Xueqing Peng
Ebisu: Benchmarking Large Language Models in Japanese Finance
Feb 01, 2026Abstract:Japanese finance combines agglutinative, head-final linguistic structure, mixed writing systems, and high-context communication norms that rely on indirect expression and implicit commitment, posing a substantial challenge for LLMs. We introduce Ebisu, a benchmark for native Japanese financial language understanding, comprising two linguistically and culturally grounded, expert-annotated tasks: JF-ICR, which evaluates implicit commitment and refusal recognition in investor-facing Q&A, and JF-TE, which assesses hierarchical extraction and ranking of nested financial terminology from professional disclosures. We evaluate a diverse set of open-source and proprietary LLMs spanning general-purpose, Japanese-adapted, and financial models. Results show that even state-of-the-art systems struggle on both tasks. While increased model scale yields limited improvements, language- and domain-specific adaptation does not reliably improve performance, leaving substantial gaps unresolved. Ebisu provides a focused benchmark for advancing linguistically and culturally grounded financial NLP. All datasets and evaluation scripts are publicly released.
MedViz: An Agent-based, Visual-guided Research Assistant for Navigating Biomedical Literature
Jan 28, 2026Abstract:Biomedical researchers face increasing challenges in navigating millions of publications in diverse domains. Traditional search engines typically return articles as ranked text lists, offering little support for global exploration or in-depth analysis. Although recent advances in generative AI and large language models have shown promise in tasks such as summarization, extraction, and question answering, their dialog-based implementations are poorly integrated with literature search workflows. To address this gap, we introduce MedViz, a visual analytics system that integrates multiple AI agents with interactive visualization to support the exploration of the large-scale biomedical literature. MedViz combines a semantic map of millions of articles with agent-driven functions for querying, summarizing, and hypothesis generation, allowing researchers to iteratively refine questions, identify trends, and uncover hidden connections. By bridging intelligent agents with interactive visualization, MedViz transforms biomedical literature search into a dynamic, exploratory process that accelerates knowledge discovery.
FinCriticalED: A Visual Benchmark for Financial Fact-Level OCR Evaluation
Nov 19, 2025Abstract:We introduce FinCriticalED (Financial Critical Error Detection), a visual benchmark for evaluating OCR and vision language models on financial documents at the fact level. Financial documents contain visually dense and table heavy layouts where numerical and temporal information is tightly coupled with structure. In high stakes settings, small OCR mistakes such as sign inversion or shifted dates can lead to materially different interpretations, while traditional OCR metrics like ROUGE and edit distance capture only surface level text similarity. \ficriticaled provides 500 image-HTML pairs with expert annotated financial facts covering over seven hundred numerical and temporal facts. It introduces three key contributions. First, it establishes the first fact level evaluation benchmark for financial document understanding, shifting evaluation from lexical overlap to domain critical factual correctness. Second, all annotations are created and verified by financial experts with strict quality control over signs, magnitudes, and temporal expressions. Third, we develop an LLM-as-Judge evaluation pipeline that performs structured fact extraction and contextual verification for visually complex financial documents. We benchmark OCR systems, open source vision language models, and proprietary models on FinCriticalED. Results show that although the strongest proprietary models achieve the highest factual accuracy, substantial errors remain in visually intricate numerical and temporal contexts. Through quantitative evaluation and expert case studies, FinCriticalED provides a rigorous foundation for advancing visual factual precision in financial and other precision critical domains.
FinTagging: An LLM-ready Benchmark for Extracting and Structuring Financial Information
May 27, 2025Abstract:We introduce FinTagging, the first full-scope, table-aware XBRL benchmark designed to evaluate the structured information extraction and semantic alignment capabilities of large language models (LLMs) in the context of XBRL-based financial reporting. Unlike prior benchmarks that oversimplify XBRL tagging as flat multi-class classification and focus solely on narrative text, FinTagging decomposes the XBRL tagging problem into two subtasks: FinNI for financial entity extraction and FinCL for taxonomy-driven concept alignment. It requires models to jointly extract facts and align them with the full 10k+ US-GAAP taxonomy across both unstructured text and structured tables, enabling realistic, fine-grained evaluation. We assess a diverse set of LLMs under zero-shot settings, systematically analyzing their performance on both subtasks and overall tagging accuracy. Our results reveal that, while LLMs demonstrate strong generalization in information extraction, they struggle with fine-grained concept alignment, particularly in disambiguating closely related taxonomy entries. These findings highlight the limitations of existing LLMs in fully automating XBRL tagging and underscore the need for improved semantic reasoning and schema-aware modeling to meet the demands of accurate financial disclosure. Code is available at our GitHub repository and data is at our Hugging Face repository.
Plutus: Benchmarking Large Language Models in Low-Resource Greek Finance
Feb 26, 2025Abstract:Despite Greece's pivotal role in the global economy, large language models (LLMs) remain underexplored for Greek financial context due to the linguistic complexity of Greek and the scarcity of domain-specific datasets. Previous efforts in multilingual financial natural language processing (NLP) have exposed considerable performance disparities, yet no dedicated Greek financial benchmarks or Greek-specific financial LLMs have been developed until now. To bridge this gap, we introduce Plutus-ben, the first Greek Financial Evaluation Benchmark, and Plutus-8B, the pioneering Greek Financial LLM, fine-tuned with Greek domain-specific data. Plutus-ben addresses five core financial NLP tasks in Greek: numeric and textual named entity recognition, question answering, abstractive summarization, and topic classification, thereby facilitating systematic and reproducible LLM assessments. To underpin these tasks, we present three novel, high-quality Greek financial datasets, thoroughly annotated by expert native Greek speakers, augmented by two existing resources. Our comprehensive evaluation of 22 LLMs on Plutus-ben reveals that Greek financial NLP remains challenging due to linguistic complexity, domain-specific terminology, and financial reasoning gaps. These findings underscore the limitations of cross-lingual transfer, the necessity for financial expertise in Greek-trained models, and the challenges of adapting financial LLMs to Greek text. We release Plutus-ben, Plutus-8B, and all associated datasets publicly to promote reproducible research and advance Greek financial NLP, fostering broader multilingual inclusivity in finance.
FLAG-Trader: Fusion LLM-Agent with Gradient-based Reinforcement Learning for Financial Trading
Feb 19, 2025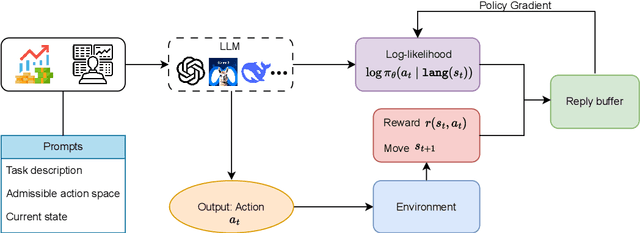
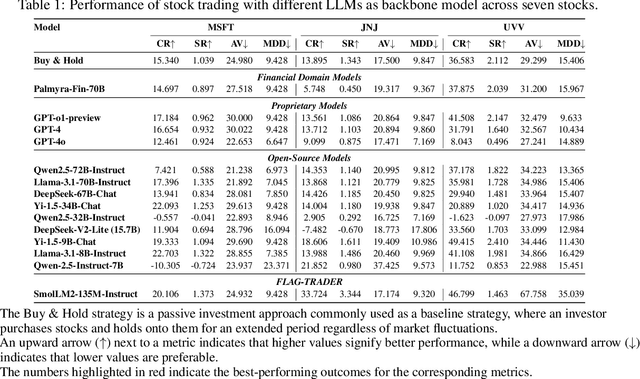
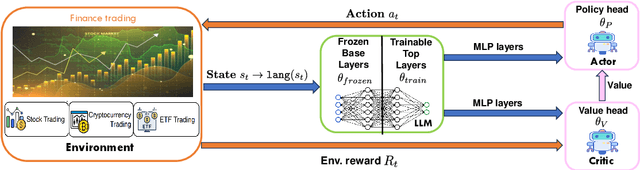
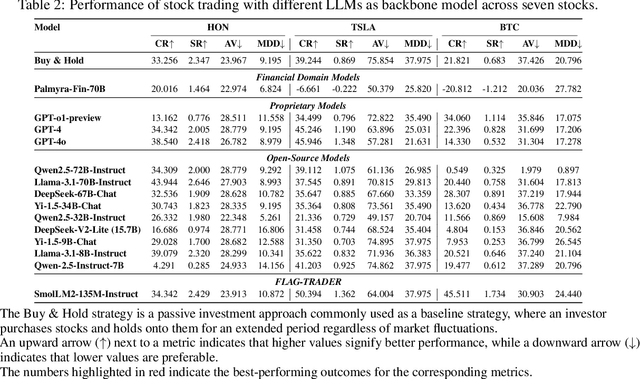
Abstract:Large language models (LLMs) fine-tuned on multimodal financial data have demonstrated impressive reasoning capabilities in various financial tasks. However, they often struggle with multi-step, goal-oriented scenarios in interactive financial markets, such as trading, where complex agentic approaches are required to improve decision-making. To address this, we propose \textsc{FLAG-Trader}, a unified architecture integrating linguistic processing (via LLMs) with gradient-driven reinforcement learning (RL) policy optimization, in which a partially fine-tuned LLM acts as the policy network, leveraging pre-trained knowledge while adapting to the financial domain through parameter-efficient fine-tuning. Through policy gradient optimization driven by trading rewards, our framework not only enhances LLM performance in trading but also improves results on other financial-domain tasks. We present extensive empirical evidence to validate these enhancements.
Fino1: On the Transferability of Reasoning Enhanced LLMs to Finance
Feb 12, 2025Abstract:Recent advancements in large language models (LLMs) have shown strong general reasoning abilities, yet their effectiveness in financial reasoning remains underexplored. In this study, we comprehensively evaluate 16 powerful reasoning and general LLMs on three complex financial tasks involving financial text, tabular data, and equations, assessing numerical reasoning, tabular interpretation, financial terminology comprehension, long-context processing, and equation-based problem solving. Our results show that while better datasets and pretraining improve financial reasoning, general enhancements like CoT fine-tuning do not always yield consistent gains. Moreover, all reasoning strategies face challenges in improving performance on long-context and multi-table tasks. To address these limitations, we develop a financial reasoning-enhanced model based on Llama-3.1-8B-Instruct, by CoT fine-tuning and reinforcement learning with domain-specific reasoning paths. Even with simple fine-tuning with one financial dataset, our model achieves a consistent 10% performance improvement across tasks, surpassing all 8B models and even Llama3-70B-Instruct and Llama3.1-70B-Instruct on average. Our results highlight the need for domain-specific adaptations in financial tasks, emphasizing future directions such as multi-table reasoning, long-context processing, and financial terminology comprehension. All our datasets, models, and codes are publicly available. Furthermore, we introduce a leaderboard for benchmarking future datasets and models.
INVESTORBENCH: A Benchmark for Financial Decision-Making Tasks with LLM-based Agent
Dec 24, 2024



Abstract:Recent advancements have underscored the potential of large language model (LLM)-based agents in financial decision-making. Despite this progress, the field currently encounters two main challenges: (1) the lack of a comprehensive LLM agent framework adaptable to a variety of financial tasks, and (2) the absence of standardized benchmarks and consistent datasets for assessing agent performance. To tackle these issues, we introduce \textsc{InvestorBench}, the first benchmark specifically designed for evaluating LLM-based agents in diverse financial decision-making contexts. InvestorBench enhances the versatility of LLM-enabled agents by providing a comprehensive suite of tasks applicable to different financial products, including single equities like stocks, cryptocurrencies and exchange-traded funds (ETFs). Additionally, we assess the reasoning and decision-making capabilities of our agent framework using thirteen different LLMs as backbone models, across various market environments and tasks. Furthermore, we have curated a diverse collection of open-source, multi-modal datasets and developed a comprehensive suite of environments for financial decision-making. This establishes a highly accessible platform for evaluating financial agents' performance across various scenarios.
Information Extraction from Clinical Notes: Are We Ready to Switch to Large Language Models?
Nov 15, 2024



Abstract:Backgrounds: Information extraction (IE) is critical in clinical natural language processing (NLP). While large language models (LLMs) excel on generative tasks, their performance on extractive tasks remains debated. Methods: We investigated Named Entity Recognition (NER) and Relation Extraction (RE) using 1,588 clinical notes from four sources (UT Physicians, MTSamples, MIMIC-III, and i2b2). We developed an annotated corpus covering 4 clinical entities and 16 modifiers, and compared instruction-tuned LLaMA-2 and LLaMA-3 against BiomedBERT in terms of performance, generalizability, computational resources, and throughput to BiomedBERT. Results: LLaMA models outperformed BiomedBERT across datasets. With sufficient training data, LLaMA showed modest improvements (1% on NER, 1.5-3.7% on RE); improvements were larger with limited training data. On unseen i2b2 data, LLaMA-3-70B outperformed BiomedBERT by 7% (F1) on NER and 4% on RE. However, LLaMA models required more computing resources and ran up to 28 times slower. We implemented "Kiwi," a clinical IE package featuring both models, available at https://kiwi.clinicalnlp.org/. Conclusion: This study is among the first to develop and evaluate a comprehensive clinical IE system using open-source LLMs. Results indicate that LLaMA models outperform BiomedBERT for clinical NER and RE but with higher computational costs and lower throughputs. These findings highlight that choosing between LLMs and traditional deep learning methods for clinical IE applications should remain task-specific, taking into account both performance metrics and practical considerations such as available computing resources and the intended use case scenarios.
Relation Extraction Using Large Language Models: A Case Study on Acupuncture Point Locations
Apr 15, 2024Abstract:In acupuncture therapy, the accurate location of acupoints is essential for its effectiveness. The advanced language understanding capabilities of large language models (LLMs) like Generative Pre-trained Transformers (GPT) present a significant opportunity for extracting relations related to acupoint locations from textual knowledge sources. This study aims to compare the performance of GPT with traditional deep learning models (Long Short-Term Memory (LSTM) and Bidirectional Encoder Representations from Transformers for Biomedical Text Mining (BioBERT)) in extracting acupoint-related location relations and assess the impact of pretraining and fine-tuning on GPT's performance. We utilized the World Health Organization Standard Acupuncture Point Locations in the Western Pacific Region (WHO Standard) as our corpus, which consists of descriptions of 361 acupoints. Five types of relations ('direction_of,' 'distance_of,' 'part_of,' 'near_acupoint,' and 'located_near') (n= 3,174) between acupoints were annotated. Five models were compared: BioBERT, LSTM, pre-trained GPT-3.5, fine-tuned GPT-3.5, as well as pre-trained GPT-4. Performance metrics included micro-average exact match precision, recall, and F1 scores. Our results demonstrate that fine-tuned GPT-3.5 consistently outperformed other models in F1 scores across all relation types. Overall, it achieved the highest micro-average F1 score of 0.92. This study underscores the effectiveness of LLMs like GPT in extracting relations related to acupoint locations, with implications for accurately modeling acupuncture knowledge and promoting standard implementation in acupuncture training and practice. The findings also contribute to advancing informatics applications in traditional and complementary medicine, showcasing the potential of LLMs in natural language processing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge