Harshita Sharma
Data Scaling Laws for Radiology Foundation Models
Sep 16, 2025Abstract:Foundation vision encoders such as CLIP and DINOv2, trained on web-scale data, exhibit strong transfer performance across tasks and datasets. However, medical imaging foundation models remain constrained by smaller datasets, limiting our understanding of how data scale and pretraining paradigms affect performance in this setting. In this work, we systematically study continual pretraining of two vision encoders, MedImageInsight (MI2) and RAD-DINO representing the two major encoder paradigms CLIP and DINOv2, on up to 3.5M chest x-rays from a single institution, holding compute and evaluation protocols constant. We evaluate on classification (radiology findings, lines and tubes), segmentation (lines and tubes), and radiology report generation. While prior work has primarily focused on tasks related to radiology findings, we include lines and tubes tasks to counterbalance this bias and evaluate a model's ability to extract features that preserve continuity along elongated structures. Our experiments show that MI2 scales more effectively for finding-related tasks, while RAD-DINO is stronger on tube-related tasks. Surprisingly, continually pretraining MI2 with both reports and structured labels using UniCL improves performance, underscoring the value of structured supervision at scale. We further show that for some tasks, as few as 30k in-domain samples are sufficient to surpass open-weights foundation models. These results highlight the utility of center-specific continual pretraining, enabling medical institutions to derive significant performance gains by utilizing in-domain data.
Lunguage: A Benchmark for Structured and Sequential Chest X-ray Interpretation
May 27, 2025Abstract:Radiology reports convey detailed clinical observations and capture diagnostic reasoning that evolves over time. However, existing evaluation methods are limited to single-report settings and rely on coarse metrics that fail to capture fine-grained clinical semantics and temporal dependencies. We introduce LUNGUAGE,a benchmark dataset for structured radiology report generation that supports both single-report evaluation and longitudinal patient-level assessment across multiple studies. It contains 1,473 annotated chest X-ray reports, each reviewed by experts, and 80 of them contain longitudinal annotations to capture disease progression and inter-study intervals, also reviewed by experts. Using this benchmark, we develop a two-stage framework that transforms generated reports into fine-grained, schema-aligned structured representations, enabling longitudinal interpretation. We also propose LUNGUAGESCORE, an interpretable metric that compares structured outputs at the entity, relation, and attribute level while modeling temporal consistency across patient timelines. These contributions establish the first benchmark dataset, structuring framework, and evaluation metric for sequential radiology reporting, with empirical results demonstrating that LUNGUAGESCORE effectively supports structured report evaluation. The code is available at: https://github.com/SuperSupermoon/Lunguage
MedHELM: Holistic Evaluation of Large Language Models for Medical Tasks
May 26, 2025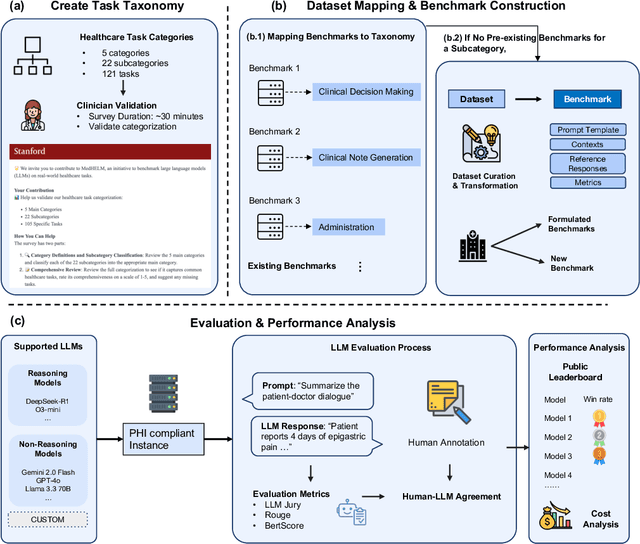
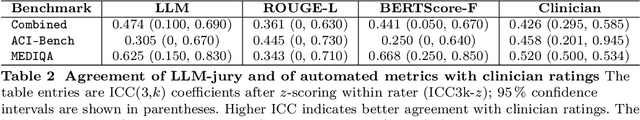
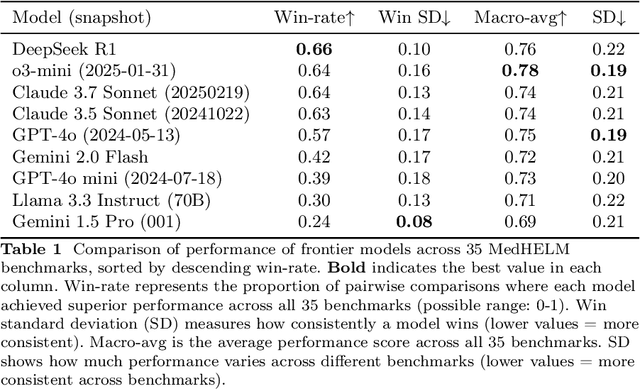
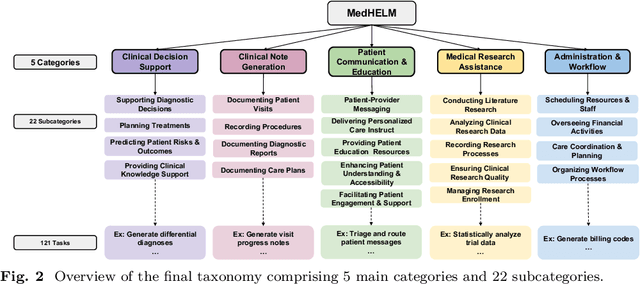

Abstract:While large language models (LLMs) achieve near-perfect scores on medical licensing exams, these evaluations inadequately reflect the complexity and diversity of real-world clinical practice. We introduce MedHELM, an extensible evaluation framework for assessing LLM performance for medical tasks with three key contributions. First, a clinician-validated taxonomy spanning 5 categories, 22 subcategories, and 121 tasks developed with 29 clinicians. Second, a comprehensive benchmark suite comprising 35 benchmarks (17 existing, 18 newly formulated) providing complete coverage of all categories and subcategories in the taxonomy. Third, a systematic comparison of LLMs with improved evaluation methods (using an LLM-jury) and a cost-performance analysis. Evaluation of 9 frontier LLMs, using the 35 benchmarks, revealed significant performance variation. Advanced reasoning models (DeepSeek R1: 66% win-rate; o3-mini: 64% win-rate) demonstrated superior performance, though Claude 3.5 Sonnet achieved comparable results at 40% lower estimated computational cost. On a normalized accuracy scale (0-1), most models performed strongly in Clinical Note Generation (0.73-0.85) and Patient Communication & Education (0.78-0.83), moderately in Medical Research Assistance (0.65-0.75), and generally lower in Clinical Decision Support (0.56-0.72) and Administration & Workflow (0.53-0.63). Our LLM-jury evaluation method achieved good agreement with clinician ratings (ICC = 0.47), surpassing both average clinician-clinician agreement (ICC = 0.43) and automated baselines including ROUGE-L (0.36) and BERTScore-F1 (0.44). Claude 3.5 Sonnet achieved comparable performance to top models at lower estimated cost. These findings highlight the importance of real-world, task-specific evaluation for medical use of LLMs and provides an open source framework to enable this.
MAIRA-Seg: Enhancing Radiology Report Generation with Segmentation-Aware Multimodal Large Language Models
Nov 18, 2024



Abstract:There is growing interest in applying AI to radiology report generation, particularly for chest X-rays (CXRs). This paper investigates whether incorporating pixel-level information through segmentation masks can improve fine-grained image interpretation of multimodal large language models (MLLMs) for radiology report generation. We introduce MAIRA-Seg, a segmentation-aware MLLM framework designed to utilize semantic segmentation masks alongside CXRs for generating radiology reports. We train expert segmentation models to obtain mask pseudolabels for radiology-specific structures in CXRs. Subsequently, building on the architectures of MAIRA, a CXR-specialised model for report generation, we integrate a trainable segmentation tokens extractor that leverages these mask pseudolabels, and employ mask-aware prompting to generate draft radiology reports. Our experiments on the publicly available MIMIC-CXR dataset show that MAIRA-Seg outperforms non-segmentation baselines. We also investigate set-of-marks prompting with MAIRA and find that MAIRA-Seg consistently demonstrates comparable or superior performance. The results confirm that using segmentation masks enhances the nuanced reasoning of MLLMs, potentially contributing to better clinical outcomes.
MAIRA-2: Grounded Radiology Report Generation
Jun 06, 2024



Abstract:Radiology reporting is a complex task that requires detailed image understanding, integration of multiple inputs, including comparison with prior imaging, and precise language generation. This makes it ideal for the development and use of generative multimodal models. Here, we extend report generation to include the localisation of individual findings on the image - a task we call grounded report generation. Prior work indicates that grounding is important for clarifying image understanding and interpreting AI-generated text. Therefore, grounded reporting stands to improve the utility and transparency of automated report drafting. To enable evaluation of grounded reporting, we propose a novel evaluation framework - RadFact - leveraging the reasoning capabilities of large language models (LLMs). RadFact assesses the factuality of individual generated sentences, as well as correctness of generated spatial localisations when present. We introduce MAIRA-2, a large multimodal model combining a radiology-specific image encoder with a LLM, and trained for the new task of grounded report generation on chest X-rays. MAIRA-2 uses more comprehensive inputs than explored previously: the current frontal image, the current lateral image, the prior frontal image and prior report, as well as the Indication, Technique and Comparison sections of the current report. We demonstrate that these additions significantly improve report quality and reduce hallucinations, establishing a new state of the art on findings generation (without grounding) on MIMIC-CXR while demonstrating the feasibility of grounded reporting as a novel and richer task.
Challenges for Responsible AI Design and Workflow Integration in Healthcare: A Case Study of Automatic Feeding Tube Qualification in Radiology
May 08, 2024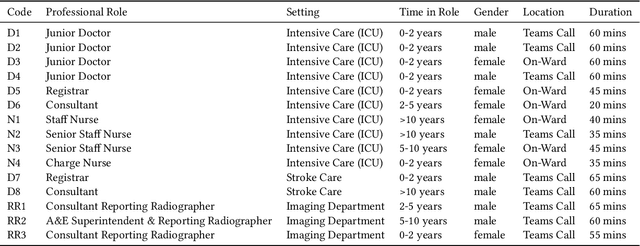
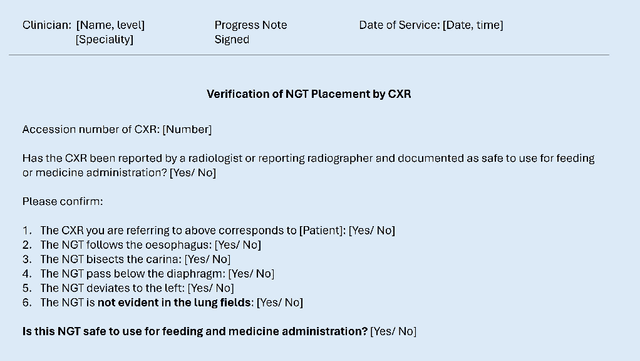
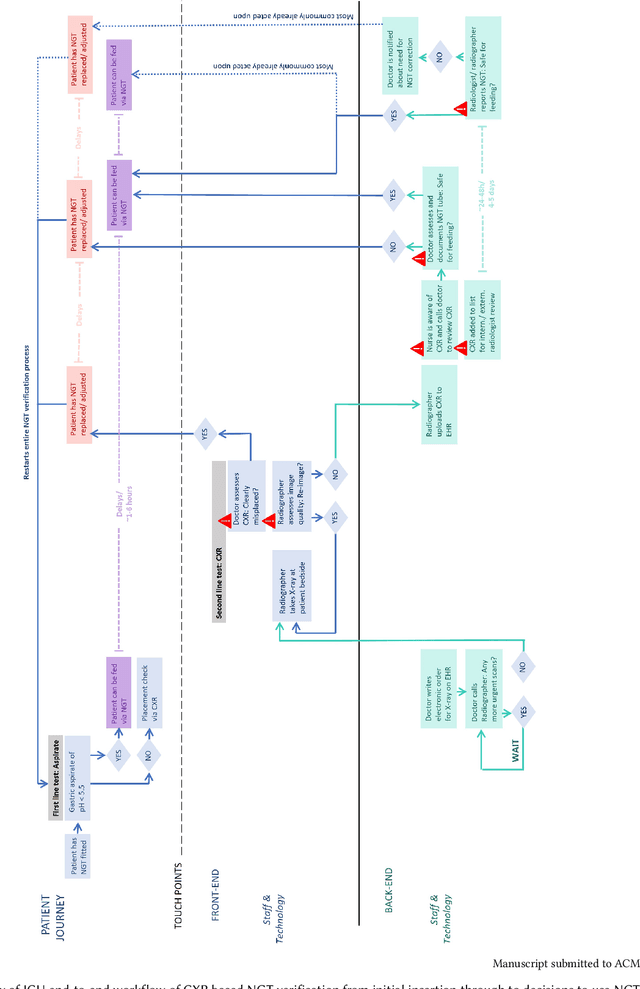


Abstract:Nasogastric tubes (NGTs) are feeding tubes that are inserted through the nose into the stomach to deliver nutrition or medication. If not placed correctly, they can cause serious harm, even death to patients. Recent AI developments demonstrate the feasibility of robustly detecting NGT placement from Chest X-ray images to reduce risks of sub-optimally or critically placed NGTs being missed or delayed in their detection, but gaps remain in clinical practice integration. In this study, we present a human-centered approach to the problem and describe insights derived following contextual inquiry and in-depth interviews with 15 clinical stakeholders. The interviews helped understand challenges in existing workflows, and how best to align technical capabilities with user needs and expectations. We discovered the trade-offs and complexities that need consideration when choosing suitable workflow stages, target users, and design configurations for different AI proposals. We explored how to balance AI benefits and risks for healthcare staff and patients within broader organizational and medical-legal constraints. We also identified data issues related to edge cases and data biases that affect model training and evaluation; how data documentation practices influence data preparation and labelling; and how to measure relevant AI outcomes reliably in future evaluations. We discuss how our work informs design and development of AI applications that are clinically useful, ethical, and acceptable in real-world healthcare services.
RAD-DINO: Exploring Scalable Medical Image Encoders Beyond Text Supervision
Jan 19, 2024Abstract:Language-supervised pre-training has proven to be a valuable method for extracting semantically meaningful features from images, serving as a foundational element in multimodal systems within the computer vision and medical imaging domains. However, resulting features are limited by the information contained within the text. This is particularly problematic in medical imaging, where radiologists' written findings focus on specific observations; a challenge compounded by the scarcity of paired imaging-text data due to concerns over leakage of personal health information. In this work, we fundamentally challenge the prevailing reliance on language supervision for learning general purpose biomedical imaging encoders. We introduce RAD-DINO, a biomedical image encoder pre-trained solely on unimodal biomedical imaging data that obtains similar or greater performance than state-of-the-art biomedical language supervised models on a diverse range of benchmarks. Specifically, the quality of learned representations is evaluated on standard imaging tasks (classification and semantic segmentation), and a vision-language alignment task (text report generation from images). To further demonstrate the drawback of language supervision, we show that features from RAD-DINO correlate with other medical records (e.g., sex or age) better than language-supervised models, which are generally not mentioned in radiology reports. Finally, we conduct a series of ablations determining the factors in RAD-DINO's performance; notably, we observe that RAD-DINO's downstream performance scales well with the quantity and diversity of training data, demonstrating that image-only supervision is a scalable approach for training a foundational biomedical image encoder.
RadEdit: stress-testing biomedical vision models via diffusion image editing
Dec 21, 2023Abstract:Biomedical imaging datasets are often small and biased, meaning that real-world performance of predictive models can be substantially lower than expected from internal testing. This work proposes using generative image editing to simulate dataset shifts and diagnose failure modes of biomedical vision models; this can be used in advance of deployment to assess readiness, potentially reducing cost and patient harm. Existing editing methods can produce undesirable changes, with spurious correlations learned due to the co-occurrence of disease and treatment interventions, limiting practical applicability. To address this, we train a text-to-image diffusion model on multiple chest X-ray datasets and introduce a new editing method RadEdit that uses multiple masks, if present, to constrain changes and ensure consistency in the edited images. We consider three types of dataset shifts: acquisition shift, manifestation shift, and population shift, and demonstrate that our approach can diagnose failures and quantify model robustness without additional data collection, complementing more qualitative tools for explainable AI.
Verb Categorisation for Hindi Word Problem Solving
Dec 18, 2023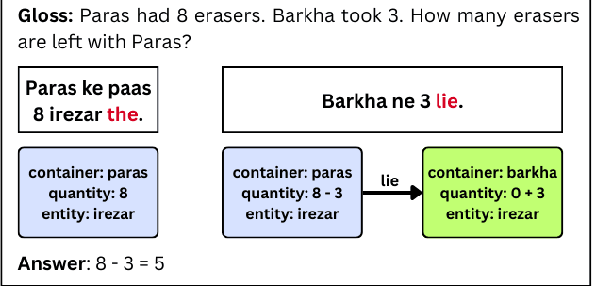



Abstract:Word problem Solving is a challenging NLP task that deals with solving mathematical problems described in natural language. Recently, there has been renewed interest in developing word problem solvers for Indian languages. As part of this paper, we have built a Hindi arithmetic word problem solver which makes use of verbs. Additionally, we have created verb categorization data for Hindi. Verbs are very important for solving word problems with addition/subtraction operations as they help us identify the set of operations required to solve the word problems. We propose a rule-based solver that uses verb categorisation to identify operations in a word problem and generate answers for it. To perform verb categorisation, we explore several approaches and present a comparative study.
Exploring the Boundaries of GPT-4 in Radiology
Oct 23, 2023



Abstract:The recent success of general-domain large language models (LLMs) has significantly changed the natural language processing paradigm towards a unified foundation model across domains and applications. In this paper, we focus on assessing the performance of GPT-4, the most capable LLM so far, on the text-based applications for radiology reports, comparing against state-of-the-art (SOTA) radiology-specific models. Exploring various prompting strategies, we evaluated GPT-4 on a diverse range of common radiology tasks and we found GPT-4 either outperforms or is on par with current SOTA radiology models. With zero-shot prompting, GPT-4 already obtains substantial gains ($\approx$ 10% absolute improvement) over radiology models in temporal sentence similarity classification (accuracy) and natural language inference ($F_1$). For tasks that require learning dataset-specific style or schema (e.g. findings summarisation), GPT-4 improves with example-based prompting and matches supervised SOTA. Our extensive error analysis with a board-certified radiologist shows GPT-4 has a sufficient level of radiology knowledge with only occasional errors in complex context that require nuanced domain knowledge. For findings summarisation, GPT-4 outputs are found to be overall comparable with existing manually-written impressions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge