Niaz Banaei
A Multi-Phase Analysis of Blood Culture Stewardship: Machine Learning Prediction, Expert Recommendation Assessment, and LLM Automation
Apr 09, 2025Abstract:Blood cultures are often over ordered without clear justification, straining healthcare resources and contributing to inappropriate antibiotic use pressures worsened by the global shortage. In study of 135483 emergency department (ED) blood culture orders, we developed machine learning (ML) models to predict the risk of bacteremia using structured electronic health record (EHR) data and provider notes via a large language model (LLM). The structured models AUC improved from 0.76 to 0.79 with note embeddings and reached 0.81 with added diagnosis codes. Compared to an expert recommendation framework applied by human reviewers and an LLM-based pipeline, our ML approach offered higher specificity without compromising sensitivity. The recommendation framework achieved sensitivity 86%, specificity 57%, while the LLM maintained high sensitivity (96%) but over classified negatives, reducing specificity (16%). These findings demonstrate that ML models integrating structured and unstructured data can outperform consensus recommendations, enhancing diagnostic stewardship beyond existing standards of care.
Antibiotic Resistance Microbiology Dataset (ARMD): A De-identified Resource for Studying Antimicrobial Resistance Using Electronic Health Records
Mar 08, 2025Abstract:The Antibiotic Resistance Microbiology Dataset (ARMD) is a de-identified resource derived from electronic health records (EHR) that facilitates research into antimicrobial resistance (AMR). ARMD encompasses data from adult patients, focusing on microbiological cultures, antibiotic susceptibilities, and associated clinical and demographic features. Key attributes include organism identification, susceptibility patterns for 55 antibiotics, implied susceptibility rules, and de-identified patient information. This dataset supports studies on antimicrobial stewardship, causal inference, and clinical decision-making. ARMD is designed to be reusable and interoperable, promoting collaboration and innovation in combating AMR. This paper describes the dataset's acquisition, structure, and utility while detailing its de-identification process.
Rapid identification of pathogenic bacteria using Raman spectroscopy and deep learning
Jan 23, 2019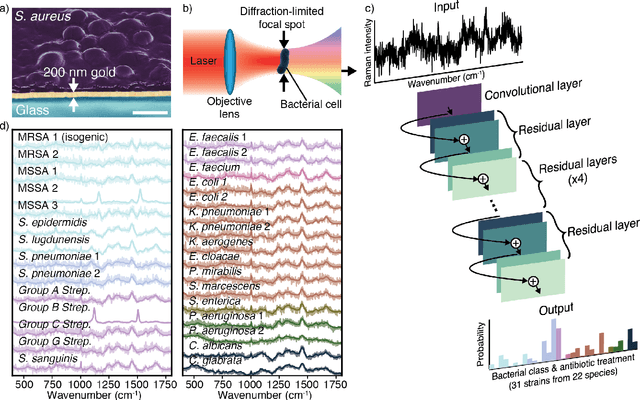


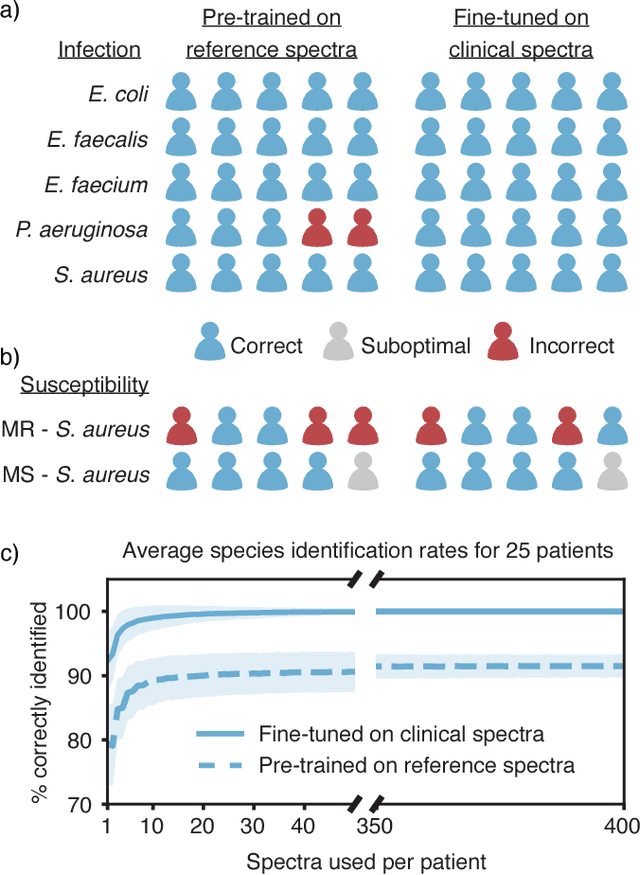
Abstract:Rapid identification of bacteria is essential to prevent the spread of infectious disease, help combat antimicrobial resistance, and improve patient outcomes. Raman optical spectroscopy promises to combine bacterial detection, identification, and antibiotic susceptibility testing in a single step. However, achieving clinically relevant speeds and accuracies remains challenging due to the weak Raman signal from bacterial cells and the large number of bacterial species and phenotypes. By amassing the largest known dataset of bacterial Raman spectra, we are able to apply state-of-the-art deep learning approaches to identify 30 of the most common bacterial pathogens from noisy Raman spectra, achieving antibiotic treatment identification accuracies of 99.0$\pm$0.1%. This novel approach distinguishes between methicillin-resistant and -susceptible isolates of Staphylococcus aureus (MRSA and MSSA) as well as a pair of isogenic MRSA and MSSA that are genetically identical apart from deletion of the mecA resistance gene, indicating the potential for culture-free detection of antibiotic resistance. Results from initial clinical validation are promising: using just 10 bacterial spectra from each of 25 isolates, we achieve 99.0$\pm$1.9% species identification accuracy. Our combined Raman-deep learning system represents an important proof-of-concept for rapid, culture-free identification of bacterial isolates and antibiotic resistance and could be readily extended for diagnostics on blood, urine, and sputum.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge