Michael M. Bronstein
A Mechanistic Analysis of Looped Reasoning Language Models
Apr 13, 2026Abstract:Reasoning has become a central capability in large language models. Recent research has shown that reasoning performance can be improved by looping an LLM's layers in the latent dimension, resulting in looped reasoning language models. Despite promising results, few works have investigated how their internal dynamics differ from those of standard feedforward models. In this paper, we conduct a mechanistic analysis of the latent states in looped language models, focusing in particular on how the stages of inference observed in feedforward models compare to those observed in looped ones. To this end, we analyze cyclic recurrence and show that for many of the studied models each layer in the cycle converges to a distinct fixed point; consequently, the recurrent block follows a consistent cyclic trajectory in the latent space. We provide evidence that as these fixed points are reached, attention-head behavior stabilizes, leading to constant behavior across recurrences. Empirically, we discover that recurrent blocks learn stages of inference that closely mirror those of feedforward models, repeating these stages in depth with each iteration. We study how recurrent block size, input injection, and normalization influence the emergence and stability of these cyclic fixed points. We believe these findings help translate mechanistic insights into practical guidance for architectural design.
Scaling Atomistic Protein Binder Design with Generative Pretraining and Test-Time Compute
Mar 30, 2026Abstract:Protein interaction modeling is central to protein design, which has been transformed by machine learning with applications in drug discovery and beyond. In this landscape, structure-based de novo binder design is cast as either conditional generative modeling or sequence optimization via structure predictors ("hallucination"). We argue that this is a false dichotomy and propose Proteina-Complexa, a novel fully atomistic binder generation method unifying both paradigms. We extend recent flow-based latent protein generation architectures and leverage the domain-domain interactions of monomeric computationally predicted protein structures to construct Teddymer, a new large-scale dataset of synthetic binder-target pairs for pretraining. Combined with high-quality experimental multimers, this enables training a strong base model. We then perform inference-time optimization with this generative prior, unifying the strengths of previously distinct generative and hallucination methods. Proteina-Complexa sets a new state of the art in computational binder design benchmarks: it delivers markedly higher in-silico success rates than existing generative approaches, and our novel test-time optimization strategies greatly outperform previous hallucination methods under normalized compute budgets. We also demonstrate interface hydrogen bond optimization, fold class-guided binder generation, and extensions to small molecule targets and enzyme design tasks, again surpassing prior methods. Code, models and new data will be publicly released.
Generalised Flow Maps for Few-Step Generative Modelling on Riemannian Manifolds
Oct 24, 2025Abstract:Geometric data and purpose-built generative models on them have become ubiquitous in high-impact deep learning application domains, ranging from protein backbone generation and computational chemistry to geospatial data. Current geometric generative models remain computationally expensive at inference -- requiring many steps of complex numerical simulation -- as they are derived from dynamical measure transport frameworks such as diffusion and flow-matching on Riemannian manifolds. In this paper, we propose Generalised Flow Maps (GFM), a new class of few-step generative models that generalises the Flow Map framework in Euclidean spaces to arbitrary Riemannian manifolds. We instantiate GFMs with three self-distillation-based training methods: Generalised Lagrangian Flow Maps, Generalised Eulerian Flow Maps, and Generalised Progressive Flow Maps. We theoretically show that GFMs, under specific design decisions, unify and elevate existing Euclidean few-step generative models, such as consistency models, shortcut models, and meanflows, to the Riemannian setting. We benchmark GFMs against other geometric generative models on a suite of geometric datasets, including geospatial data, RNA torsion angles, and hyperbolic manifolds, and achieve state-of-the-art sample quality for single- and few-step evaluations, and superior or competitive log-likelihoods using the implicit probability flow.
ResCP: Reservoir Conformal Prediction for Time Series Forecasting
Oct 06, 2025Abstract:Conformal prediction offers a powerful framework for building distribution-free prediction intervals for exchangeable data. Existing methods that extend conformal prediction to sequential data rely on fitting a relatively complex model to capture temporal dependencies. However, these methods can fail if the sample size is small and often require expensive retraining when the underlying data distribution changes. To overcome these limitations, we propose Reservoir Conformal Prediction (ResCP), a novel training-free conformal prediction method for time series. Our approach leverages the efficiency and representation learning capabilities of reservoir computing to dynamically reweight conformity scores. In particular, we compute similarity scores among reservoir states and use them to adaptively reweight the observed residuals at each step. With this approach, ResCP enables us to account for local temporal dynamics when modeling the error distribution without compromising computational scalability. We prove that, under reasonable assumptions, ResCP achieves asymptotic conditional coverage, and we empirically demonstrate its effectiveness across diverse forecasting tasks.
Amortized Sampling with Transferable Normalizing Flows
Aug 25, 2025Abstract:Efficient equilibrium sampling of molecular conformations remains a core challenge in computational chemistry and statistical inference. Classical approaches such as molecular dynamics or Markov chain Monte Carlo inherently lack amortization; the computational cost of sampling must be paid in-full for each system of interest. The widespread success of generative models has inspired interest into overcoming this limitation through learning sampling algorithms. Despite performing on par with conventional methods when trained on a single system, learned samplers have so far demonstrated limited ability to transfer across systems. We prove that deep learning enables the design of scalable and transferable samplers by introducing Prose, a 280 million parameter all-atom transferable normalizing flow trained on a corpus of peptide molecular dynamics trajectories up to 8 residues in length. Prose draws zero-shot uncorrelated proposal samples for arbitrary peptide systems, achieving the previously intractable transferability across sequence length, whilst retaining the efficient likelihood evaluation of normalizing flows. Through extensive empirical evaluation we demonstrate the efficacy of Prose as a proposal for a variety of sampling algorithms, finding a simple importance sampling-based finetuning procedure to achieve superior performance to established methods such as sequential Monte Carlo on unseen tetrapeptides. We open-source the Prose codebase, model weights, and training dataset, to further stimulate research into amortized sampling methods and finetuning objectives.
Over-squashing in Spatiotemporal Graph Neural Networks
Jun 18, 2025


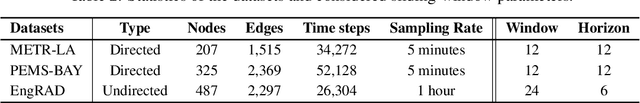
Abstract:Graph Neural Networks (GNNs) have achieved remarkable success across various domains. However, recent theoretical advances have identified fundamental limitations in their information propagation capabilities, such as over-squashing, where distant nodes fail to effectively exchange information. While extensively studied in static contexts, this issue remains unexplored in Spatiotemporal GNNs (STGNNs), which process sequences associated with graph nodes. Nonetheless, the temporal dimension amplifies this challenge by increasing the information that must be propagated. In this work, we formalize the spatiotemporal over-squashing problem and demonstrate its distinct characteristics compared to the static case. Our analysis reveals that counterintuitively, convolutional STGNNs favor information propagation from points temporally distant rather than close in time. Moreover, we prove that architectures that follow either time-and-space or time-then-space processing paradigms are equally affected by this phenomenon, providing theoretical justification for computationally efficient implementations. We validate our findings on synthetic and real-world datasets, providing deeper insights into their operational dynamics and principled guidance for more effective designs.
HYPER: A Foundation Model for Inductive Link Prediction with Knowledge Hypergraphs
Jun 14, 2025Abstract:Inductive link prediction with knowledge hypergraphs is the task of predicting missing hyperedges involving completely novel entities (i.e., nodes unseen during training). Existing methods for inductive link prediction with knowledge hypergraphs assume a fixed relational vocabulary and, as a result, cannot generalize to knowledge hypergraphs with novel relation types (i.e., relations unseen during training). Inspired by knowledge graph foundation models, we propose HYPER as a foundation model for link prediction, which can generalize to any knowledge hypergraph, including novel entities and novel relations. Importantly, HYPER can learn and transfer across different relation types of varying arities, by encoding the entities of each hyperedge along with their respective positions in the hyperedge. To evaluate HYPER, we construct 16 new inductive datasets from existing knowledge hypergraphs, covering a diverse range of relation types of varying arities. Empirically, HYPER consistently outperforms all existing methods in both node-only and node-and-relation inductive settings, showing strong generalization to unseen, higher-arity relational structures.
On Measuring Long-Range Interactions in Graph Neural Networks
Jun 06, 2025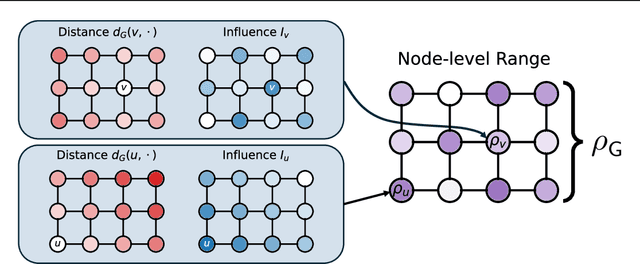
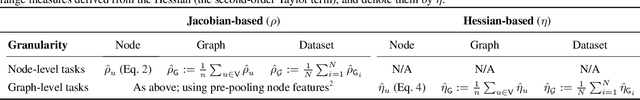
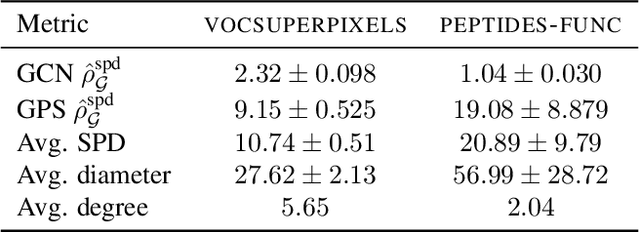



Abstract:Long-range graph tasks -- those dependent on interactions between distant nodes -- are an open problem in graph neural network research. Real-world benchmark tasks, especially the Long Range Graph Benchmark, have become popular for validating the long-range capability of proposed architectures. However, this is an empirical approach that lacks both robustness and theoretical underpinning; a more principled characterization of the long-range problem is required. To bridge this gap, we formalize long-range interactions in graph tasks, introduce a range measure for operators on graphs, and validate it with synthetic experiments. We then leverage our measure to examine commonly used tasks and architectures, and discuss to what extent they are, in fact, long-range. We believe our work advances efforts to define and address the long-range problem on graphs, and that our range measure will aid evaluation of new datasets and architectures.
Scalable Equilibrium Sampling with Sequential Boltzmann Generators
Feb 25, 2025

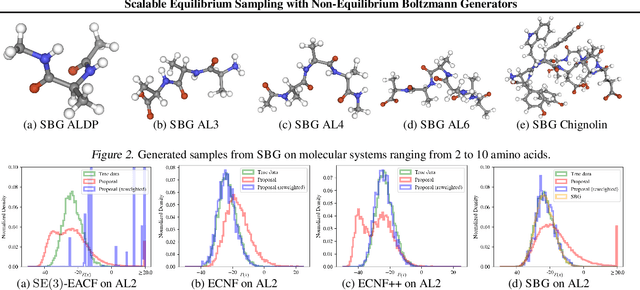

Abstract:Scalable sampling of molecular states in thermodynamic equilibrium is a long-standing challenge in statistical physics. Boltzmann generators tackle this problem by pairing powerful normalizing flows with importance sampling to obtain statistically independent samples under the target distribution. In this paper, we extend the Boltzmann generator framework and introduce Sequential Boltzmann generators (SBG) with two key improvements. The first is a highly efficient non-equivariant Transformer-based normalizing flow operating directly on all-atom Cartesian coordinates. In contrast to equivariant continuous flows of prior methods, we leverage exactly invertible non-equivariant architectures which are highly efficient both during sample generation and likelihood computation. As a result, this unlocks more sophisticated inference strategies beyond standard importance sampling. More precisely, as a second key improvement we perform inference-time scaling of flow samples using annealed Langevin dynamics which transports samples toward the target distribution leading to lower variance (annealed) importance weights which enable higher fidelity resampling with sequential Monte Carlo. SBG achieves state-of-the-art performance w.r.t. all metrics on molecular systems, demonstrating the first equilibrium sampling in Cartesian coordinates of tri, tetra, and hexapeptides that were so far intractable for prior Boltzmann generators.
Position: Graph Learning Will Lose Relevance Due To Poor Benchmarks
Feb 20, 2025
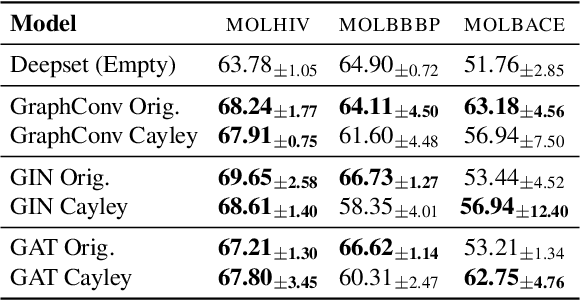
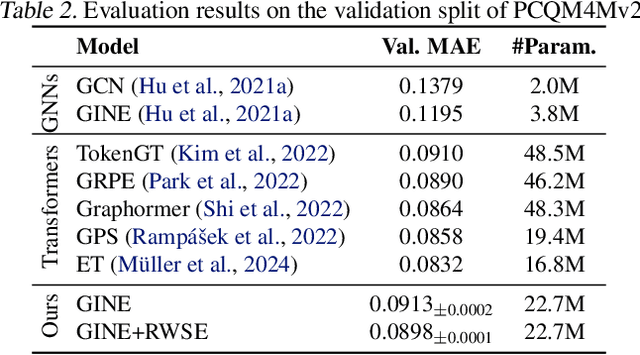
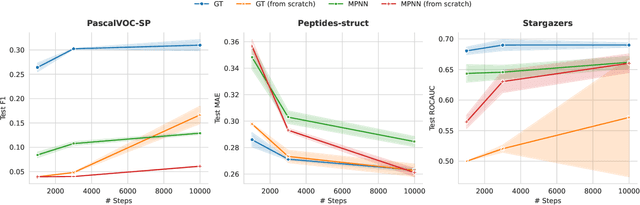
Abstract:While machine learning on graphs has demonstrated promise in drug design and molecular property prediction, significant benchmarking challenges hinder its further progress and relevance. Current benchmarking practices often lack focus on transformative, real-world applications, favoring narrow domains like two-dimensional molecular graphs over broader, impactful areas such as combinatorial optimization, relational databases, or chip design. Additionally, many benchmark datasets poorly represent the underlying data, leading to inadequate abstractions and misaligned use cases. Fragmented evaluations and an excessive focus on accuracy further exacerbate these issues, incentivizing overfitting rather than fostering generalizable insights. These limitations have prevented the development of truly useful graph foundation models. This position paper calls for a paradigm shift toward more meaningful benchmarks, rigorous evaluation protocols, and stronger collaboration with domain experts to drive impactful and reliable advances in graph learning research, unlocking the potential of graph learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge