Minu Tizabi
Computer Assisted Medical Interventions
Deployment of Image Analysis Algorithms under Prevalence Shifts
Mar 22, 2023



Abstract:Domain gaps are among the most relevant roadblocks in the clinical translation of machine learning (ML)-based solutions for medical image analysis. While current research focuses on new training paradigms and network architectures, little attention is given to the specific effect of prevalence shifts on an algorithm deployed in practice. Such discrepancies between class frequencies in the data used for a method's development/validation and that in its deployment environment(s) are of great importance, for example in the context of artificial intelligence (AI) democratization, as disease prevalences may vary widely across time and location. Our contribution is twofold. First, we empirically demonstrate the potentially severe consequences of missing prevalence handling by analyzing (i) the extent of miscalibration, (ii) the deviation of the decision threshold from the optimum, and (iii) the ability of validation metrics to reflect neural network performance on the deployment population as a function of the discrepancy between development and deployment prevalence. Second, we propose a workflow for prevalence-aware image classification that uses estimated deployment prevalences to adjust a trained classifier to a new environment, without requiring additional annotated deployment data. Comprehensive experiments based on a diverse set of 30 medical classification tasks showcase the benefit of the proposed workflow in generating better classifier decisions and more reliable performance estimates compared to current practice.
Robust deep learning-based semantic organ segmentation in hyperspectral images
Nov 09, 2021Abstract:Semantic image segmentation is an important prerequisite for context-awareness and autonomous robotics in surgery. The state of the art has focused on conventional RGB video data acquired during minimally invasive surgery, but full-scene semantic segmentation based on spectral imaging data and obtained during open surgery has received almost no attention to date. To address this gap in the literature, we are investigating the following research questions based on hyperspectral imaging (HSI) data of pigs acquired in an open surgery setting: (1) What is an adequate representation of HSI data for neural network-based fully automated organ segmentation, especially with respect to the spatial granularity of the data (pixels vs. superpixels vs. patches vs. full images)? (2) Is there a benefit of using HSI data compared to other modalities, namely RGB data and processed HSI data (e.g. tissue parameters like oxygenation), when performing semantic organ segmentation? According to a comprehensive validation study based on 506 HSI images from 20 pigs, annotated with a total of 19 classes, deep learning-based segmentation performance increases - consistently across modalities - with the spatial context of the input data. Unprocessed HSI data offers an advantage over RGB data or processed data from the camera provider, with the advantage increasing with decreasing size of the input to the neural network. Maximum performance (HSI applied to whole images) yielded a mean dice similarity coefficient (DSC) of 0.89 (standard deviation (SD) 0.04), which is in the range of the inter-rater variability (DSC of 0.89 (SD 0.07)). We conclude that HSI could become a powerful image modality for fully-automatic surgical scene understanding with many advantages over traditional imaging, including the ability to recover additional functional tissue information.
Machine learning-based analysis of hyperspectral images for automated sepsis diagnosis
Jun 15, 2021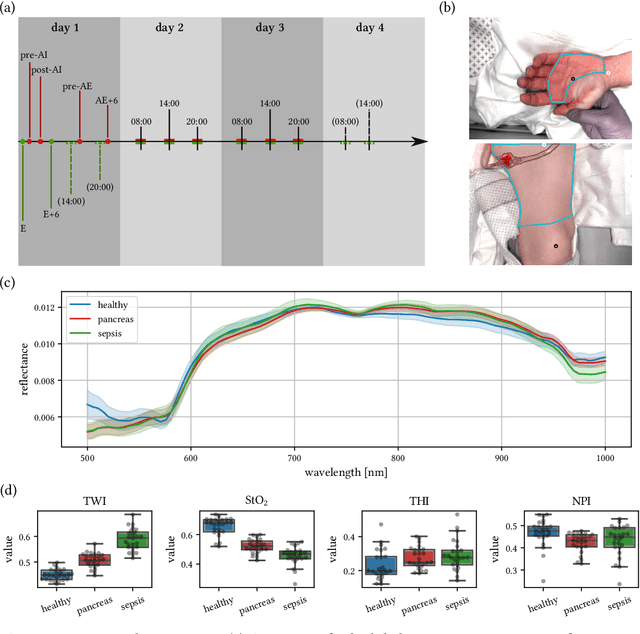
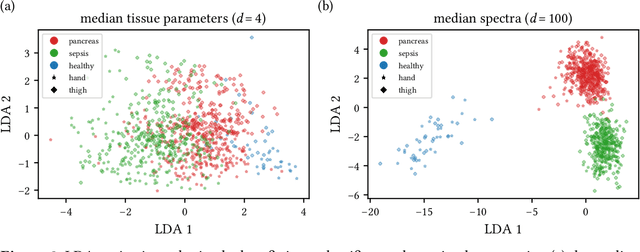
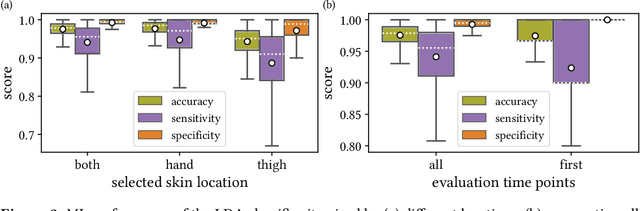
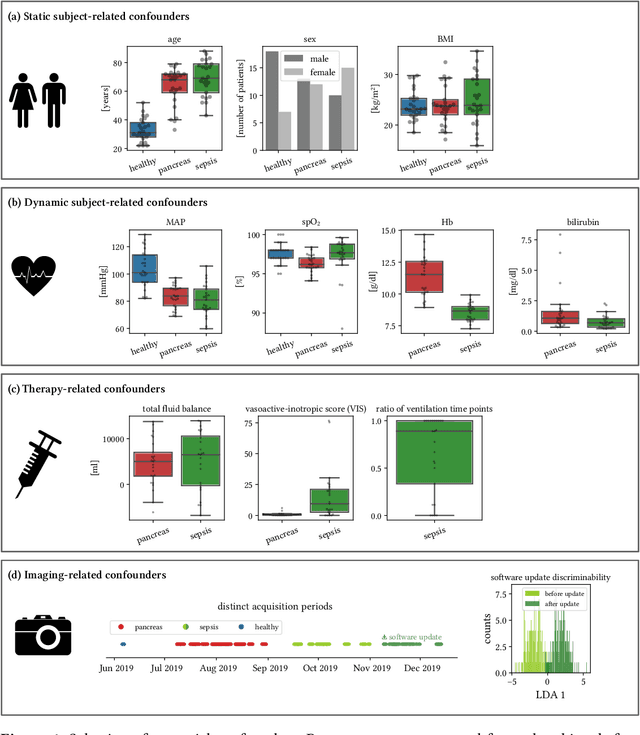
Abstract:Sepsis is a leading cause of mortality and critical illness worldwide. While robust biomarkers for early diagnosis are still missing, recent work indicates that hyperspectral imaging (HSI) has the potential to overcome this bottleneck by monitoring microcirculatory alterations. Automated machine learning-based diagnosis of sepsis based on HSI data, however, has not been explored to date. Given this gap in the literature, we leveraged an existing data set to (1) investigate whether HSI-based automated diagnosis of sepsis is possible and (2) put forth a list of possible confounders relevant for HSI-based tissue classification. While we were able to classify sepsis with an accuracy of over $98\,\%$ using the existing data, our research also revealed several subject-, therapy- and imaging-related confounders that may lead to an overestimation of algorithm performance when not balanced across the patient groups. We conclude that further prospective studies, carefully designed with respect to these confounders, are necessary to confirm the preliminary results obtained in this study.
Heidelberg Colorectal Data Set for Surgical Data Science in the Sensor Operating Room
May 28, 2020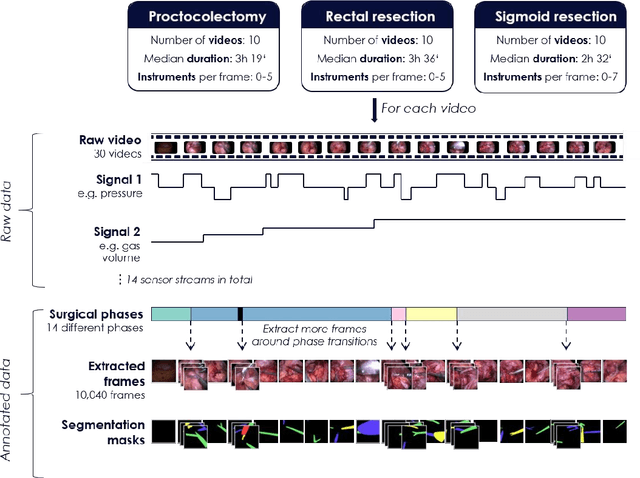
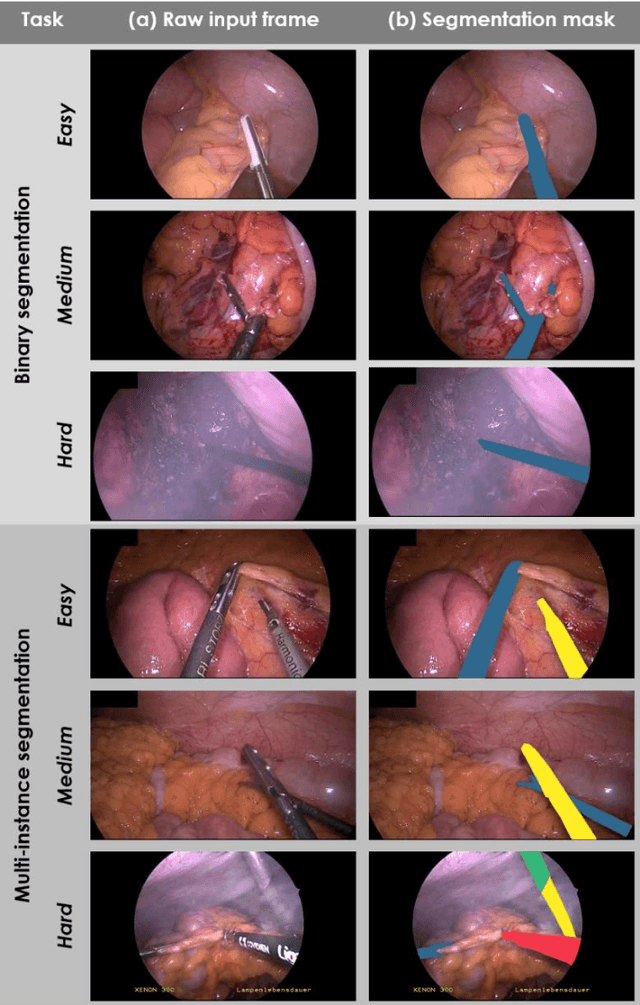
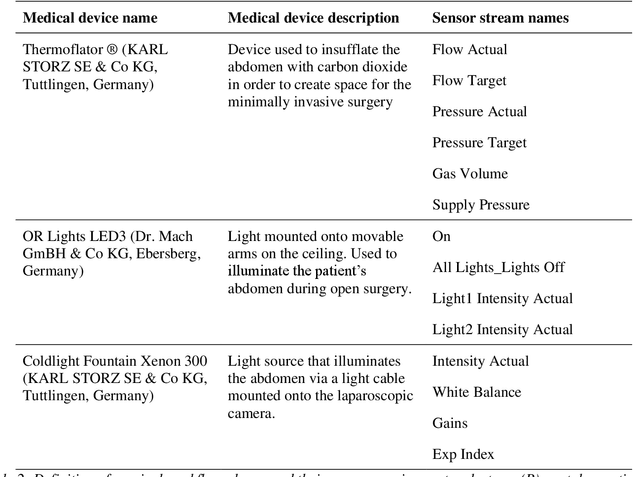
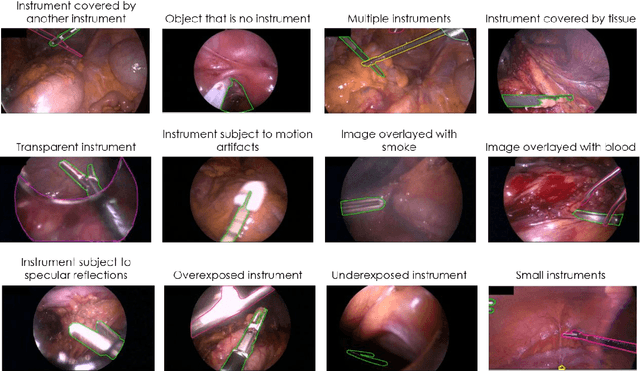
Abstract:Image-based tracking of medical instruments is an integral part of many surgical data science applications. Previous research has addressed the tasks of detecting, segmenting and tracking medical instruments based on laparoscopic video data. However, the methods proposed still tend to fail when applied to challenging images and do not generalize well to data they have not been trained on. This paper introduces the Heidelberg Colorectal (HeiCo) data set - the first publicly available data set enabling comprehensive benchmarking of medical instrument detection and segmentation algorithms with a specific emphasis on robustness and generalization capabilities of the methods. Our data set comprises 30 laparoscopic videos and corresponding sensor data from medical devices in the operating room for three different types of laparoscopic surgery. Annotations include surgical phase labels for all frames in the videos as well as instance-wise segmentation masks for surgical instruments in more than 10,000 individual frames. The data has successfully been used to organize international competitions in the scope of the Endoscopic Vision Challenges (EndoVis) 2017 and 2019.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge