Yunya Song
Multimodal Clinical Reasoning through Knowledge-augmented Rationale Generation
Nov 12, 2024Abstract:Clinical rationales play a pivotal role in accurate disease diagnosis; however, many models predominantly use discriminative methods and overlook the importance of generating supportive rationales. Rationale distillation is a process that transfers knowledge from large language models (LLMs) to smaller language models (SLMs), thereby enhancing the latter's ability to break down complex tasks. Despite its benefits, rationale distillation alone is inadequate for addressing domain knowledge limitations in tasks requiring specialized expertise, such as disease diagnosis. Effectively embedding domain knowledge in SLMs poses a significant challenge. While current LLMs are primarily geared toward processing textual data, multimodal LLMs that incorporate time series data, especially electronic health records (EHRs), are still evolving. To tackle these limitations, we introduce ClinRaGen, an SLM optimized for multimodal rationale generation in disease diagnosis. ClinRaGen incorporates a unique knowledge-augmented attention mechanism to merge domain knowledge with time series EHR data, utilizing a stepwise rationale distillation strategy to produce both textual and time series-based clinical rationales. Our evaluations show that ClinRaGen markedly improves the SLM's capability to interpret multimodal EHR data and generate accurate clinical rationales, supporting more reliable disease diagnosis, advancing LLM applications in healthcare, and narrowing the performance divide between LLMs and SLMs.
Improved Target-specific Stance Detection on Social Media Platforms by Delving into Conversation Threads
Nov 06, 2022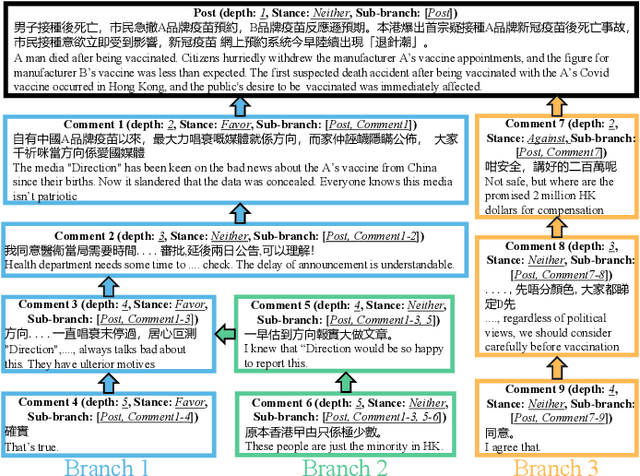
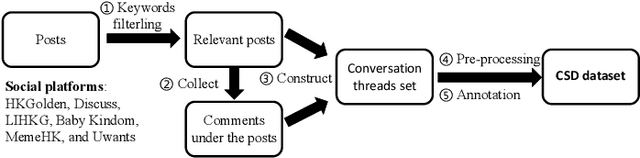
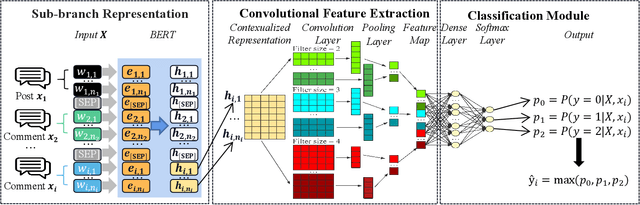
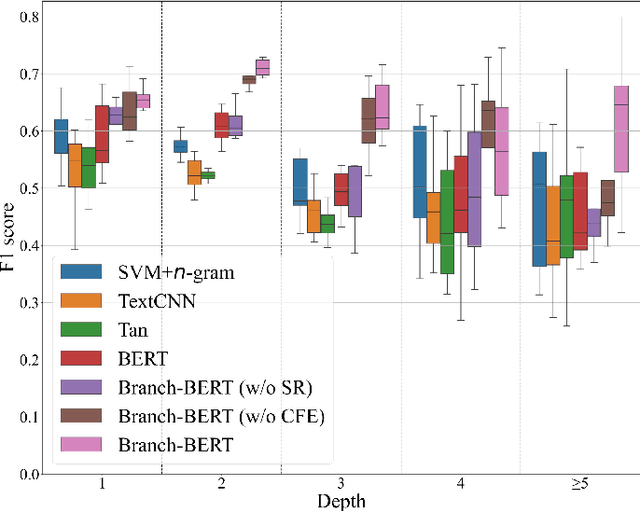
Abstract:Target-specific stance detection on social media, which aims at classifying a textual data instance such as a post or a comment into a stance class of a target issue, has become an emerging opinion mining paradigm of importance. An example application would be to overcome vaccine hesitancy in combating the coronavirus pandemic. However, existing stance detection strategies rely merely on the individual instances which cannot always capture the expressed stance of a given target. In response, we address a new task called conversational stance detection which is to infer the stance towards a given target (e.g., COVID-19 vaccination) when given a data instance and its corresponding conversation thread. To tackle the task, we first propose a benchmarking conversational stance detection (CSD) dataset with annotations of stances and the structures of conversation threads among the instances based on six major social media platforms in Hong Kong. To infer the desired stances from both data instances and conversation threads, we propose a model called Branch-BERT that incorporates contextual information in conversation threads. Extensive experiments on our CSD dataset show that our proposed model outperforms all the baseline models that do not make use of contextual information. Specifically, it improves the F1 score by 10.3% compared with the state-of-the-art method in the SemEval-2016 Task 6 competition. This shows the potential of incorporating rich contextual information on detecting target-specific stances on social media platforms and implies a more practical way to construct future stance detection tasks.
Label-dependent and event-guided interpretable disease risk prediction using EHRs
Jan 18, 2022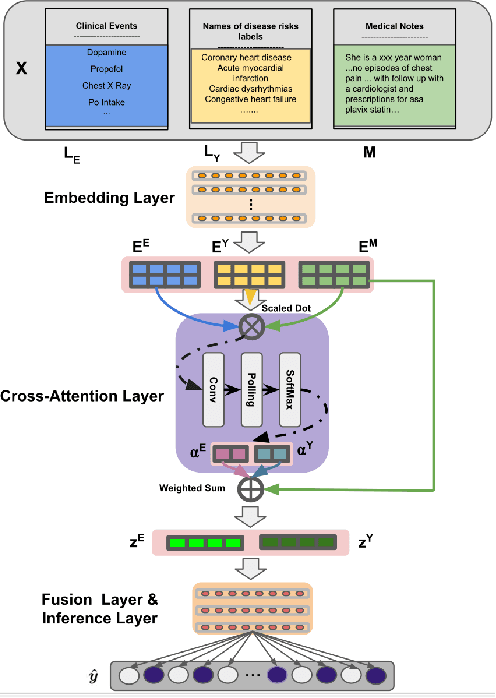
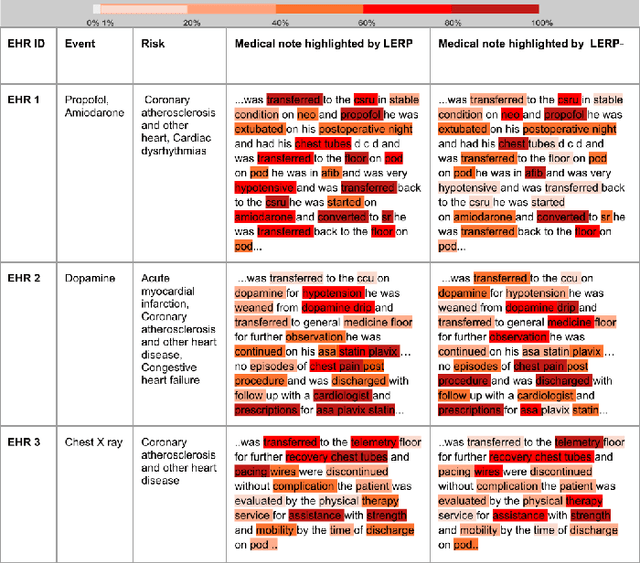
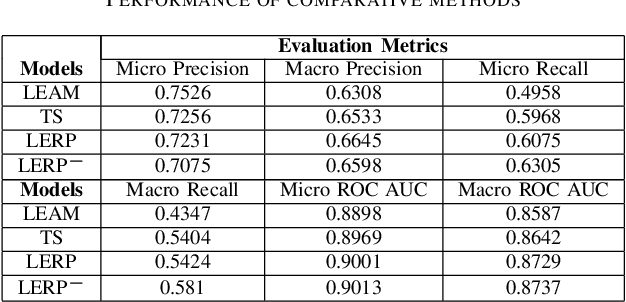
Abstract:Electronic health records (EHRs) contain patients' heterogeneous data that are collected from medical providers involved in the patient's care, including medical notes, clinical events, laboratory test results, symptoms, and diagnoses. In the field of modern healthcare, predicting whether patients would experience any risks based on their EHRs has emerged as a promising research area, in which artificial intelligence (AI) plays a key role. To make AI models practically applicable, it is required that the prediction results should be both accurate and interpretable. To achieve this goal, this paper proposed a label-dependent and event-guided risk prediction model (LERP) to predict the presence of multiple disease risks by mainly extracting information from unstructured medical notes. Our model is featured in the following aspects. First, we adopt a label-dependent mechanism that gives greater attention to words from medical notes that are semantically similar to the names of risk labels. Secondly, as the clinical events (e.g., treatments and drugs) can also indicate the health status of patients, our model utilizes the information from events and uses them to generate an event-guided representation of medical notes. Thirdly, both label-dependent and event-guided representations are integrated to make a robust prediction, in which the interpretability is enabled by the attention weights over words from medical notes. To demonstrate the applicability of the proposed method, we apply it to the MIMIC-III dataset, which contains real-world EHRs collected from hospitals. Our method is evaluated in both quantitative and qualitative ways.
Label Dependent Attention Model for Disease Risk Prediction Using Multimodal Electronic Health Records
Jan 18, 2022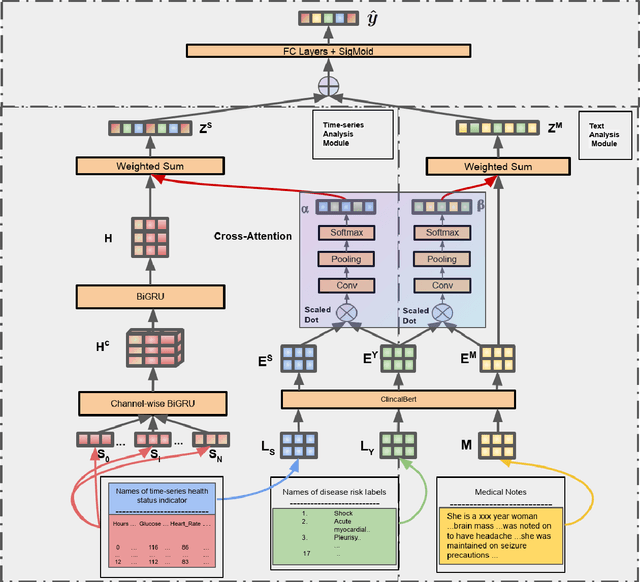
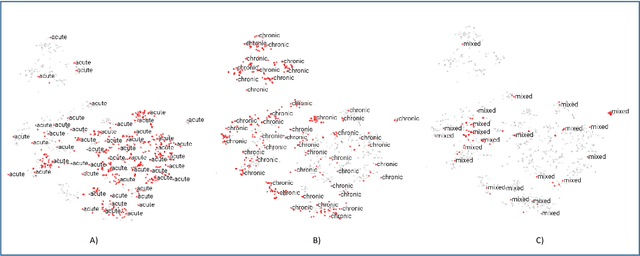
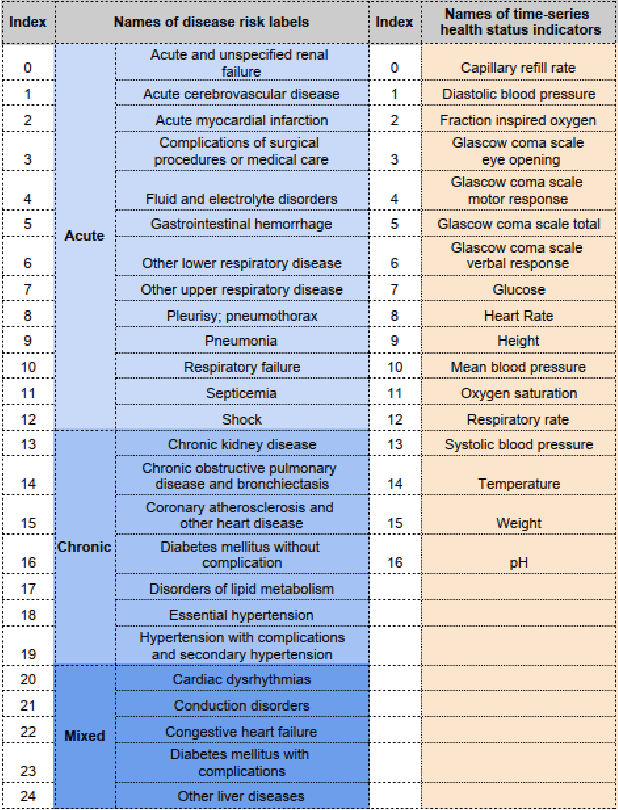

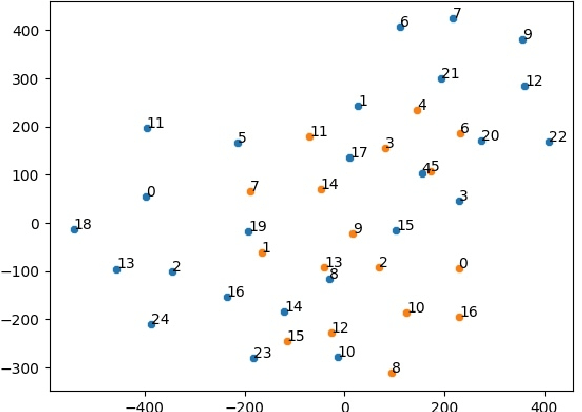
Abstract:Disease risk prediction has attracted increasing attention in the field of modern healthcare, especially with the latest advances in artificial intelligence (AI). Electronic health records (EHRs), which contain heterogeneous patient information, are widely used in disease risk prediction tasks. One challenge of applying AI models for risk prediction lies in generating interpretable evidence to support the prediction results while retaining the prediction ability. In order to address this problem, we propose the method of jointly embedding words and labels whereby attention modules learn the weights of words from medical notes according to their relevance to the names of risk prediction labels. This approach boosts interpretability by employing an attention mechanism and including the names of prediction tasks in the model. However, its application is only limited to the handling of textual inputs such as medical notes. In this paper, we propose a label dependent attention model LDAM to 1) improve the interpretability by exploiting Clinical-BERT (a biomedical language model pre-trained on a large clinical corpus) to encode biomedically meaningful features and labels jointly; 2) extend the idea of joint embedding to the processing of time-series data, and develop a multi-modal learning framework for integrating heterogeneous information from medical notes and time-series health status indicators. To demonstrate our method, we apply LDAM to the MIMIC-III dataset to predict different disease risks. We evaluate our method both quantitatively and qualitatively. Specifically, the predictive power of LDAM will be shown, and case studies will be carried out to illustrate its interpretability.
A Synthetic Approach for Recommendation: Combining Ratings, Social Relations, and Reviews
Jan 11, 2016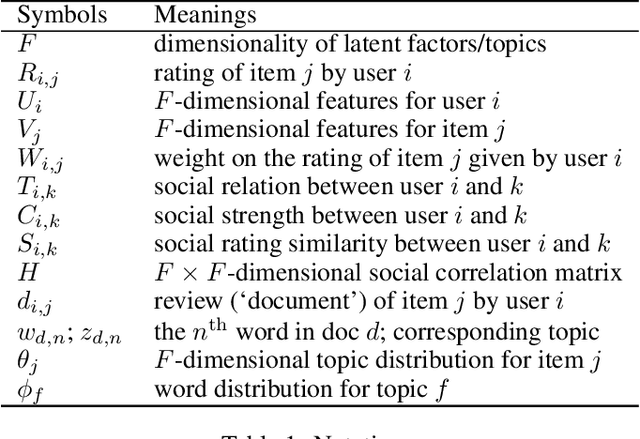
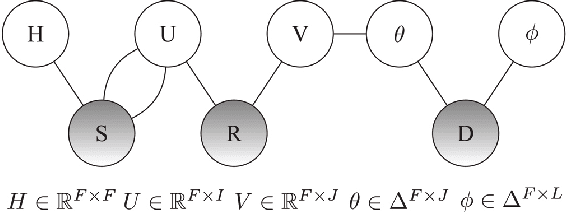
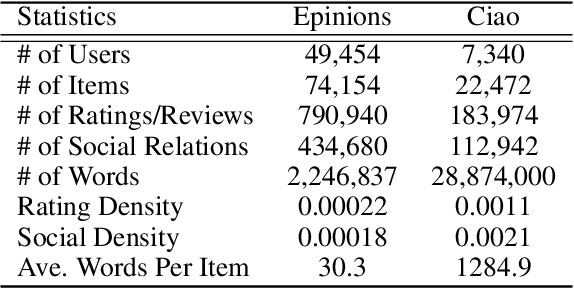
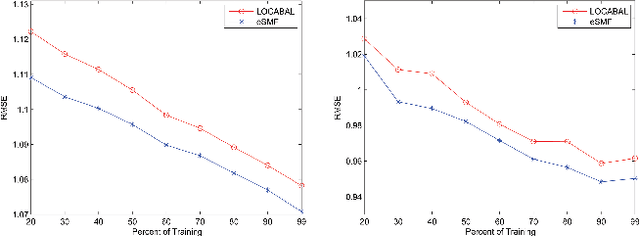
Abstract:Recommender systems (RSs) provide an effective way of alleviating the information overload problem by selecting personalized choices. Online social networks and user-generated content provide diverse sources for recommendation beyond ratings, which present opportunities as well as challenges for traditional RSs. Although social matrix factorization (Social MF) can integrate ratings with social relations and topic matrix factorization can integrate ratings with item reviews, both of them ignore some useful information. In this paper, we investigate the effective data fusion by combining the two approaches, in two steps. First, we extend Social MF to exploit the graph structure of neighbors. Second, we propose a novel framework MR3 to jointly model these three types of information effectively for rating prediction by aligning latent factors and hidden topics. We achieve more accurate rating prediction on two real-life datasets. Furthermore, we measure the contribution of each data source to the proposed framework.
* 7 pages, 8 figures
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge