Patrick Godau
German Cancer Research Center, National Center for Tumor Diseases
Performance uncertainty in medical image analysis: a large-scale investigation of confidence intervals
Jan 23, 2026Abstract:Performance uncertainty quantification is essential for reliable validation and eventual clinical translation of medical imaging artificial intelligence (AI). Confidence intervals (CIs) play a central role in this process by indicating how precise a reported performance estimate is. Yet, due to the limited amount of work examining CI behavior in medical imaging, the community remains largely unaware of how many diverse CI methods exist and how they behave in specific settings. The purpose of this study is to close this gap. To this end, we conducted a large-scale empirical analysis across a total of 24 segmentation and classification tasks, using 19 trained models per task group, a broad spectrum of commonly used performance metrics, multiple aggregation strategies, and several widely adopted CI methods. Reliability (coverage) and precision (width) of each CI method were estimated across all settings to characterize their dependence on study characteristics. Our analysis revealed five principal findings: 1) the sample size required for reliable CIs varies from a few dozens to several thousands of cases depending on study parameters; 2) CI behavior is strongly affected by the choice of performance metric; 3) aggregation strategy substantially influences the reliability of CIs, e.g. they require more observations for macro than for micro; 4) the machine learning problem (segmentation versus classification) modulates these effects; 5) different CI methods are not equally reliable and precise depending on the use case. These results form key components for the development of future guidelines on reporting performance uncertainty in medical imaging AI.
Challenging Vision-Language Models with Surgical Data: A New Dataset and Broad Benchmarking Study
Jun 06, 2025Abstract:While traditional computer vision models have historically struggled to generalize to endoscopic domains, the emergence of foundation models has shown promising cross-domain performance. In this work, we present the first large-scale study assessing the capabilities of Vision Language Models (VLMs) for endoscopic tasks with a specific focus on laparoscopic surgery. Using a diverse set of state-of-the-art models, multiple surgical datasets, and extensive human reference annotations, we address three key research questions: (1) Can current VLMs solve basic perception tasks on surgical images? (2) Can they handle advanced frame-based endoscopic scene understanding tasks? and (3) How do specialized medical VLMs compare to generalist models in this context? Our results reveal that VLMs can effectively perform basic surgical perception tasks, such as object counting and localization, with performance levels comparable to general domain tasks. However, their performance deteriorates significantly when the tasks require medical knowledge. Notably, we find that specialized medical VLMs currently underperform compared to generalist models across both basic and advanced surgical tasks, suggesting that they are not yet optimized for the complexity of surgical environments. These findings highlight the need for further advancements to enable VLMs to handle the unique challenges posed by surgery. Overall, our work provides important insights for the development of next-generation endoscopic AI systems and identifies key areas for improvement in medical visual language models.
False Promises in Medical Imaging AI? Assessing Validity of Outperformance Claims
May 07, 2025Abstract:Performance comparisons are fundamental in medical imaging Artificial Intelligence (AI) research, often driving claims of superiority based on relative improvements in common performance metrics. However, such claims frequently rely solely on empirical mean performance. In this paper, we investigate whether newly proposed methods genuinely outperform the state of the art by analyzing a representative cohort of medical imaging papers. We quantify the probability of false claims based on a Bayesian approach that leverages reported results alongside empirically estimated model congruence to estimate whether the relative ranking of methods is likely to have occurred by chance. According to our results, the majority (>80%) of papers claims outperformance when introducing a new method. Our analysis further revealed a high probability (>5%) of false outperformance claims in 86% of classification papers and 53% of segmentation papers. These findings highlight a critical flaw in current benchmarking practices: claims of outperformance in medical imaging AI are frequently unsubstantiated, posing a risk of misdirecting future research efforts.
Beyond Knowledge Silos: Task Fingerprinting for Democratization of Medical Imaging AI
Dec 11, 2024Abstract:The field of medical imaging AI is currently undergoing rapid transformations, with methodical research increasingly translated into clinical practice. Despite these successes, research suffers from knowledge silos, hindering collaboration and progress: Existing knowledge is scattered across publications and many details remain unpublished, while privacy regulations restrict data sharing. In the spirit of democratizing of AI, we propose a framework for secure knowledge transfer in the field of medical image analysis. The key to our approach is dataset "fingerprints", structured representations of feature distributions, that enable quantification of task similarity. We tested our approach across 71 distinct tasks and 12 medical imaging modalities by transferring neural architectures, pretraining, augmentation policies, and multi-task learning. According to comprehensive analyses, our method outperforms traditional methods for identifying relevant knowledge and facilitates collaborative model training. Our framework fosters the democratization of AI in medical imaging and could become a valuable tool for promoting faster scientific advancement.
PitVis-2023 Challenge: Workflow Recognition in videos of Endoscopic Pituitary Surgery
Sep 02, 2024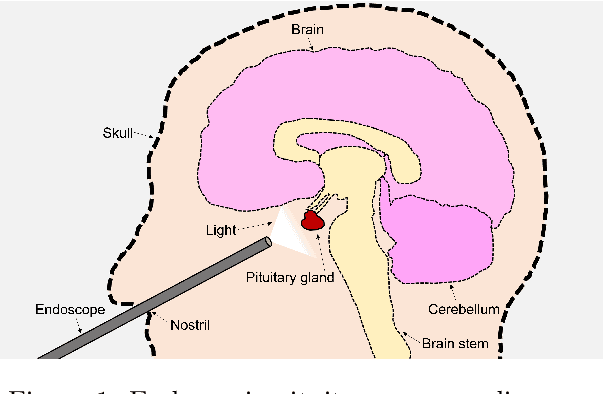
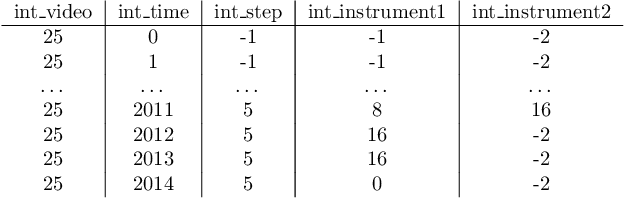

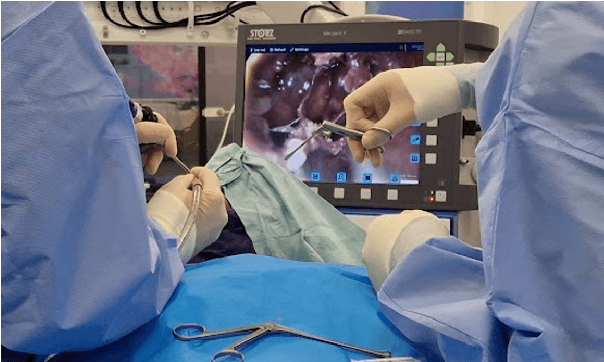
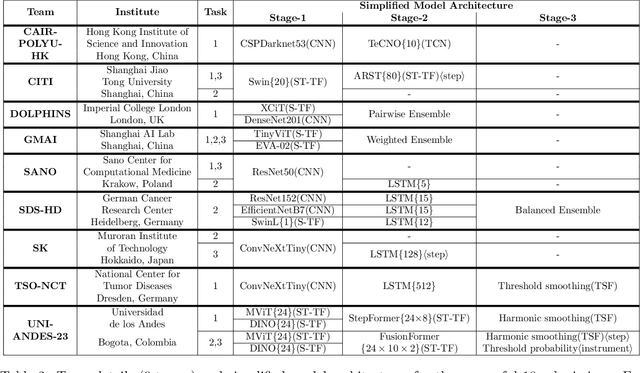
Abstract:The field of computer vision applied to videos of minimally invasive surgery is ever-growing. Workflow recognition pertains to the automated recognition of various aspects of a surgery: including which surgical steps are performed; and which surgical instruments are used. This information can later be used to assist clinicians when learning the surgery; during live surgery; and when writing operation notes. The Pituitary Vision (PitVis) 2023 Challenge tasks the community to step and instrument recognition in videos of endoscopic pituitary surgery. This is a unique task when compared to other minimally invasive surgeries due to the smaller working space, which limits and distorts vision; and higher frequency of instrument and step switching, which requires more precise model predictions. Participants were provided with 25-videos, with results presented at the MICCAI-2023 conference as part of the Endoscopic Vision 2023 Challenge in Vancouver, Canada, on 08-Oct-2023. There were 18-submissions from 9-teams across 6-countries, using a variety of deep learning models. A commonality between the top performing models was incorporating spatio-temporal and multi-task methods, with greater than 50% and 10% macro-F1-score improvement over purely spacial single-task models in step and instrument recognition respectively. The PitVis-2023 Challenge therefore demonstrates state-of-the-art computer vision models in minimally invasive surgery are transferable to a new dataset, with surgery specific techniques used to enhance performance, progressing the field further. Benchmark results are provided in the paper, and the dataset is publicly available at: https://doi.org/10.5522/04/26531686.
Why is the winner the best?
Mar 30, 2023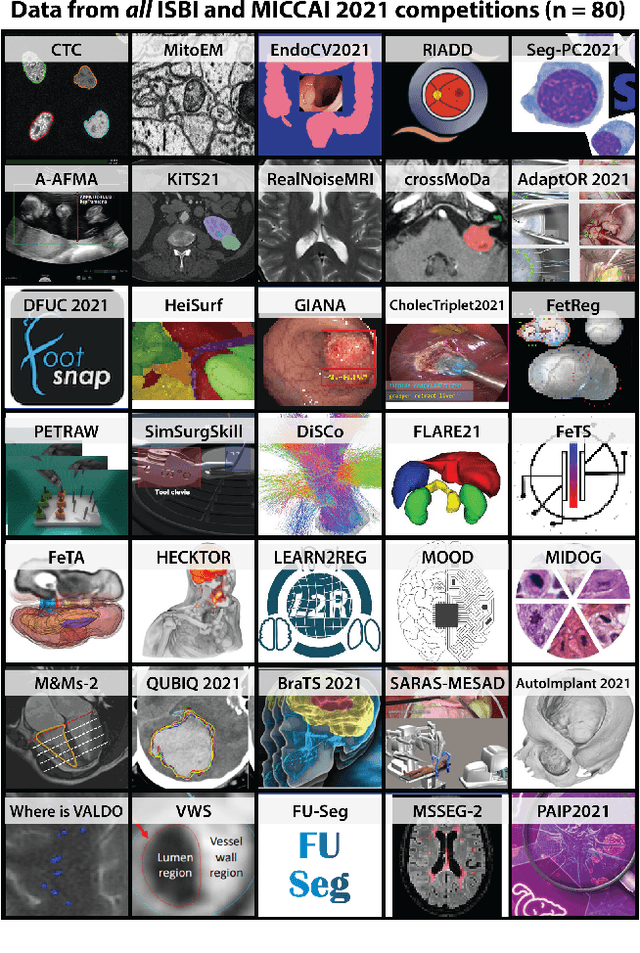
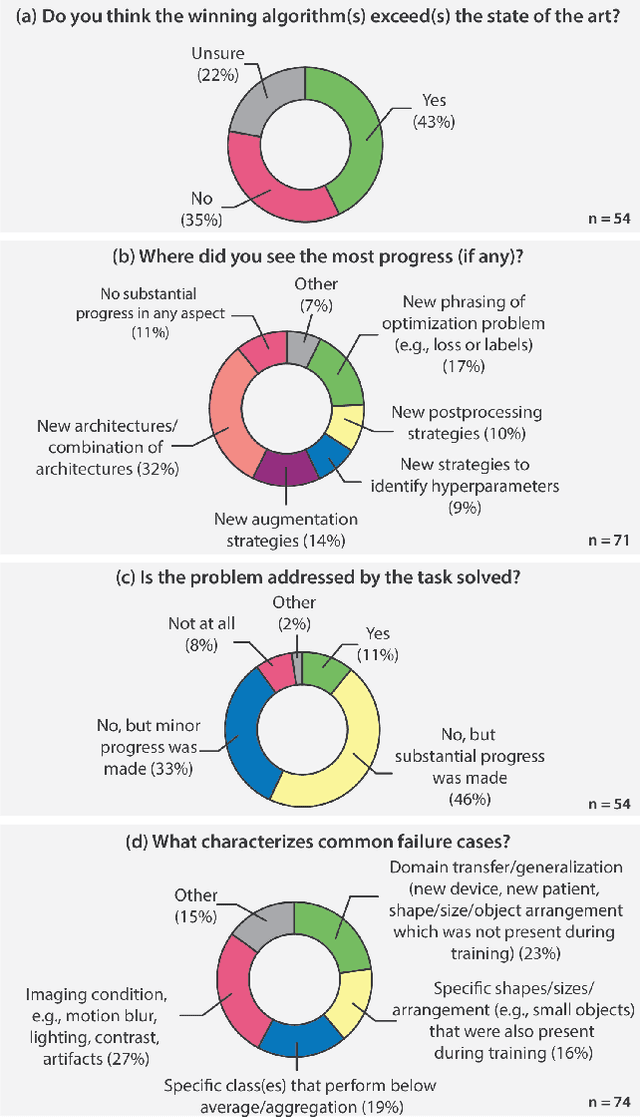
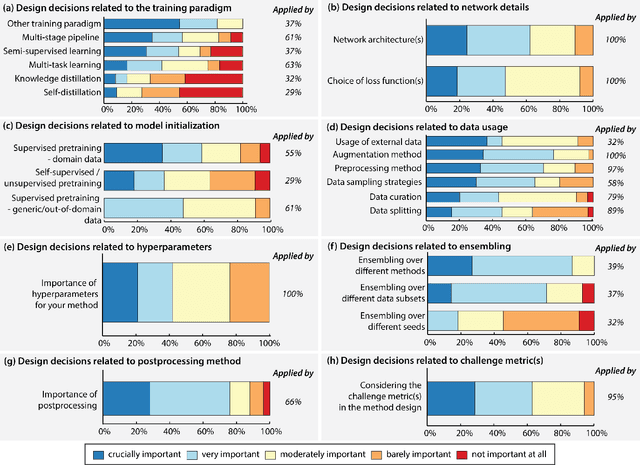
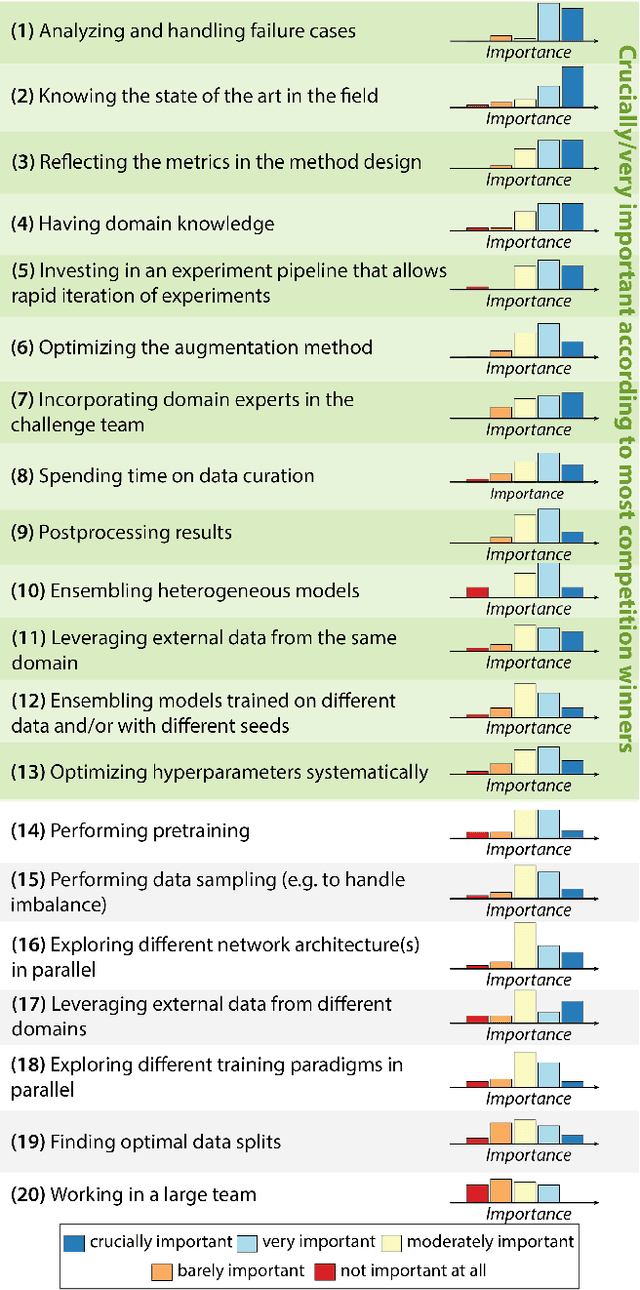
Abstract:International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
Deployment of Image Analysis Algorithms under Prevalence Shifts
Mar 22, 2023



Abstract:Domain gaps are among the most relevant roadblocks in the clinical translation of machine learning (ML)-based solutions for medical image analysis. While current research focuses on new training paradigms and network architectures, little attention is given to the specific effect of prevalence shifts on an algorithm deployed in practice. Such discrepancies between class frequencies in the data used for a method's development/validation and that in its deployment environment(s) are of great importance, for example in the context of artificial intelligence (AI) democratization, as disease prevalences may vary widely across time and location. Our contribution is twofold. First, we empirically demonstrate the potentially severe consequences of missing prevalence handling by analyzing (i) the extent of miscalibration, (ii) the deviation of the decision threshold from the optimum, and (iii) the ability of validation metrics to reflect neural network performance on the deployment population as a function of the discrepancy between development and deployment prevalence. Second, we propose a workflow for prevalence-aware image classification that uses estimated deployment prevalences to adjust a trained classifier to a new environment, without requiring additional annotated deployment data. Comprehensive experiments based on a diverse set of 30 medical classification tasks showcase the benefit of the proposed workflow in generating better classifier decisions and more reliable performance estimates compared to current practice.
Self-distillation for surgical action recognition
Mar 22, 2023



Abstract:Surgical scene understanding is a key prerequisite for contextaware decision support in the operating room. While deep learning-based approaches have already reached or even surpassed human performance in various fields, the task of surgical action recognition remains a major challenge. With this contribution, we are the first to investigate the concept of self-distillation as a means of addressing class imbalance and potential label ambiguity in surgical video analysis. Our proposed method is a heterogeneous ensemble of three models that use Swin Transfomers as backbone and the concepts of self-distillation and multi-task learning as core design choices. According to ablation studies performed with the CholecT45 challenge data via cross-validation, the biggest performance boost is achieved by the usage of soft labels obtained by self-distillation. External validation of our method on an independent test set was achieved by providing a Docker container of our inference model to the challenge organizers. According to their analysis, our method outperforms all other solutions submitted to the latest challenge in the field. Our approach thus shows the potential of self-distillation for becoming an important tool in medical image analysis applications.
CholecTriplet2022: Show me a tool and tell me the triplet -- an endoscopic vision challenge for surgical action triplet detection
Feb 13, 2023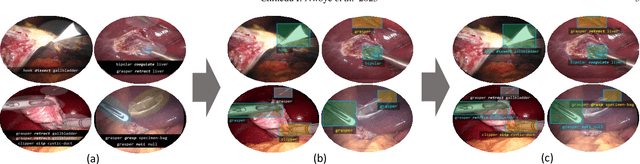
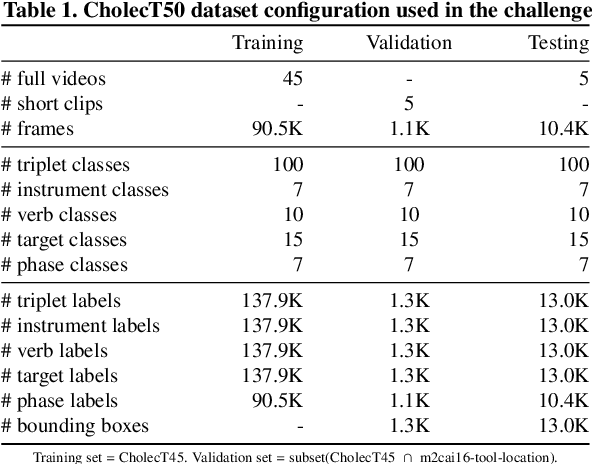
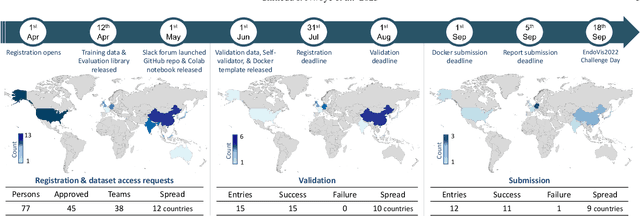
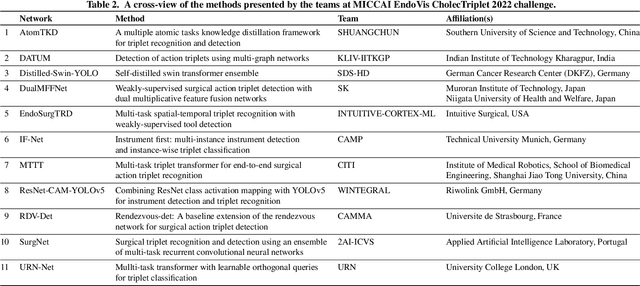
Abstract:Formalizing surgical activities as triplets of the used instruments, actions performed, and target anatomies is becoming a gold standard approach for surgical activity modeling. The benefit is that this formalization helps to obtain a more detailed understanding of tool-tissue interaction which can be used to develop better Artificial Intelligence assistance for image-guided surgery. Earlier efforts and the CholecTriplet challenge introduced in 2021 have put together techniques aimed at recognizing these triplets from surgical footage. Estimating also the spatial locations of the triplets would offer a more precise intraoperative context-aware decision support for computer-assisted intervention. This paper presents the CholecTriplet2022 challenge, which extends surgical action triplet modeling from recognition to detection. It includes weakly-supervised bounding box localization of every visible surgical instrument (or tool), as the key actors, and the modeling of each tool-activity in the form of <instrument, verb, target> triplet. The paper describes a baseline method and 10 new deep learning algorithms presented at the challenge to solve the task. It also provides thorough methodological comparisons of the methods, an in-depth analysis of the obtained results, their significance, and useful insights for future research directions and applications in surgery.
Understanding metric-related pitfalls in image analysis validation
Feb 09, 2023Abstract:Validation metrics are key for the reliable tracking of scientific progress and for bridging the current chasm between artificial intelligence (AI) research and its translation into practice. However, increasing evidence shows that particularly in image analysis, metrics are often chosen inadequately in relation to the underlying research problem. This could be attributed to a lack of accessibility of metric-related knowledge: While taking into account the individual strengths, weaknesses, and limitations of validation metrics is a critical prerequisite to making educated choices, the relevant knowledge is currently scattered and poorly accessible to individual researchers. Based on a multi-stage Delphi process conducted by a multidisciplinary expert consortium as well as extensive community feedback, the present work provides the first reliable and comprehensive common point of access to information on pitfalls related to validation metrics in image analysis. Focusing on biomedical image analysis but with the potential of transfer to other fields, the addressed pitfalls generalize across application domains and are categorized according to a newly created, domain-agnostic taxonomy. To facilitate comprehension, illustrations and specific examples accompany each pitfall. As a structured body of information accessible to researchers of all levels of expertise, this work enhances global comprehension of a key topic in image analysis validation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge