Jonathan H. Chen
PhysicianBench: Evaluating LLM Agents in Real-World EHR Environments
May 04, 2026Abstract:We introduce PhysicianBench, a benchmark for evaluating LLM agents on physician tasks grounded in real clinical setting within electronic health record (EHR) environments. Existing medical agent benchmarks primarily focus on static knowledge recall, single-step atomic actions, or action intent without verifiable execution against the environment. As a result, they fail to capture the long-horizon, composite workflows that characterize real clinical systems. PhysicianBench comprises 100 long-horizon tasks adapted from real consultation cases between primary care and subspecialty physicians, with each task independently reviewed by a separate panel of physicians. Tasks are instantiated in an EHR environment with real patient records and accessed through the same standard APIs used by commercial EHR vendors. Tasks span 21 specialties (e.g., cardiology, endocrinology, oncology, psychiatry) and diverse workflow types (e.g., diagnosis interpretation, medication prescribing, treatment planning), requiring an average of 27 tool calls per task. Solving each task requires retrieving data across encounters, reasoning over heterogeneous clinical information, executing consequential clinical actions, and producing clinical documentation. Each task is decomposed into structured checkpoints (670 in total across the benchmark) capturing distinct stages of completion graded by task-specific scripts with execution-grounded verification. Across 13 proprietary and open-source LLM agents, the best-performing model achieves only 46% success rate (pass@1), while open-source models reach at most 19%, revealing a substantial gap between current agent capabilities and the demands of real-world clinical workflows. PhysicianBench provides a realistic and execution-grounded benchmark for measuring progress toward autonomous clinical agents.
Clinician input steers frontier AI models toward both accurate and harmful decisions
Mar 14, 2026Abstract:Large language models (LLMs) are entering clinician workflows, yet evaluations rarely measure how clinician reasoning shapes model behavior during clinical interactions. We combined 61 New England Journal of Medicine Case Records with 92 real-world clinician-AI interactions to evaluate 21 reasoning LLM variants across 8 frontier models on differential diagnosis generation and next step recommendations under three conditions: reasoning alone, after expert clinician context, and after adversarial clinician context. LLM-clinician concordance increased substantially after clinician exposure, with simulations sharing >=3 differential diagnosis items rising from 65.8% to 93.5% and >=3 next step recommendations from 20.3% to 53.8%. Expert context significantly improved correct final diagnosis inclusion across all 21 models (mean +20.4 percentage points), reflecting both reasoning improvement and passive content echoing, while adversarial context caused significant diagnostic degradation in 14 models (mean -5.4 percentage points). Multi-turn disagreement probes revealed distinct model phenotypes ranging from highly conformist to dogmatic, with adversarial arguments remaining a persistent vulnerability even for otherwise resilient models. Inference-time scaling reduced harmful echoing of clinician-introduced recommendations across WHO-defined harm severity tiers (relative reductions: 62.7% mild, 57.9% moderate, 76.3% severe, 83.5% death-tier). In GPT-4o experiments, explicit clinician uncertainty signals improved diagnostic performance after adversarial context (final diagnosis inclusion 27% to 42%) and reduced alignment with incorrect arguments by 21%. These findings establish a foundation for evaluating clinician-AI collaboration, introducing interactive metrics and mitigation strategies essential for safety and robustness.
Monitoring Deployed AI Systems in Health Care
Dec 09, 2025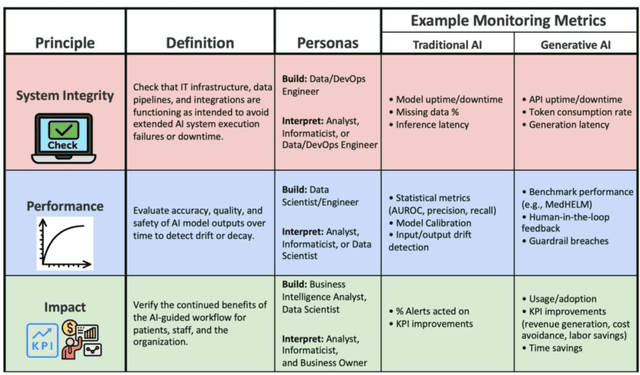
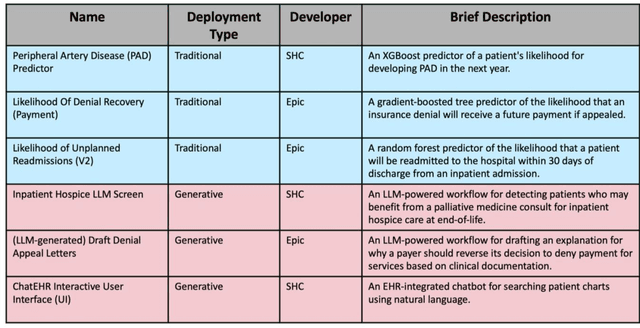
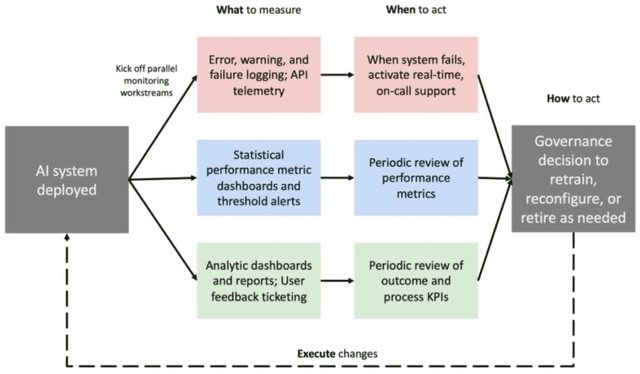
Abstract:Post-deployment monitoring of artificial intelligence (AI) systems in health care is essential to ensure their safety, quality, and sustained benefit-and to support governance decisions about which systems to update, modify, or decommission. Motivated by these needs, we developed a framework for monitoring deployed AI systems grounded in the mandate to take specific actions when they fail to behave as intended. This framework, which is now actively used at Stanford Health Care, is organized around three complementary principles: system integrity, performance, and impact. System integrity monitoring focuses on maximizing system uptime, detecting runtime errors, and identifying when changes to the surrounding IT ecosystem have unintended effects. Performance monitoring focuses on maintaining accurate system behavior in the face of changing health care practices (and thus input data) over time. Impact monitoring assesses whether a deployed system continues to have value in the form of benefit to clinicians and patients. Drawing on examples of deployed AI systems at our academic medical center, we provide practical guidance for creating monitoring plans based on these principles that specify which metrics to measure, when those metrics should be reviewed, who is responsible for acting when metrics change, and what concrete follow-up actions should be taken-for both traditional and generative AI. We also discuss challenges to implementing this framework, including the effort and cost of monitoring for health systems with limited resources and the difficulty of incorporating data-driven monitoring practices into complex organizations where conflicting priorities and definitions of success often coexist. This framework offers a practical template and starting point for health systems seeking to ensure that AI deployments remain safe and effective over time.
Retrieval-Augmented Guardrails for AI-Drafted Patient-Portal Messages: Error Taxonomy Construction and Large-Scale Evaluation
Sep 26, 2025Abstract:Asynchronous patient-clinician messaging via EHR portals is a growing source of clinician workload, prompting interest in large language models (LLMs) to assist with draft responses. However, LLM outputs may contain clinical inaccuracies, omissions, or tone mismatches, making robust evaluation essential. Our contributions are threefold: (1) we introduce a clinically grounded error ontology comprising 5 domains and 59 granular error codes, developed through inductive coding and expert adjudication; (2) we develop a retrieval-augmented evaluation pipeline (RAEC) that leverages semantically similar historical message-response pairs to improve judgment quality; and (3) we provide a two-stage prompting architecture using DSPy to enable scalable, interpretable, and hierarchical error detection. Our approach assesses the quality of drafts both in isolation and with reference to similar past message-response pairs retrieved from institutional archives. Using a two-stage DSPy pipeline, we compared baseline and reference-enhanced evaluations on over 1,500 patient messages. Retrieval context improved error identification in domains such as clinical completeness and workflow appropriateness. Human validation on 100 messages demonstrated superior agreement (concordance = 50% vs. 33%) and performance (F1 = 0.500 vs. 0.256) of context-enhanced labels vs. baseline, supporting the use of our RAEC pipeline as AI guardrails for patient messaging.
BRIDGE: Benchmarking Large Language Models for Understanding Real-world Clinical Practice Text
May 01, 2025



Abstract:Large language models (LLMs) hold great promise for medical applications and are evolving rapidly, with new models being released at an accelerated pace. However, current evaluations of LLMs in clinical contexts remain limited. Most existing benchmarks rely on medical exam-style questions or PubMed-derived text, failing to capture the complexity of real-world electronic health record (EHR) data. Others focus narrowly on specific application scenarios, limiting their generalizability across broader clinical use. To address this gap, we present BRIDGE, a comprehensive multilingual benchmark comprising 87 tasks sourced from real-world clinical data sources across nine languages. We systematically evaluated 52 state-of-the-art LLMs (including DeepSeek-R1, GPT-4o, Gemini, and Llama 4) under various inference strategies. With a total of 13,572 experiments, our results reveal substantial performance variation across model sizes, languages, natural language processing tasks, and clinical specialties. Notably, we demonstrate that open-source LLMs can achieve performance comparable to proprietary models, while medically fine-tuned LLMs based on older architectures often underperform versus updated general-purpose models. The BRIDGE and its corresponding leaderboard serve as a foundational resource and a unique reference for the development and evaluation of new LLMs in real-world clinical text understanding. The BRIDGE leaderboard: https://huggingface.co/spaces/YLab-Open/BRIDGE-Medical-Leaderboard
Deconver: A Deconvolutional Network for Medical Image Segmentation
Apr 01, 2025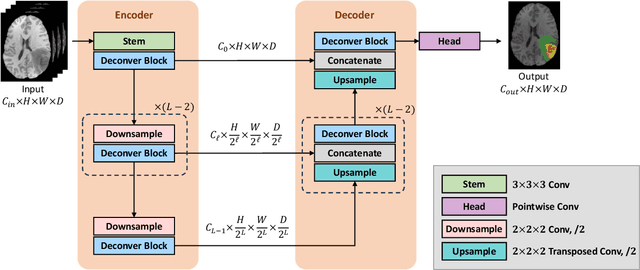
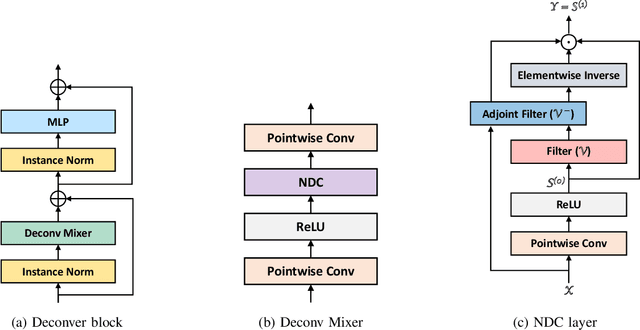
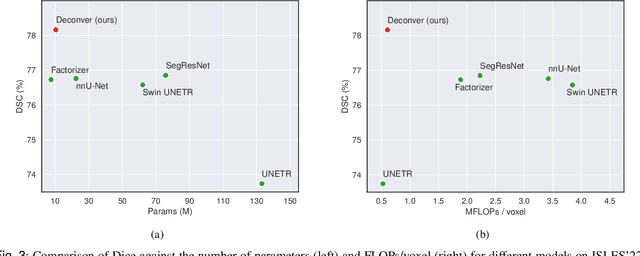
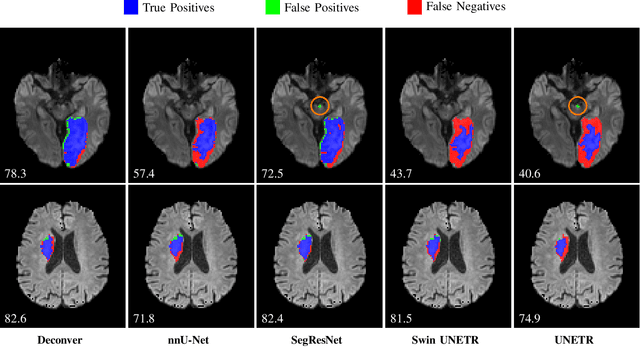
Abstract:While convolutional neural networks (CNNs) and vision transformers (ViTs) have advanced medical image segmentation, they face inherent limitations such as local receptive fields in CNNs and high computational complexity in ViTs. This paper introduces Deconver, a novel network that integrates traditional deconvolution techniques from image restoration as a core learnable component within a U-shaped architecture. Deconver replaces computationally expensive attention mechanisms with efficient nonnegative deconvolution (NDC) operations, enabling the restoration of high-frequency details while suppressing artifacts. Key innovations include a backpropagation-friendly NDC layer based on a provably monotonic update rule and a parameter-efficient design. Evaluated across four datasets (ISLES'22, BraTS'23, GlaS, FIVES) covering both 2D and 3D segmentation tasks, Deconver achieves state-of-the-art performance in Dice scores and Hausdorff distance while reducing computational costs (FLOPs) by up to 90% compared to leading baselines. By bridging traditional image restoration with deep learning, this work offers a practical solution for high-precision segmentation in resource-constrained clinical workflows. The project is available at https://github.com/pashtari/deconver.
Optimizing Large Language Models for Detecting Symptoms of Comorbid Depression or Anxiety in Chronic Diseases: Insights from Patient Messages
Mar 14, 2025



Abstract:Patients with diabetes are at increased risk of comorbid depression or anxiety, complicating their management. This study evaluated the performance of large language models (LLMs) in detecting these symptoms from secure patient messages. We applied multiple approaches, including engineered prompts, systemic persona, temperature adjustments, and zero-shot and few-shot learning, to identify the best-performing model and enhance performance. Three out of five LLMs demonstrated excellent performance (over 90% of F-1 and accuracy), with Llama 3.1 405B achieving 93% in both F-1 and accuracy using a zero-shot approach. While LLMs showed promise in binary classification and handling complex metrics like Patient Health Questionnaire-4, inconsistencies in challenging cases warrant further real-life assessment. The findings highlight the potential of LLMs to assist in timely screening and referrals, providing valuable empirical knowledge for real-world triage systems that could improve mental health care for patients with chronic diseases.
Antibiotic Resistance Microbiology Dataset (ARMD): A De-identified Resource for Studying Antimicrobial Resistance Using Electronic Health Records
Mar 08, 2025Abstract:The Antibiotic Resistance Microbiology Dataset (ARMD) is a de-identified resource derived from electronic health records (EHR) that facilitates research into antimicrobial resistance (AMR). ARMD encompasses data from adult patients, focusing on microbiological cultures, antibiotic susceptibilities, and associated clinical and demographic features. Key attributes include organism identification, susceptibility patterns for 55 antibiotics, implied susceptibility rules, and de-identified patient information. This dataset supports studies on antimicrobial stewardship, causal inference, and clinical decision-making. ARMD is designed to be reusable and interoperable, promoting collaboration and innovation in combating AMR. This paper describes the dataset's acquisition, structure, and utility while detailing its de-identification process.
MedAgentBench: Dataset for Benchmarking LLMs as Agents in Medical Applications
Jan 24, 2025Abstract:Recent large language models (LLMs) have demonstrated significant advancements, particularly in their ability to serve as agents thereby surpassing their traditional role as chatbots. These agents can leverage their planning and tool utilization capabilities to address tasks specified at a high level. However, a standardized dataset to benchmark the agent capabilities of LLMs in medical applications is currently lacking, making the evaluation of LLMs on complex tasks in interactive healthcare environments challenging. To address this gap, we introduce MedAgentBench, a broad evaluation suite designed to assess the agent capabilities of large language models within medical records contexts. MedAgentBench encompasses 100 patient-specific clinically-derived tasks from 10 categories written by human physicians, realistic profiles of 100 patients with over 700,000 data elements, a FHIR-compliant interactive environment, and an accompanying codebase. The environment uses the standard APIs and communication infrastructure used in modern EMR systems, so it can be easily migrated into live EMR systems. MedAgentBench presents an unsaturated agent-oriented benchmark that current state-of-the-art LLMs exhibit some ability to succeed at. The best model (GPT-4o) achieves a success rate of 72%. However, there is still substantial space for improvement to give the community a next direction to optimize. Furthermore, there is significant variation in performance across task categories. MedAgentBench establishes this and is publicly available at https://github.com/stanfordmlgroup/MedAgentBench , offering a valuable framework for model developers to track progress and drive continuous improvements in the agent capabilities of large language models within the medical domain.
Quantization-free Lossy Image Compression Using Integer Matrix Factorization
Aug 22, 2024



Abstract:Lossy image compression is essential for efficient transmission and storage. Traditional compression methods mainly rely on discrete cosine transform (DCT) or singular value decomposition (SVD), both of which represent image data in continuous domains and therefore necessitate carefully designed quantizers. Notably, SVD-based methods are more sensitive to quantization errors than DCT-based methods like JPEG. To address this issue, we introduce a variant of integer matrix factorization (IMF) to develop a novel quantization-free lossy image compression method. IMF provides a low-rank representation of the image data as a product of two smaller factor matrices with bounded integer elements, thereby eliminating the need for quantization. We propose an efficient, provably convergent iterative algorithm for IMF using a block coordinate descent (BCD) scheme, with subproblems having closed-form solutions. Our experiments on the Kodak and CLIC 2024 datasets demonstrate that our IMF compression method consistently outperforms JPEG at low bit rates below 0.25 bits per pixel (bpp) and remains comparable at higher bit rates. We also assessed our method's capability to preserve visual semantics by evaluating an ImageNet pre-trained classifier on compressed images. Remarkably, our method improved top-1 accuracy by over 5 percentage points compared to JPEG at bit rates under 0.25 bpp. The project is available at https://github.com/pashtari/lrf .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge