Zhihua Wang
LoViF 2026 Challenge on Human-oriented Semantic Image Quality Assessment: Methods and Results
Apr 13, 2026Abstract:This paper reviews the LoViF 2026 Challenge on Human-oriented Semantic Image Quality Assessment. This challenge aims to raise a new direction, i.e., how to evaluate the loss of semantic information from the human perspective, intending to promote the development of some new directions, like semantic coding, processing, and semantic-oriented optimization, etc. Unlike existing datasets of quality assessment, we form a dataset of human-oriented semantic quality assessment, termed the SeIQA dataset. This dataset is divided into three parts for this competition: (i) training data: 510 pairs of degraded images and their corresponding ground truth references; (ii) validation data: 80 pairs of degraded images and their corresponding ground-truth references; (iii) testing data: 160 pairs of degraded images and their corresponding ground-truth references. The primary objective of this challenge is to establish a new and powerful benchmark for human-oriented semantic image quality assessment. There are a total of 58 teams registered in this competition, and 6 teams submitted valid solutions and fact sheets for the final testing phase. These submissions achieved state-of-the-art (SOTA) performance on the SeIQA dataset.
PhysInOne: Visual Physics Learning and Reasoning in One Suite
Apr 10, 2026Abstract:We present PhysInOne, a large-scale synthetic dataset addressing the critical scarcity of physically-grounded training data for AI systems. Unlike existing datasets limited to merely hundreds or thousands of examples, PhysInOne provides 2 million videos across 153,810 dynamic 3D scenes, covering 71 basic physical phenomena in mechanics, optics, fluid dynamics, and magnetism. Distinct from previous works, our scenes feature multiobject interactions against complex backgrounds, with comprehensive ground-truth annotations including 3D geometry, semantics, dynamic motion, physical properties, and text descriptions. We demonstrate PhysInOne's efficacy across four emerging applications: physics-aware video generation, long-/short-term future frame prediction, physical property estimation, and motion transfer. Experiments show that fine-tuning foundation models on PhysInOne significantly enhances physical plausibility, while also exposing critical gaps in modeling complex physical dynamics and estimating intrinsic properties. As the largest dataset of its kind, orders of magnitude beyond prior works, PhysInOne establishes a new benchmark for advancing physics-grounded world models in generation, simulation, and embodied AI.
Development and multi-center evaluation of domain-adapted speech recognition for human-AI teaming in real-world gastrointestinal endoscopy
Apr 02, 2026Abstract:Automatic speech recognition (ASR) is a critical interface for human-AI interaction in gastrointestinal endoscopy, yet its reliability in real-world clinical settings is limited by domain-specific terminology and complex acoustic conditions. Here, we present EndoASR, a domain-adapted ASR system designed for real-time deployment in endoscopic workflows. We develop a two-stage adaptation strategy based on synthetic endoscopy reports, targeting domain-specific language modeling and noise robustness. In retrospective evaluation across six endoscopists, EndoASR substantially improves both transcription accuracy and clinical usability, reducing character error rate (CER) from 20.52% to 14.14% and increasing medical term accuracy (Med ACC) from 54.30% to 87.59%. In a prospective multi-center study spanning five independent endoscopy centers, EndoASR demonstrates consistent generalization under heterogeneous real-world conditions. Compared with the baseline Paraformer model, CER is reduced from 16.20% to 14.97%, while Med ACC is improved from 61.63% to 84.16%, confirming its robustness in practical deployment scenarios. Notably, EndoASR achieves a real-time factor (RTF) of 0.005, significantly faster than Whisper-large-v3 (RTF 0.055), while maintaining a compact model size of 220M parameters, enabling efficient edge deployment. Furthermore, integration with large language models demonstrates that improved ASR quality directly enhances downstream structured information extraction and clinician-AI interaction. These results demonstrate that domain-adapted ASR can serve as a reliable interface for human-AI teaming in gastrointestinal endoscopy, with consistent performance validated across multi-center real-world clinical settings.
MDS-VQA: Model-Informed Data Selection for Video Quality Assessment
Mar 12, 2026Abstract:Learning-based video quality assessment (VQA) has advanced rapidly, yet progress is increasingly constrained by a disconnect between model design and dataset curation. Model-centric approaches often iterate on fixed benchmarks, while data-centric efforts collect new human labels without systematically targeting the weaknesses of existing VQA models. Here, we describe MDS-VQA, a model-informed data selection mechanism for curating unlabeled videos that are both difficult for the base VQA model and diverse in content. Difficulty is estimated by a failure predictor trained with a ranking objective, and diversity is measured using deep semantic video features, with a greedy procedure balancing the two under a constrained labeling budget. Experiments across multiple VQA datasets and models demonstrate that MDS-VQA identifies diverse, challenging samples that are particularly informative for active fine-tuning. With only a 5% selected subset per target domain, the fine-tuned model improves mean SRCC from 0.651 to 0.722 and achieves the top gMAD rank, indicating strong adaptation and generalization.
One-shot synthesis of rare gastrointestinal lesions improves diagnostic accuracy and clinical training
Dec 30, 2025Abstract:Rare gastrointestinal lesions are infrequently encountered in routine endoscopy, restricting the data available for developing reliable artificial intelligence (AI) models and training novice clinicians. Here we present EndoRare, a one-shot, retraining-free generative framework that synthesizes diverse, high-fidelity lesion exemplars from a single reference image. By leveraging language-guided concept disentanglement, EndoRare separates pathognomonic lesion features from non-diagnostic attributes, encoding the former into a learnable prototype embedding while varying the latter to ensure diversity. We validated the framework across four rare pathologies (calcifying fibrous tumor, juvenile polyposis syndrome, familial adenomatous polyposis, and Peutz-Jeghers syndrome). Synthetic images were judged clinically plausible by experts and, when used for data augmentation, significantly enhanced downstream AI classifiers, improving the true positive rate at low false-positive rates. Crucially, a blinded reader study demonstrated that novice endoscopists exposed to EndoRare-generated cases achieved a 0.400 increase in recall and a 0.267 increase in precision. These results establish a practical, data-efficient pathway to bridge the rare-disease gap in both computer-aided diagnostics and clinical education.
RoboMatch: A Mobile-Manipulation Teleoperation Platform with Auto-Matching Network Architecture for Long-Horizon Manipulation
Sep 10, 2025



Abstract:This paper presents RoboMatch, a novel unified teleoperation platform for mobile manipulation with an auto-matching network architecture, designed to tackle long-horizon tasks in dynamic environments. Our system enhances teleoperation performance, data collection efficiency, task accuracy, and operational stability. The core of RoboMatch is a cockpit-style control interface that enables synchronous operation of the mobile base and dual arms, significantly improving control precision and data collection. Moreover, we introduce the Proprioceptive-Visual Enhanced Diffusion Policy (PVE-DP), which leverages Discrete Wavelet Transform (DWT) for multi-scale visual feature extraction and integrates high-precision IMUs at the end-effector to enrich proprioceptive feedback, substantially boosting fine manipulation performance. Furthermore, we propose an Auto-Matching Network (AMN) architecture that decomposes long-horizon tasks into logical sequences and dynamically assigns lightweight pre-trained models for distributed inference. Experimental results demonstrate that our approach improves data collection efficiency by over 20%, increases task success rates by 20-30% with PVE-DP, and enhances long-horizon inference performance by approximately 40% with AMN, offering a robust solution for complex manipulation tasks.
EndoFinder: Online Lesion Retrieval for Explainable Colorectal Polyp Diagnosis Leveraging Latent Scene Representations
Jul 23, 2025Abstract:Colorectal cancer (CRC) remains a leading cause of cancer-related mortality, underscoring the importance of timely polyp detection and diagnosis. While deep learning models have improved optical-assisted diagnostics, they often demand extensive labeled datasets and yield "black-box" outputs with limited interpretability. In this paper, we propose EndoFinder, an online polyp retrieval framework that leverages multi-view scene representations for explainable and scalable CRC diagnosis. First, we develop a Polyp-aware Image Encoder by combining contrastive learning and a reconstruction task, guided by polyp segmentation masks. This self-supervised approach captures robust features without relying on large-scale annotated data. Next, we treat each polyp as a three-dimensional "scene" and introduce a Scene Representation Transformer, which fuses multiple views of the polyp into a single latent representation. By discretizing this representation through a hashing layer, EndoFinder enables real-time retrieval from a compiled database of historical polyp cases, where diagnostic information serves as interpretable references for new queries. We evaluate EndoFinder on both public and newly collected polyp datasets for re-identification and pathology classification. Results show that EndoFinder outperforms existing methods in accuracy while providing transparent, retrieval-based insights for clinical decision-making. By contributing a novel dataset and a scalable, explainable framework, our work addresses key challenges in polyp diagnosis and offers a promising direction for more efficient AI-driven colonoscopy workflows. The source code is available at https://github.com/ku262/EndoFinder-Scene.
Endo-CLIP: Progressive Self-Supervised Pre-training on Raw Colonoscopy Records
May 14, 2025



Abstract:Pre-training on image-text colonoscopy records offers substantial potential for improving endoscopic image analysis, but faces challenges including non-informative background images, complex medical terminology, and ambiguous multi-lesion descriptions. We introduce Endo-CLIP, a novel self-supervised framework that enhances Contrastive Language-Image Pre-training (CLIP) for this domain. Endo-CLIP's three-stage framework--cleansing, attunement, and unification--addresses these challenges by (1) removing background frames, (2) leveraging large language models to extract clinical attributes for fine-grained contrastive learning, and (3) employing patient-level cross-attention to resolve multi-polyp ambiguities. Extensive experiments demonstrate that Endo-CLIP significantly outperforms state-of-the-art pre-training methods in zero-shot and few-shot polyp detection and classification, paving the way for more accurate and clinically relevant endoscopic analysis.
Towards Realistic Low-Light Image Enhancement via ISP Driven Data Modeling
Apr 16, 2025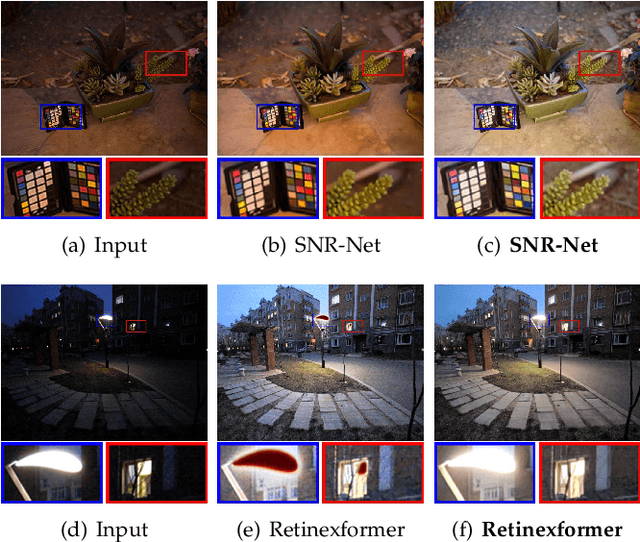
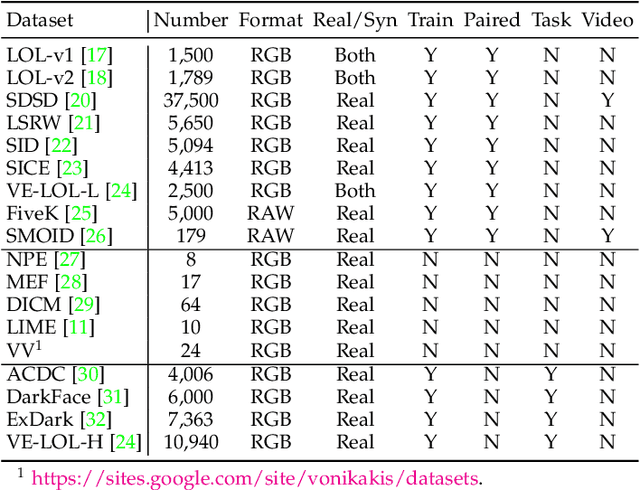
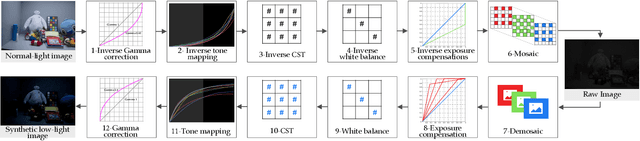
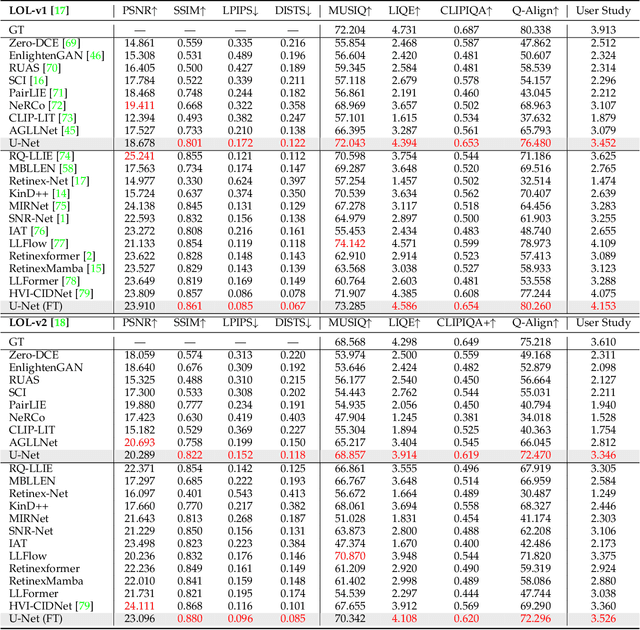
Abstract:Deep neural networks (DNNs) have recently become the leading method for low-light image enhancement (LLIE). However, despite significant progress, their outputs may still exhibit issues such as amplified noise, incorrect white balance, or unnatural enhancements when deployed in real world applications. A key challenge is the lack of diverse, large scale training data that captures the complexities of low-light conditions and imaging pipelines. In this paper, we propose a novel image signal processing (ISP) driven data synthesis pipeline that addresses these challenges by generating unlimited paired training data. Specifically, our pipeline begins with easily collected high-quality normal-light images, which are first unprocessed into the RAW format using a reverse ISP. We then synthesize low-light degradations directly in the RAW domain. The resulting data is subsequently processed through a series of ISP stages, including white balance adjustment, color space conversion, tone mapping, and gamma correction, with controlled variations introduced at each stage. This broadens the degradation space and enhances the diversity of the training data, enabling the generated data to capture a wide range of degradations and the complexities inherent in the ISP pipeline. To demonstrate the effectiveness of our synthetic pipeline, we conduct extensive experiments using a vanilla UNet model consisting solely of convolutional layers, group normalization, GeLU activation, and convolutional block attention modules (CBAMs). Extensive testing across multiple datasets reveals that the vanilla UNet model trained with our data synthesis pipeline delivers high fidelity, visually appealing enhancement results, surpassing state-of-the-art (SOTA) methods both quantitatively and qualitatively.
Robust Polyp Detection and Diagnosis through Compositional Prompt-Guided Diffusion Models
Feb 25, 2025Abstract:Colorectal cancer (CRC) is a significant global health concern, and early detection through screening plays a critical role in reducing mortality. While deep learning models have shown promise in improving polyp detection, classification, and segmentation, their generalization across diverse clinical environments, particularly with out-of-distribution (OOD) data, remains a challenge. Multi-center datasets like PolypGen have been developed to address these issues, but their collection is costly and time-consuming. Traditional data augmentation techniques provide limited variability, failing to capture the complexity of medical images. Diffusion models have emerged as a promising solution for generating synthetic polyp images, but the image generation process in current models mainly relies on segmentation masks as the condition, limiting their ability to capture the full clinical context. To overcome these limitations, we propose a Progressive Spectrum Diffusion Model (PSDM) that integrates diverse clinical annotations-such as segmentation masks, bounding boxes, and colonoscopy reports-by transforming them into compositional prompts. These prompts are organized into coarse and fine components, allowing the model to capture both broad spatial structures and fine details, generating clinically accurate synthetic images. By augmenting training data with PSDM-generated samples, our model significantly improves polyp detection, classification, and segmentation. For instance, on the PolypGen dataset, PSDM increases the F1 score by 2.12% and the mean average precision by 3.09%, demonstrating superior performance in OOD scenarios and enhanced generalization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge