduced to systematically query pertinent bases on a global scale. Furthermore, to enhance expressive capacity, we propose the Multi-Spectrum Enhancement Knowledge (MSEK), which fosters local information flow within the channels of latent code and provides additional embedding for reconstruction. Through comprehensive experimentation on both publicly available datasets and a private downstream task dataset, the proposed approach demonstrates noteworthy performance enhancements compared to existing methods. Extensive ablation studies further substantiate the effectiveness of each introduced module.
Protein Representation Learning via Knowledge Enhanced Primary Structure Modeling
Feb 15, 2023Abstract:Protein representation learning has primarily benefited from the remarkable development of language models (LMs). Accordingly, pre-trained protein models also suffer from a problem in LMs: a lack of factual knowledge. The recent solution models the relationships between protein and associated knowledge terms as the knowledge encoding objective. However, it fails to explore the relationships at a more granular level, i.e., the token level. To mitigate this, we propose Knowledge-exploited Auto-encoder for Protein (KeAP), which performs token-level knowledge graph exploration for protein representation learning. In practice, non-masked amino acids iteratively query the associated knowledge tokens to extract and integrate helpful information for restoring masked amino acids via attention. We show that KeAP can consistently outperform the previous counterpart on 9 representative downstream applications, sometimes surpassing it by large margins. These results suggest that KeAP provides an alternative yet effective way to perform knowledge enhanced protein representation learning.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Domain Adaptation Meets Zero-Shot Learning: An Annotation-Efficient Approach to Multi-Modality Medical Image Segmentation
Mar 19, 2022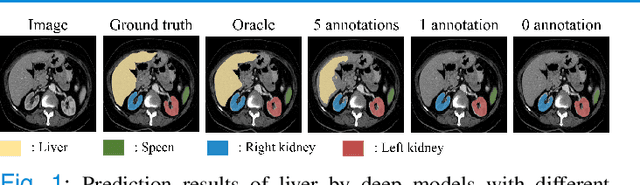
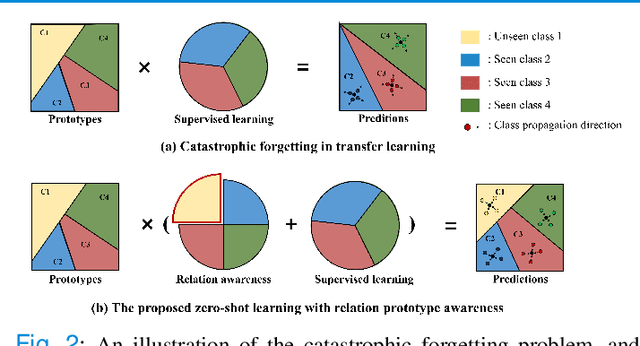
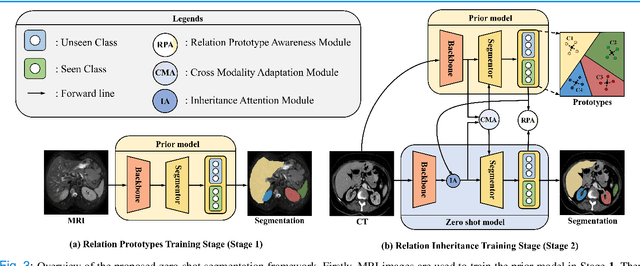
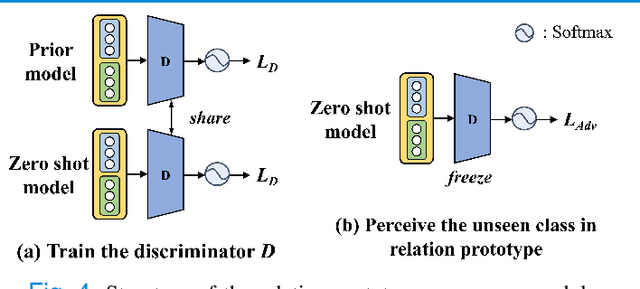
Abstract:Due to the lack of properly annotated medical data, exploring the generalization capability of the deep model is becoming a public concern. Zero-shot learning (ZSL) has emerged in recent years to equip the deep model with the ability to recognize unseen classes. However, existing studies mainly focus on natural images, which utilize linguistic models to extract auxiliary information for ZSL. It is impractical to apply the natural image ZSL solutions directly to medical images, since the medical terminology is very domain-specific, and it is not easy to acquire linguistic models for the medical terminology. In this work, we propose a new paradigm of ZSL specifically for medical images utilizing cross-modality information. We make three main contributions with the proposed paradigm. First, we extract the prior knowledge about the segmentation targets, called relation prototypes, from the prior model and then propose a cross-modality adaptation module to inherit the prototypes to the zero-shot model. Second, we propose a relation prototype awareness module to make the zero-shot model aware of information contained in the prototypes. Last but not least, we develop an inheritance attention module to recalibrate the relation prototypes to enhance the inheritance process. The proposed framework is evaluated on two public cross-modality datasets including a cardiac dataset and an abdominal dataset. Extensive experiments show that the proposed framework significantly outperforms the state of the arts.
Label-efficient Hybrid-supervised Learning for Medical Image Segmentation
Mar 10, 2022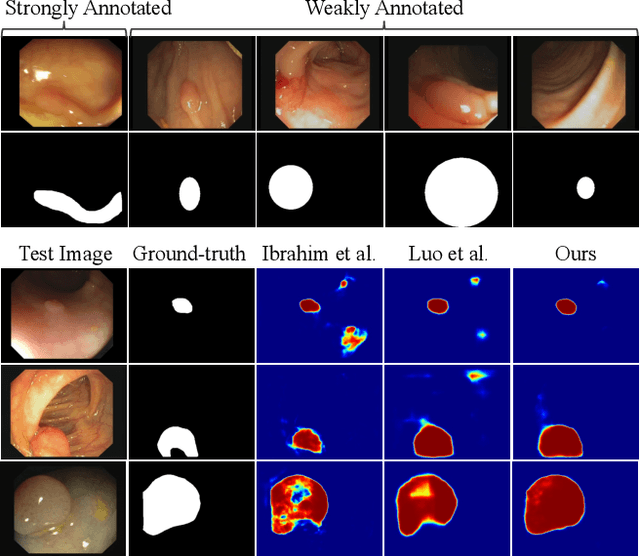
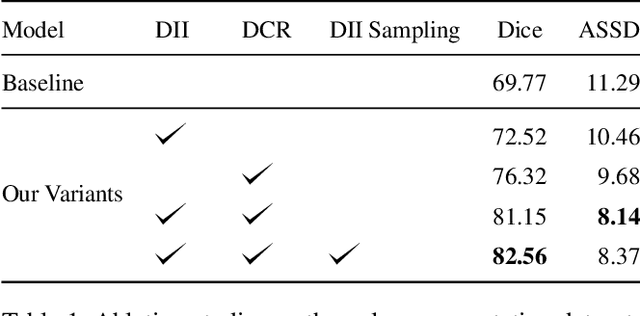

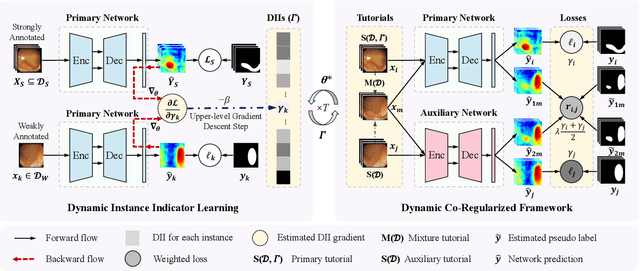
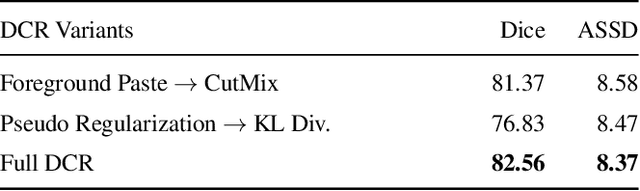
Abstract:Due to the lack of expertise for medical image annotation, the investigation of label-efficient methodology for medical image segmentation becomes a heated topic. Recent progresses focus on the efficient utilization of weak annotations together with few strongly-annotated labels so as to achieve comparable segmentation performance in many unprofessional scenarios. However, these approaches only concentrate on the supervision inconsistency between strongly- and weakly-annotated instances but ignore the instance inconsistency inside the weakly-annotated instances, which inevitably leads to performance degradation. To address this problem, we propose a novel label-efficient hybrid-supervised framework, which considers each weakly-annotated instance individually and learns its weight guided by the gradient direction of the strongly-annotated instances, so that the high-quality prior in the strongly-annotated instances is better exploited and the weakly-annotated instances are depicted more precisely. Specially, our designed dynamic instance indicator (DII) realizes the above objectives, and is adapted to our dynamic co-regularization (DCR) framework further to alleviate the erroneous accumulation from distortions of weak annotations. Extensive experiments on two hybrid-supervised medical segmentation datasets demonstrate that with only 10% strong labels, the proposed framework can leverage the weak labels efficiently and achieve competitive performance against the 100% strong-label supervised scenario.
REFUGE2 Challenge: Treasure for Multi-Domain Learning in Glaucoma Assessment
Feb 24, 2022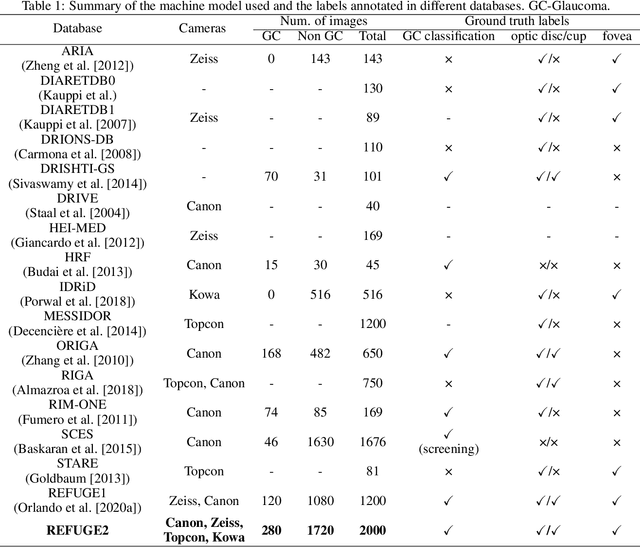
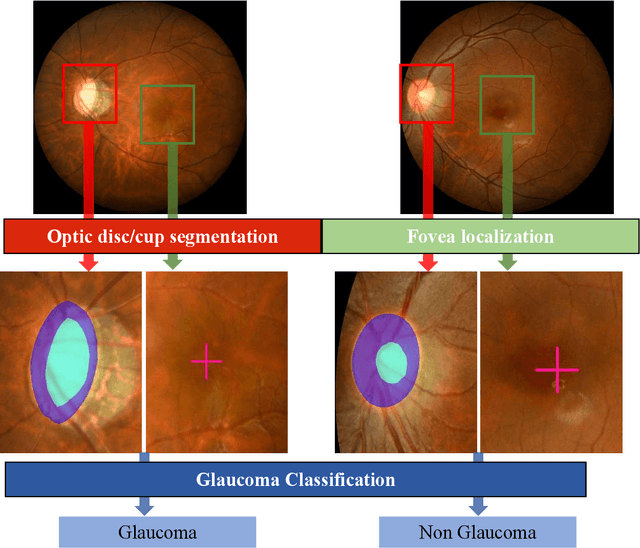
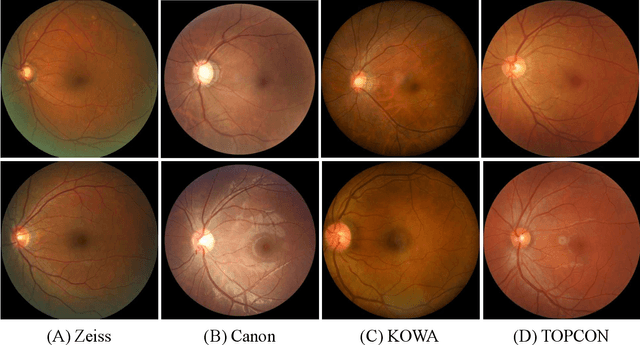
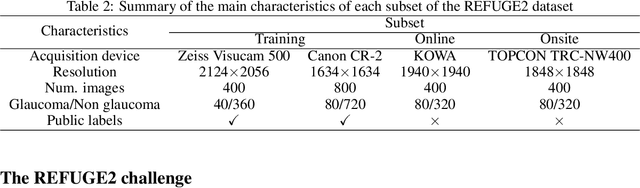
Abstract:Glaucoma is the second leading cause of blindness and is the leading cause of irreversible blindness disease in the world. Early screening for glaucoma in the population is significant. Color fundus photography is the most cost effective imaging modality to screen for ocular diseases. Deep learning network is often used in color fundus image analysis due to its powful feature extraction capability. However, the model training of deep learning method needs a large amount of data, and the distribution of data should be abundant for the robustness of model performance. To promote the research of deep learning in color fundus photography and help researchers further explore the clinical application signification of AI technology, we held a REFUGE2 challenge. This challenge released 2,000 color fundus images of four models, including Zeiss, Canon, Kowa and Topcon, which can validate the stabilization and generalization of algorithms on multi-domain. Moreover, three sub-tasks were designed in the challenge, including glaucoma classification, cup/optic disc segmentation, and macular fovea localization. These sub-tasks technically cover the three main problems of computer vision and clinicly cover the main researchs of glaucoma diagnosis. Over 1,300 international competitors joined the REFUGE2 challenge, 134 teams submitted more than 3,000 valid preliminary results, and 22 teams reached the final. This article summarizes the methods of some of the finalists and analyzes their results. In particular, we observed that the teams using domain adaptation strategies had high and robust performance on the dataset with multi-domain. This indicates that UDA and other multi-domain related researches will be the trend of deep learning field in the future, and our REFUGE2 datasets will play an important role in these researches.
Multi-Anchor Active Domain Adaptation for Semantic Segmentation
Aug 18, 2021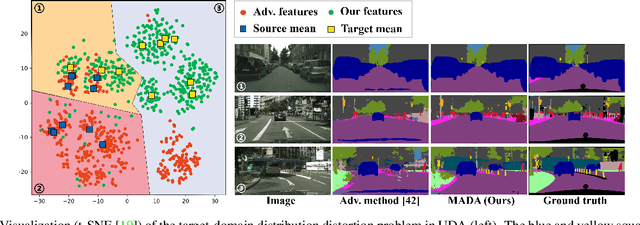
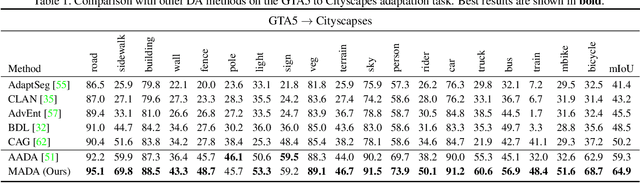
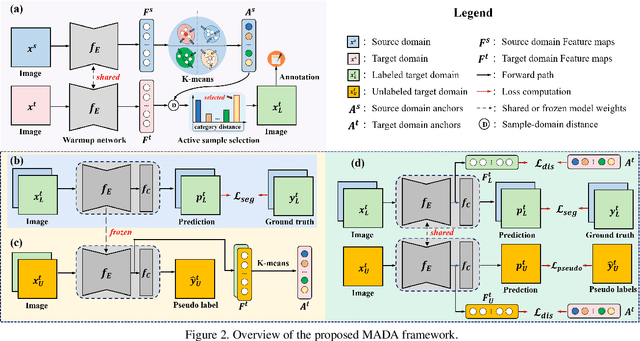
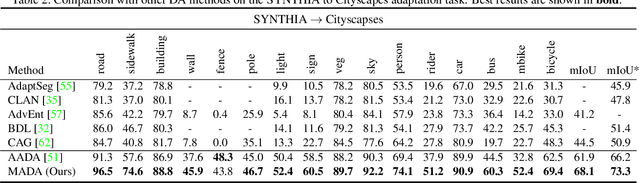
Abstract:Unsupervised domain adaption has proven to be an effective approach for alleviating the intensive workload of manual annotation by aligning the synthetic source-domain data and the real-world target-domain samples. Unfortunately, mapping the target-domain distribution to the source-domain unconditionally may distort the essential structural information of the target-domain data. To this end, we firstly propose to introduce a novel multi-anchor based active learning strategy to assist domain adaptation regarding the semantic segmentation task. By innovatively adopting multiple anchors instead of a single centroid, the source domain can be better characterized as a multimodal distribution, thus more representative and complimentary samples are selected from the target domain. With little workload to manually annotate these active samples, the distortion of the target-domain distribution can be effectively alleviated, resulting in a large performance gain. The multi-anchor strategy is additionally employed to model the target-distribution. By regularizing the latent representation of the target samples compact around multiple anchors through a novel soft alignment loss, more precise segmentation can be achieved. Extensive experiments are conducted on public datasets to demonstrate that the proposed approach outperforms state-of-the-art methods significantly, along with thorough ablation study to verify the effectiveness of each component.
A New Bidirectional Unsupervised Domain Adaptation Segmentation Framework
Aug 18, 2021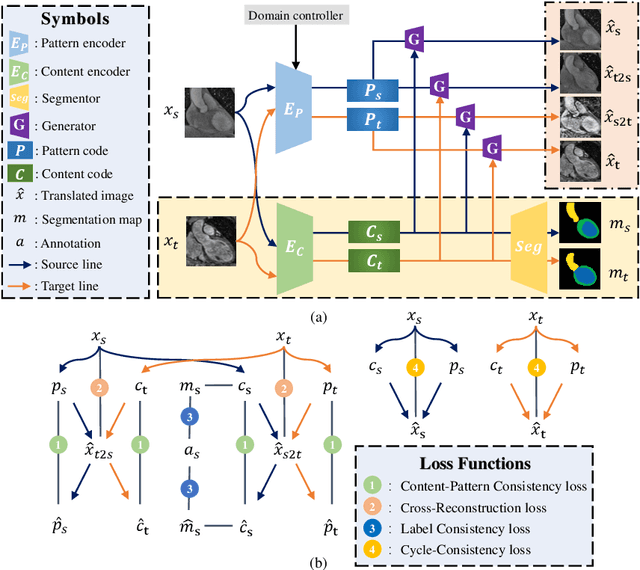
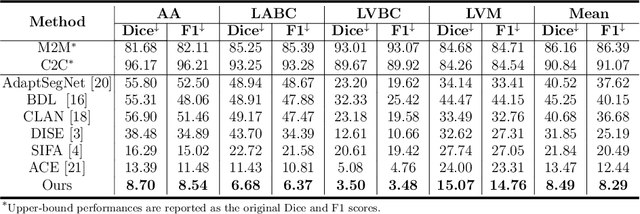
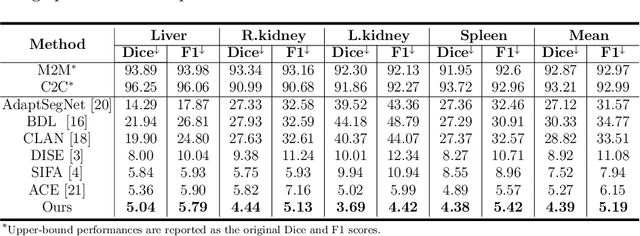
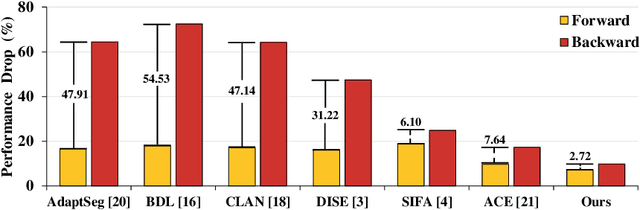
Abstract:Domain shift happens in cross-domain scenarios commonly because of the wide gaps between different domains: when applying a deep learning model well-trained in one domain to another target domain, the model usually performs poorly. To tackle this problem, unsupervised domain adaptation (UDA) techniques are proposed to bridge the gap between different domains, for the purpose of improving model performance without annotation in the target domain. Particularly, UDA has a great value for multimodal medical image analysis, where annotation difficulty is a practical concern. However, most existing UDA methods can only achieve satisfactory improvements in one adaptation direction (e.g., MRI to CT), but often perform poorly in the other (CT to MRI), limiting their practical usage. In this paper, we propose a bidirectional UDA (BiUDA) framework based on disentangled representation learning for equally competent two-way UDA performances. This framework employs a unified domain-aware pattern encoder which not only can adaptively encode images in different domains through a domain controller, but also improve model efficiency by eliminating redundant parameters. Furthermore, to avoid distortion of contents and patterns of input images during the adaptation process, a content-pattern consistency loss is introduced. Additionally, for better UDA segmentation performance, a label consistency strategy is proposed to provide extra supervision by recomposing target-domain-styled images and corresponding source-domain annotations. Comparison experiments and ablation studies conducted on two public datasets demonstrate the superiority of our BiUDA framework to current state-of-the-art UDA methods and the effectiveness of its novel designs. By successfully addressing two-way adaptations, our BiUDA framework offers a flexible solution of UDA techniques to the real-world scenario.
Ensembled ResUnet for Anatomical Brain Barriers Segmentation
Jan 04, 2021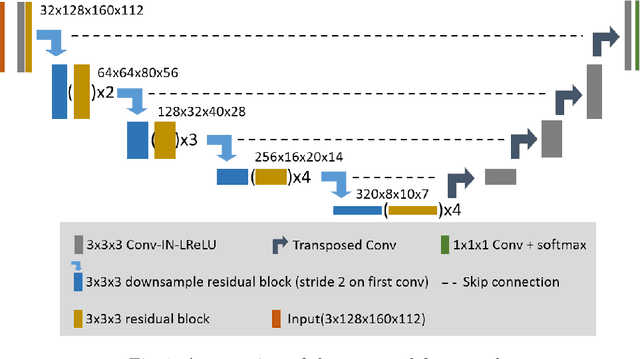
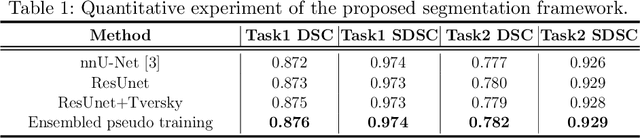
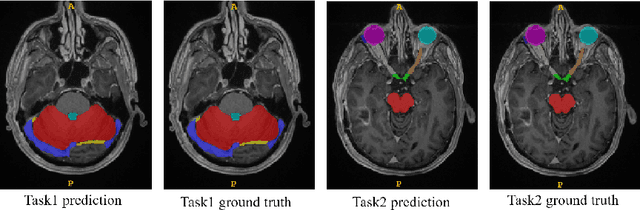
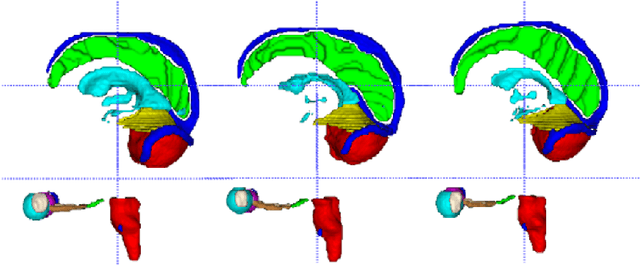
Abstract:Accuracy segmentation of brain structures could be helpful for glioma and radiotherapy planning. However, due to the visual and anatomical differences between different modalities, the accurate segmentation of brain structures becomes challenging. To address this problem, we first construct a residual block based U-shape network with a deep encoder and shallow decoder, which can trade off the framework performance and efficiency. Then, we introduce the Tversky loss to address the issue of the class imbalance between different foreground and the background classes. Finally, a model ensemble strategy is utilized to remove outliers and further boost performance.
TR-GAN: Topology Ranking GAN with Triplet Loss for Retinal Artery/Vein Classification
Jul 29, 2020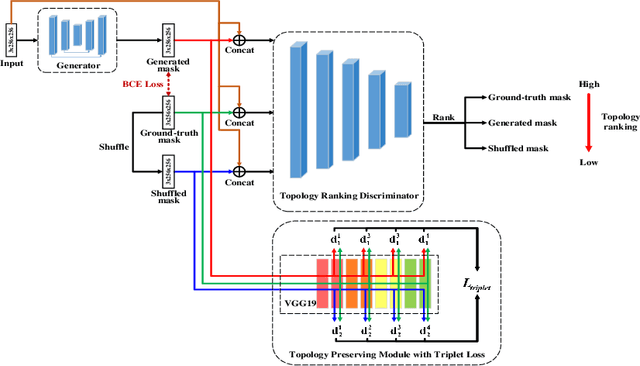
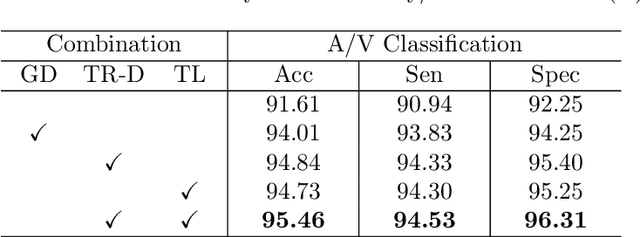
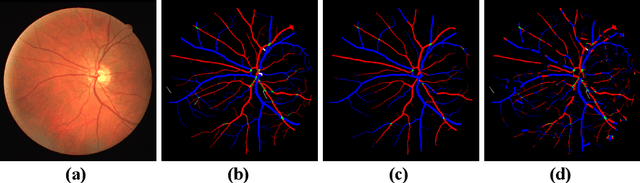
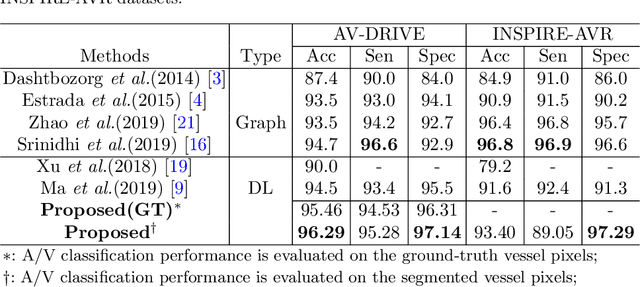
Abstract:Retinal artery/vein (A/V) classification lays the foundation for the quantitative analysis of retinal vessels, which is associated with potential risks of various cardiovascular and cerebral diseases. The topological connection relationship, which has been proved effective in improving the A/V classification performance for the conventional graph based method, has not been exploited by the deep learning based method. In this paper, we propose a Topology Ranking Generative Adversarial Network (TR-GAN) to improve the topology connectivity of the segmented arteries and veins, and further to boost the A/V classification performance. A topology ranking discriminator based on ordinal regression is proposed to rank the topological connectivity level of the ground-truth, the generated A/V mask and the intentionally shuffled mask. The ranking loss is further back-propagated to the generator to generate better connected A/V masks. In addition, a topology preserving module with triplet loss is also proposed to extract the high-level topological features and further to narrow the feature distance between the predicted A/V mask and the ground-truth. The proposed framework effectively increases the topological connectivity of the predicted A/V masks and achieves state-of-the-art A/V classification performance on the publicly available AV-DRIVE dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge