Naoto Usuyama
Learning Sparse Visual Representations via Spatial-Semantic Factorization
Feb 02, 2026Abstract:Self-supervised learning (SSL) faces a fundamental conflict between semantic understanding and image reconstruction. High-level semantic SSL (e.g., DINO) relies on global tokens that are forced to be location-invariant for augmentation alignment, a process that inherently discards the spatial coordinates required for reconstruction. Conversely, generative SSL (e.g., MAE) preserves dense feature grids for reconstruction but fails to produce high-level abstractions. We introduce STELLAR, a framework that resolves this tension by factorizing visual features into a low-rank product of semantic concepts and their spatial distributions. This disentanglement allows us to perform DINO-style augmentation alignment on the semantic tokens while maintaining the precise spatial mapping in the localization matrix necessary for pixel-level reconstruction. We demonstrate that as few as 16 sparse tokens under this factorized form are sufficient to simultaneously support high-quality reconstruction (2.60 FID) and match the semantic performance of dense backbones (79.10% ImageNet accuracy). Our results highlight STELLAR as a versatile sparse representation that bridges the gap between discriminative and generative vision by strategically separating semantic identity from spatial geometry. Code available at https://aka.ms/stellar.
Exploring Scaling Laws for EHR Foundation Models
May 29, 2025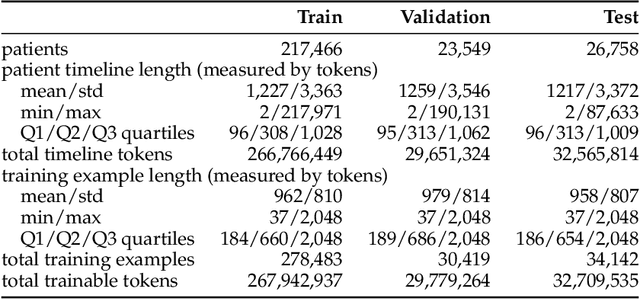
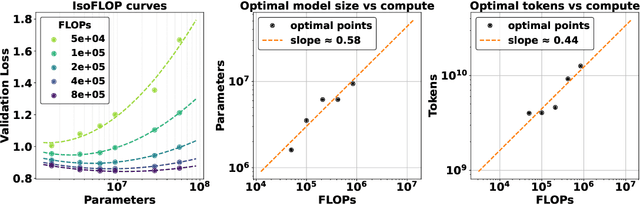
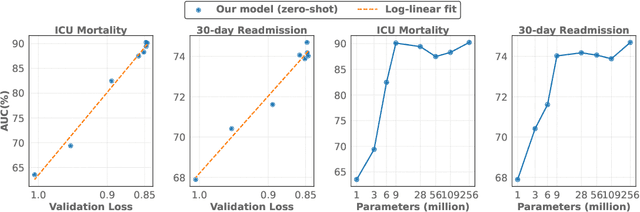
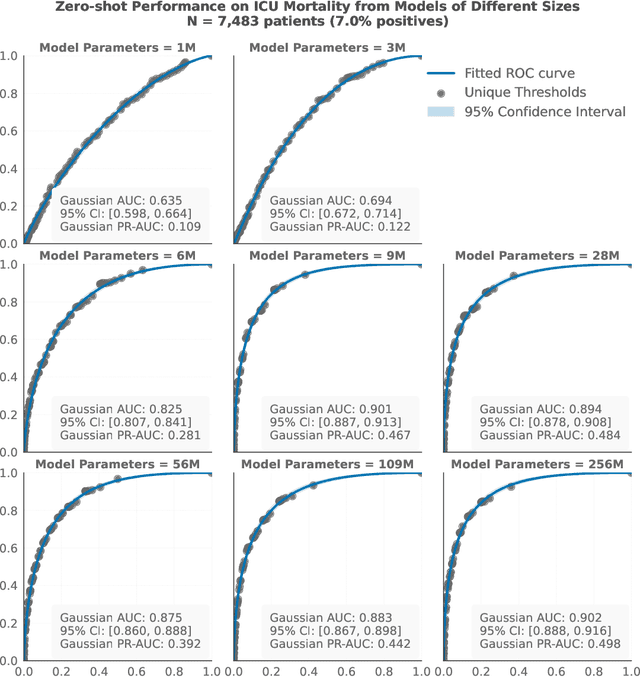
Abstract:The emergence of scaling laws has profoundly shaped the development of large language models (LLMs), enabling predictable performance gains through systematic increases in model size, dataset volume, and compute. Yet, these principles remain largely unexplored in the context of electronic health records (EHRs) -- a rich, sequential, and globally abundant data source that differs structurally from natural language. In this work, we present the first empirical investigation of scaling laws for EHR foundation models. By training transformer architectures on patient timeline data from the MIMIC-IV database across varying model sizes and compute budgets, we identify consistent scaling patterns, including parabolic IsoFLOPs curves and power-law relationships between compute, model parameters, data size, and clinical utility. These findings demonstrate that EHR models exhibit scaling behavior analogous to LLMs, offering predictive insights into resource-efficient training strategies. Our results lay the groundwork for developing powerful EHR foundation models capable of transforming clinical prediction tasks and advancing personalized healthcare.
Boltzmann Attention Sampling for Image Analysis with Small Objects
Mar 04, 2025Abstract:Detecting and segmenting small objects, such as lung nodules and tumor lesions, remains a critical challenge in image analysis. These objects often occupy less than 0.1% of an image, making traditional transformer architectures inefficient and prone to performance degradation due to redundant attention computations on irrelevant regions. Existing sparse attention mechanisms rely on rigid hierarchical structures, which are poorly suited for detecting small, variable, and uncertain object locations. In this paper, we propose BoltzFormer, a novel transformer-based architecture designed to address these challenges through dynamic sparse attention. BoltzFormer identifies and focuses attention on relevant areas by modeling uncertainty using a Boltzmann distribution with an annealing schedule. Initially, a higher temperature allows broader area sampling in early layers, when object location uncertainty is greatest. As the temperature decreases in later layers, attention becomes more focused, enhancing efficiency and accuracy. BoltzFormer seamlessly integrates into existing transformer architectures via a modular Boltzmann attention sampling mechanism. Comprehensive evaluations on benchmark datasets demonstrate that BoltzFormer significantly improves segmentation performance for small objects while reducing attention computation by an order of magnitude compared to previous state-of-the-art methods.
From Medprompt to o1: Exploration of Run-Time Strategies for Medical Challenge Problems and Beyond
Nov 06, 2024



Abstract:Run-time steering strategies like Medprompt are valuable for guiding large language models (LLMs) to top performance on challenging tasks. Medprompt demonstrates that a general LLM can be focused to deliver state-of-the-art performance on specialized domains like medicine by using a prompt to elicit a run-time strategy involving chain of thought reasoning and ensembling. OpenAI's o1-preview model represents a new paradigm, where a model is designed to do run-time reasoning before generating final responses. We seek to understand the behavior of o1-preview on a diverse set of medical challenge problem benchmarks. Following on the Medprompt study with GPT-4, we systematically evaluate the o1-preview model across various medical benchmarks. Notably, even without prompting techniques, o1-preview largely outperforms the GPT-4 series with Medprompt. We further systematically study the efficacy of classic prompt engineering strategies, as represented by Medprompt, within the new paradigm of reasoning models. We found that few-shot prompting hinders o1's performance, suggesting that in-context learning may no longer be an effective steering approach for reasoning-native models. While ensembling remains viable, it is resource-intensive and requires careful cost-performance optimization. Our cost and accuracy analysis across run-time strategies reveals a Pareto frontier, with GPT-4o representing a more affordable option and o1-preview achieving state-of-the-art performance at higher cost. Although o1-preview offers top performance, GPT-4o with steering strategies like Medprompt retains value in specific contexts. Moreover, we note that the o1-preview model has reached near-saturation on many existing medical benchmarks, underscoring the need for new, challenging benchmarks. We close with reflections on general directions for inference-time computation with LLMs.
BiomedParse: a biomedical foundation model for image parsing of everything everywhere all at once
May 21, 2024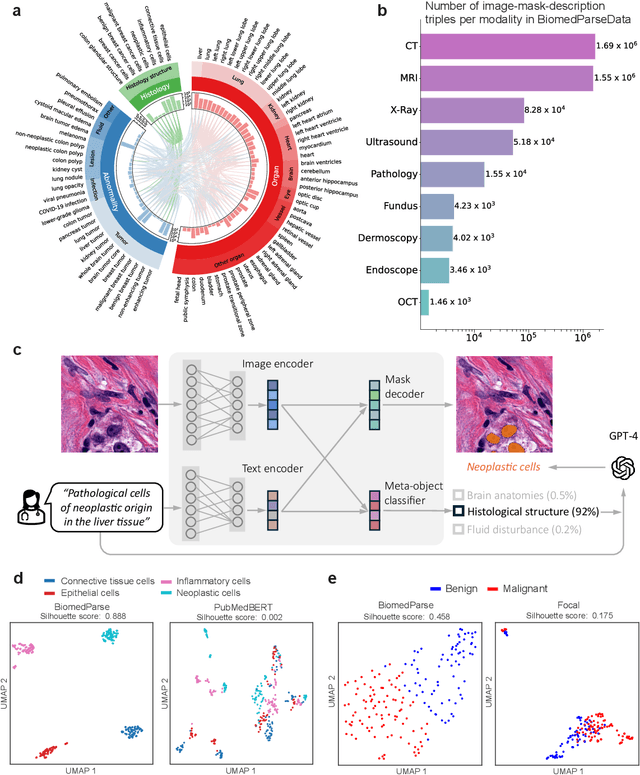
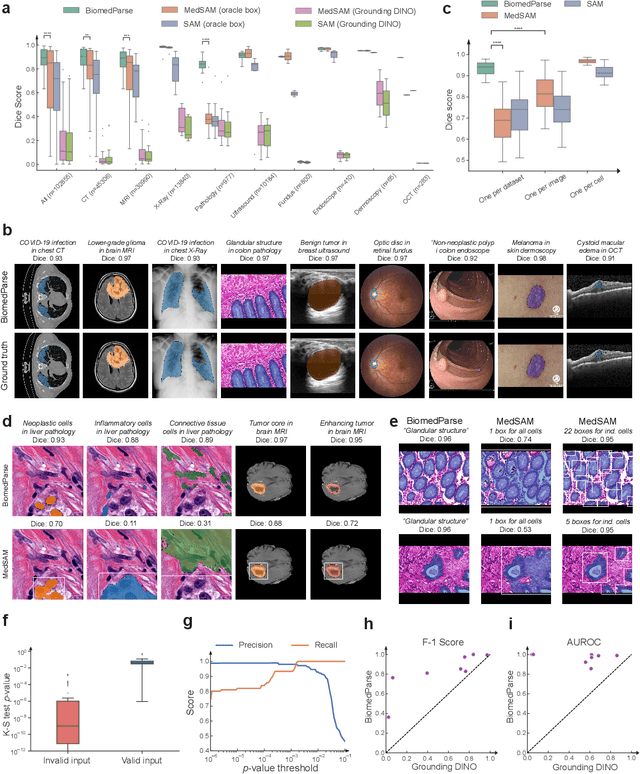
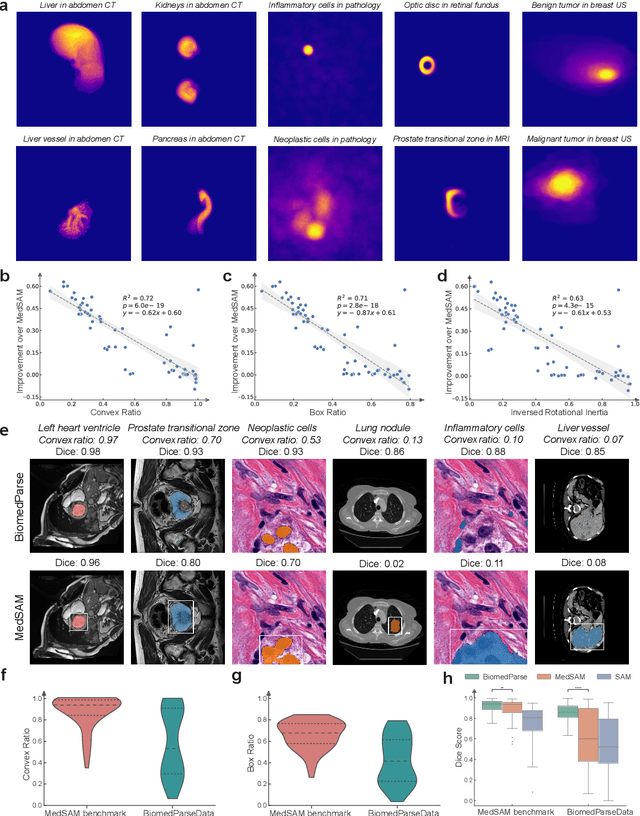
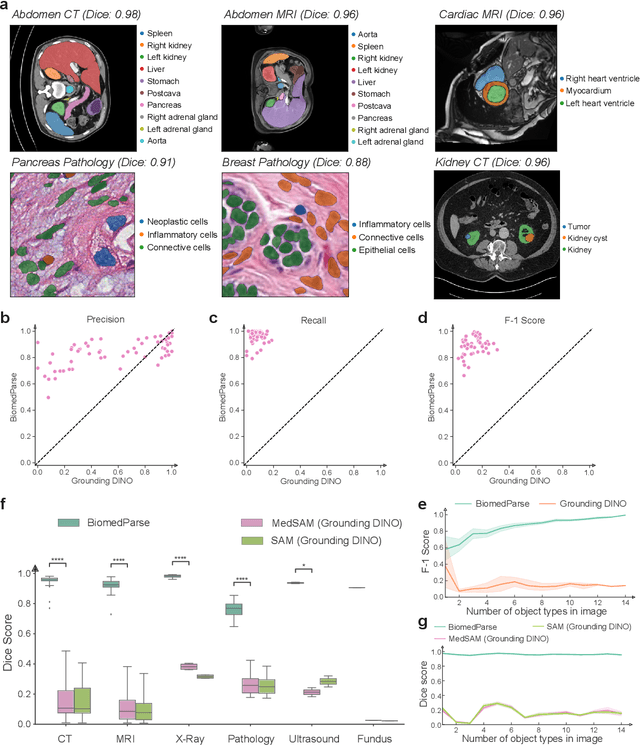
Abstract:Biomedical image analysis is fundamental for biomedical discovery in cell biology, pathology, radiology, and many other biomedical domains. Holistic image analysis comprises interdependent subtasks such as segmentation, detection, and recognition of relevant objects. Here, we propose BiomedParse, a biomedical foundation model for imaging parsing that can jointly conduct segmentation, detection, and recognition for 82 object types across 9 imaging modalities. Through joint learning, we can improve accuracy for individual tasks and enable novel applications such as segmenting all relevant objects in an image through a text prompt, rather than requiring users to laboriously specify the bounding box for each object. We leveraged readily available natural-language labels or descriptions accompanying those datasets and use GPT-4 to harmonize the noisy, unstructured text information with established biomedical object ontologies. We created a large dataset comprising over six million triples of image, segmentation mask, and textual description. On image segmentation, we showed that BiomedParse is broadly applicable, outperforming state-of-the-art methods on 102,855 test image-mask-label triples across 9 imaging modalities (everything). On object detection, which aims to locate a specific object of interest, BiomedParse again attained state-of-the-art performance, especially on objects with irregular shapes (everywhere). On object recognition, which aims to identify all objects in a given image along with their semantic types, we showed that BiomedParse can simultaneously segment and label all biomedical objects in an image (all at once). In summary, BiomedParse is an all-in-one tool for biomedical image analysis by jointly solving segmentation, detection, and recognition for all major biomedical image modalities, paving the path for efficient and accurate image-based biomedical discovery.
Training Small Multimodal Models to Bridge Biomedical Competency Gap: A Case Study in Radiology Imaging
Mar 20, 2024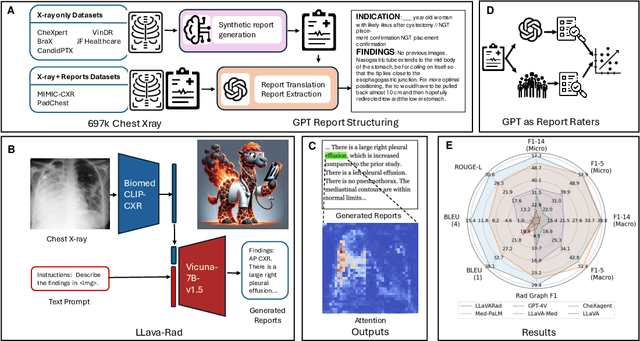
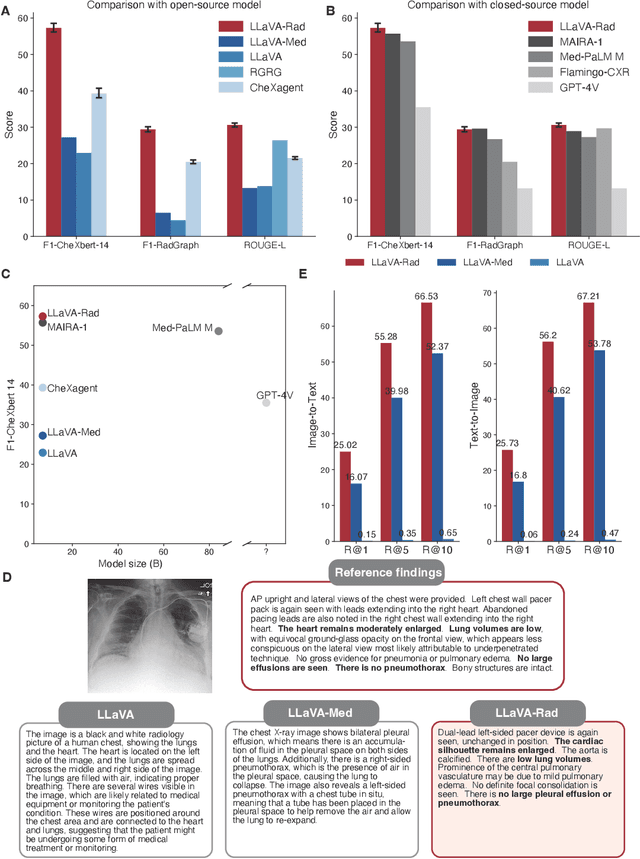
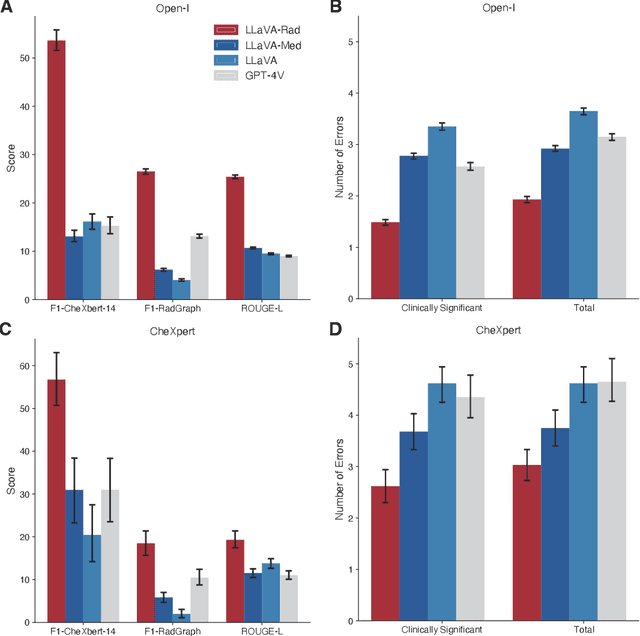
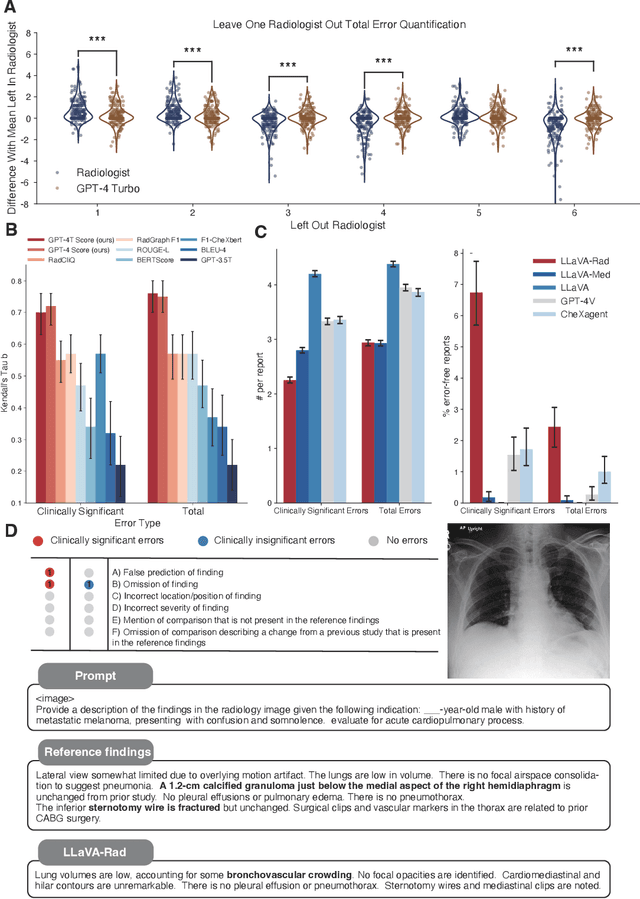
Abstract:The scaling laws and extraordinary performance of large foundation models motivate the development and utilization of such large models in biomedicine. However, despite early promising results on some biomedical benchmarks, there are still major challenges that need to be addressed before these models can be used in real-world applications. Frontier models such as GPT-4V still have major competency gaps in multimodal capabilities for biomedical applications. Moreover, pragmatic issues such as access, cost, latency, and compliance make it hard for clinicians to use privately-hosted state-of-the-art large models directly on private patient data. In this paper, we explore training open-source small multimodal models (SMMs) to bridge biomedical competency gaps for unmet clinical needs. To maximize data efficiency, we adopt a modular approach by incorporating state-of-the-art pre-trained models for image and text modalities, and focusing on training a lightweight adapter to ground each modality to the text embedding space. We conduct a comprehensive study of this approach on radiology imaging. For training, we assemble a large dataset with over 1 million image-text pairs. For evaluation, we propose a clinically driven novel approach using GPT-4 and demonstrate its parity with expert evaluation. We also study grounding qualitatively using attention. For best practice, we conduct a systematic ablation study on various choices in data engineering and multimodal training. The resulting LLaVA-Rad (7B) model attains state-of-the-art results on radiology tasks such as report generation and cross-modal retrieval, even outperforming much larger models such as GPT-4V and Med-PaLM M (84B). LLaVA-Rad is fast and can be run on a single V100 GPU in private settings, offering a promising state-of-the-art tool for real-world clinical applications.
Foundation Models for Biomedical Image Segmentation: A Survey
Jan 15, 2024Abstract:Recent advancements in biomedical image analysis have been significantly driven by the Segment Anything Model (SAM). This transformative technology, originally developed for general-purpose computer vision, has found rapid application in medical image processing. Within the last year, marked by over 100 publications, SAM has demonstrated its prowess in zero-shot learning adaptations for medical imaging. The fundamental premise of SAM lies in its capability to segment or identify objects in images without prior knowledge of the object type or imaging modality. This approach aligns well with tasks achievable by the human visual system, though its application in non-biological vision contexts remains more theoretically challenging. A notable feature of SAM is its ability to adjust segmentation according to a specified resolution scale or area of interest, akin to semantic priming. This adaptability has spurred a wave of creativity and innovation in applying SAM to medical imaging. Our review focuses on the period from April 1, 2023, to September 30, 2023, a critical first six months post-initial publication. We examine the adaptations and integrations of SAM necessary to address longstanding clinical challenges, particularly in the context of 33 open datasets covered in our analysis. While SAM approaches or achieves state-of-the-art performance in numerous applications, it falls short in certain areas, such as segmentation of the carotid artery, adrenal glands, optic nerve, and mandible bone. Our survey delves into the innovative techniques where SAM's foundational approach excels and explores the core concepts in translating and applying these models effectively in diverse medical imaging scenarios.
When an Image is Worth 1,024 x 1,024 Words: A Case Study in Computational Pathology
Dec 06, 2023Abstract:This technical report presents LongViT, a vision Transformer that can process gigapixel images in an end-to-end manner. Specifically, we split the gigapixel image into a sequence of millions of patches and project them linearly into embeddings. LongNet is then employed to model the extremely long sequence, generating representations that capture both short-range and long-range dependencies. The linear computation complexity of LongNet, along with its distributed algorithm, enables us to overcome the constraints of both computation and memory. We apply LongViT in the field of computational pathology, aiming for cancer diagnosis and prognosis within gigapixel whole-slide images. Experimental results demonstrate that LongViT effectively encodes gigapixel images and outperforms previous state-of-the-art methods on cancer subtyping and survival prediction. Code and models will be available at https://aka.ms/LongViT.
Can Generalist Foundation Models Outcompete Special-Purpose Tuning? Case Study in Medicine
Nov 28, 2023
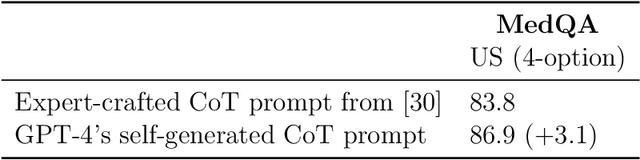
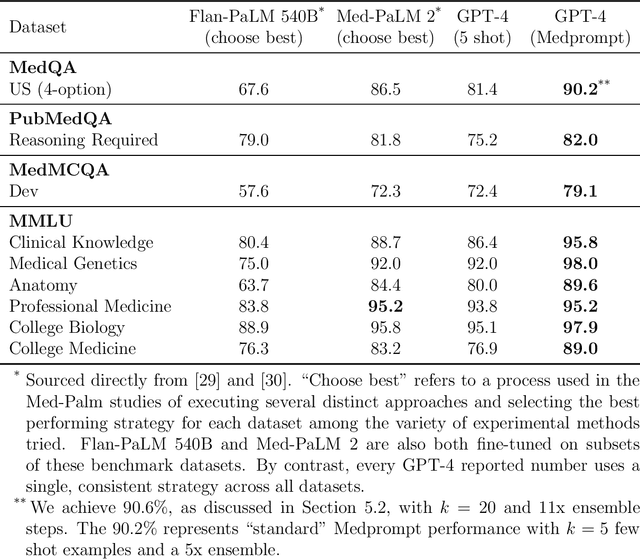


Abstract:Generalist foundation models such as GPT-4 have displayed surprising capabilities in a wide variety of domains and tasks. Yet, there is a prevalent assumption that they cannot match specialist capabilities of fine-tuned models. For example, most explorations to date on medical competency benchmarks have leveraged domain-specific training, as exemplified by efforts on BioGPT and Med-PaLM. We build on a prior study of GPT-4's capabilities on medical challenge benchmarks in the absence of special training. Rather than using simple prompting to highlight the model's out-of-the-box capabilities, we perform a systematic exploration of prompt engineering. We find that prompting innovation can unlock deeper specialist capabilities and show that GPT-4 easily tops prior leading results for medical benchmarks. The prompting methods we explore are general purpose, and make no specific use of domain expertise, removing the need for expert-curated content. Our experimental design carefully controls for overfitting during the prompt engineering process. We introduce Medprompt, based on a composition of several prompting strategies. With Medprompt, GPT-4 achieves state-of-the-art results on all nine of the benchmark datasets in the MultiMedQA suite. The method outperforms leading specialist models such as Med-PaLM 2 by a significant margin with an order of magnitude fewer calls to the model. Steering GPT-4 with Medprompt achieves a 27% reduction in error rate on the MedQA dataset over the best methods to date achieved with specialist models and surpasses a score of 90% for the first time. Beyond medical problems, we show the power of Medprompt to generalize to other domains and provide evidence for the broad applicability of the approach via studies of the strategy on exams in electrical engineering, machine learning, philosophy, accounting, law, nursing, and clinical psychology.
Exploring the Boundaries of GPT-4 in Radiology
Oct 23, 2023



Abstract:The recent success of general-domain large language models (LLMs) has significantly changed the natural language processing paradigm towards a unified foundation model across domains and applications. In this paper, we focus on assessing the performance of GPT-4, the most capable LLM so far, on the text-based applications for radiology reports, comparing against state-of-the-art (SOTA) radiology-specific models. Exploring various prompting strategies, we evaluated GPT-4 on a diverse range of common radiology tasks and we found GPT-4 either outperforms or is on par with current SOTA radiology models. With zero-shot prompting, GPT-4 already obtains substantial gains ($\approx$ 10% absolute improvement) over radiology models in temporal sentence similarity classification (accuracy) and natural language inference ($F_1$). For tasks that require learning dataset-specific style or schema (e.g. findings summarisation), GPT-4 improves with example-based prompting and matches supervised SOTA. Our extensive error analysis with a board-certified radiologist shows GPT-4 has a sufficient level of radiology knowledge with only occasional errors in complex context that require nuanced domain knowledge. For findings summarisation, GPT-4 outputs are found to be overall comparable with existing manually-written impressions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge