Training Small Multimodal Models to Bridge Biomedical Competency Gap: A Case Study in Radiology Imaging
Paper and Code
Mar 20, 2024
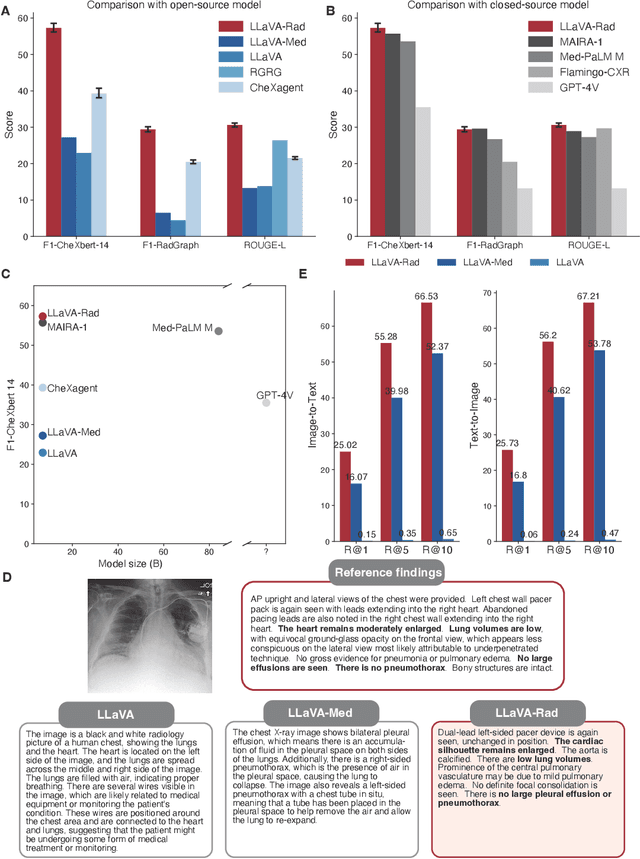
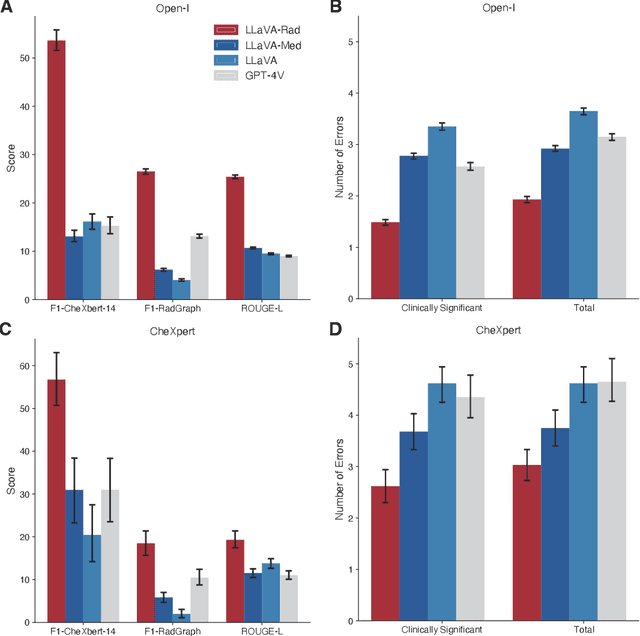
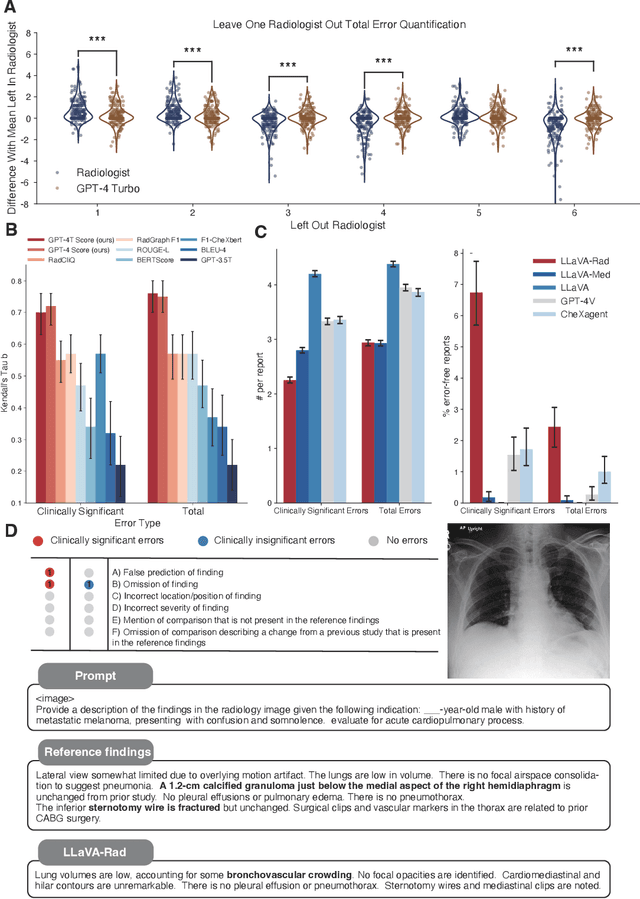
The scaling laws and extraordinary performance of large foundation models motivate the development and utilization of such large models in biomedicine. However, despite early promising results on some biomedical benchmarks, there are still major challenges that need to be addressed before these models can be used in real-world applications. Frontier models such as GPT-4V still have major competency gaps in multimodal capabilities for biomedical applications. Moreover, pragmatic issues such as access, cost, latency, and compliance make it hard for clinicians to use privately-hosted state-of-the-art large models directly on private patient data. In this paper, we explore training open-source small multimodal models (SMMs) to bridge biomedical competency gaps for unmet clinical needs. To maximize data efficiency, we adopt a modular approach by incorporating state-of-the-art pre-trained models for image and text modalities, and focusing on training a lightweight adapter to ground each modality to the text embedding space. We conduct a comprehensive study of this approach on radiology imaging. For training, we assemble a large dataset with over 1 million image-text pairs. For evaluation, we propose a clinically driven novel approach using GPT-4 and demonstrate its parity with expert evaluation. We also study grounding qualitatively using attention. For best practice, we conduct a systematic ablation study on various choices in data engineering and multimodal training. The resulting LLaVA-Rad (7B) model attains state-of-the-art results on radiology tasks such as report generation and cross-modal retrieval, even outperforming much larger models such as GPT-4V and Med-PaLM M (84B). LLaVA-Rad is fast and can be run on a single V100 GPU in private settings, offering a promising state-of-the-art tool for real-world clinical applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge