Shanghua Gao
AutoScientists: Self-Organizing Agent Teams for Long-Running Scientific Experimentation
May 27, 2026Abstract:Scientific research proceeds through iterative cycles of hypothesis generation, experiment design, execution, and revision. AI agents can automate parts of this process, but existing approaches typically follow a single research trajectory or coordinate through a central planner with fixed objectives. As a result, they struggle to sustain parallel exploration, adapt as experimental evidence changes, or preserve knowledge of failed directions over long-running experiments. We introduce AutoScientists, a decentralized team of AI agents for long-running computational scientific experimentation. Agents interpret a shared experimental state, self-organize into teams around promising hypotheses, critique proposals before using experimental compute, and share successes and failures to reduce redundant exploration. Under matched experimental budgets, AutoScientists improves over prior AI agents across biomedical machine learning, language-model training optimization, and protein fitness prediction. On BioML-Bench, spanning biomedical imaging, protein engineering, single-cell omics, and drug discovery, AutoScientists achieves a mean leaderboard percentile of 74.4% across 24 tasks, improving over the strongest AI agent by +8.33%. On GPT training optimization, AutoScientists reaches a target validation bits-per-byte 1.9x faster than Autoresearch and continues discovering improvements from a starting champion where the single-agent approach finds none (7 vs. 0 accepted improvements). On ProteinGym fitness prediction, AutoScientists discovers a method for ACE2-Spike binding that improves over the current state-of-the-art model by +12.5% in Spearman correlation. Applied without modification across all 217 ProteinGym assays, the same method improves over the prior state of the art by +6.5% (Spearman correlation).
When Sensors Fail: Temporal Sequence Models for Robust PPO under Sensor Drift
Mar 04, 2026Abstract:Real-world reinforcement learning systems must operate under distributional drift in their observation streams, yet most policy architectures implicitly assume fully observed and noise-free states. We study robustness of Proximal Policy Optimization (PPO) under temporally persistent sensor failures that induce partial observability and representation shift. To respond to this drift, we augment PPO with temporal sequence models, including Transformers and State Space Models (SSMs), to enable policies to infer missing information from history and maintain performance. Under a stochastic sensor failure process, we prove a high-probability bound on infinite-horizon reward degradation that quantifies how robustness depends on policy smoothness and failure persistence. Empirically, on MuJoCo continuous-control benchmarks with severe sensor dropout, we show Transformer-based sequence policies substantially outperform MLP, RNN, and SSM baselines in robustness, maintaining high returns even when large fractions of sensors are unavailable. These results demonstrate that temporal sequence reasoning provides a principled and practical mechanism for reliable operation under observation drift caused by sensor unreliability.
STRAND: Sequence-Conditioned Transport for Single-Cell Perturbations
Feb 10, 2026Abstract:Predicting how genetic perturbations change cellular state is a core problem for building controllable models of gene regulation. Perturbations targeting the same gene can produce different transcriptional responses depending on their genomic locus, including different transcription start sites and regulatory elements. Gene-level perturbation models collapse these distinct interventions into the same representation. We introduce STRAND, a generative model that predicts single-cell transcriptional responses by conditioning on regulatory DNA sequence. STRAND represents a perturbation by encoding the sequence at its genomic locus and uses this representation to parameterize a conditional transport process from control to perturbed cell states. Representing perturbations by sequence, rather than by a fixed set of gene identifiers, supports zero-shot inference at loci not seen during training and expands inference-time genomic coverage from ~1.5% for gene-level single-cell foundation models to ~95% of the genome. We evaluate STRAND on CRISPR perturbation datasets in K562, Jurkat, and RPE1 cells. STRAND improves discrimination scores by up to 33% in low-sample regimes, achieves the best average rank on unseen gene perturbation benchmarks, and improves transfer to novel cell lines by up to 0.14 in Pearson correlation. Ablations isolate the gains to sequence conditioning and transport, and case studies show that STRAND resolves functionally alternative transcription start sites missed by gene-level models.
Representation Entanglement for Generation:Training Diffusion Transformers Is Much Easier Than You Think
Jul 02, 2025
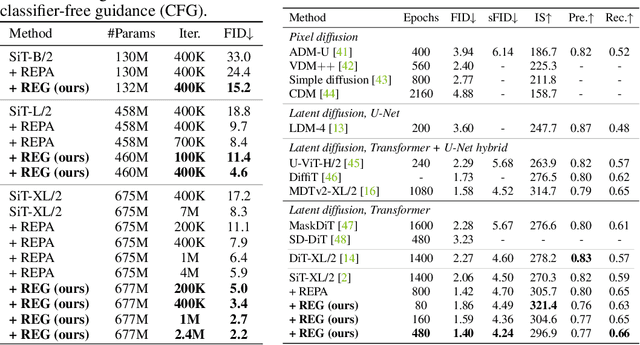
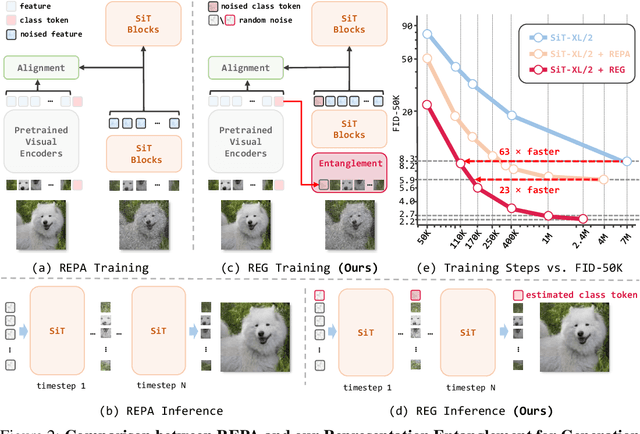
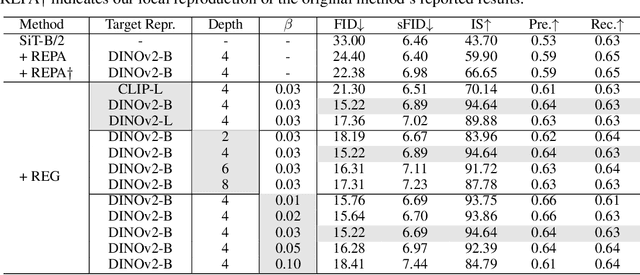
Abstract:REPA and its variants effectively mitigate training challenges in diffusion models by incorporating external visual representations from pretrained models, through alignment between the noisy hidden projections of denoising networks and foundational clean image representations. We argue that the external alignment, which is absent during the entire denoising inference process, falls short of fully harnessing the potential of discriminative representations. In this work, we propose a straightforward method called Representation Entanglement for Generation (REG), which entangles low-level image latents with a single high-level class token from pretrained foundation models for denoising. REG acquires the capability to produce coherent image-class pairs directly from pure noise, substantially improving both generation quality and training efficiency. This is accomplished with negligible additional inference overhead, requiring only one single additional token for denoising (<0.5\% increase in FLOPs and latency). The inference process concurrently reconstructs both image latents and their corresponding global semantics, where the acquired semantic knowledge actively guides and enhances the image generation process. On ImageNet 256$\times$256, SiT-XL/2 + REG demonstrates remarkable convergence acceleration, achieving $\textbf{63}\times$ and $\textbf{23}\times$ faster training than SiT-XL/2 and SiT-XL/2 + REPA, respectively. More impressively, SiT-L/2 + REG trained for merely 400K iterations outperforms SiT-XL/2 + REPA trained for 4M iterations ($\textbf{10}\times$ longer). Code is available at: https://github.com/Martinser/REG.
TxAgent: An AI Agent for Therapeutic Reasoning Across a Universe of Tools
Mar 14, 2025Abstract:Precision therapeutics require multimodal adaptive models that generate personalized treatment recommendations. We introduce TxAgent, an AI agent that leverages multi-step reasoning and real-time biomedical knowledge retrieval across a toolbox of 211 tools to analyze drug interactions, contraindications, and patient-specific treatment strategies. TxAgent evaluates how drugs interact at molecular, pharmacokinetic, and clinical levels, identifies contraindications based on patient comorbidities and concurrent medications, and tailors treatment strategies to individual patient characteristics. It retrieves and synthesizes evidence from multiple biomedical sources, assesses interactions between drugs and patient conditions, and refines treatment recommendations through iterative reasoning. It selects tools based on task objectives and executes structured function calls to solve therapeutic tasks that require clinical reasoning and cross-source validation. The ToolUniverse consolidates 211 tools from trusted sources, including all US FDA-approved drugs since 1939 and validated clinical insights from Open Targets. TxAgent outperforms leading LLMs, tool-use models, and reasoning agents across five new benchmarks: DrugPC, BrandPC, GenericPC, TreatmentPC, and DescriptionPC, covering 3,168 drug reasoning tasks and 456 personalized treatment scenarios. It achieves 92.1% accuracy in open-ended drug reasoning tasks, surpassing GPT-4o and outperforming DeepSeek-R1 (671B) in structured multi-step reasoning. TxAgent generalizes across drug name variants and descriptions. By integrating multi-step inference, real-time knowledge grounding, and tool-assisted decision-making, TxAgent ensures that treatment recommendations align with established clinical guidelines and real-world evidence, reducing the risk of adverse events and improving therapeutic decision-making.
Multimodal Medical Code Tokenizer
Feb 06, 2025



Abstract:Foundation models trained on patient electronic health records (EHRs) require tokenizing medical data into sequences of discrete vocabulary items. Existing tokenizers treat medical codes from EHRs as isolated textual tokens. However, each medical code is defined by its textual description, its position in ontological hierarchies, and its relationships to other codes, such as disease co-occurrences and drug-treatment associations. Medical vocabularies contain more than 600,000 codes with critical information for clinical reasoning. We introduce MedTok, a multimodal medical code tokenizer that uses the text descriptions and relational context of codes. MedTok processes text using a language model encoder and encodes the relational structure with a graph encoder. It then quantizes both modalities into a unified token space, preserving modality-specific and cross-modality information. We integrate MedTok into five EHR models and evaluate it on operational and clinical tasks across in-patient and out-patient datasets, including outcome prediction, diagnosis classification, drug recommendation, and risk stratification. Swapping standard EHR tokenizers with MedTok improves AUPRC across all EHR models, by 4.10% on MIMIC-III, 4.78% on MIMIC-IV, and 11.30% on EHRShot, with the largest gains in drug recommendation. Beyond EHR modeling, we demonstrate using MedTok tokenizer with medical QA systems. Our results demonstrate the potential of MedTok as a unified tokenizer for medical codes, improving tokenization for medical foundation models.
A Causality-aware Paradigm for Evaluating Creativity of Multimodal Large Language Models
Jan 25, 2025



Abstract:Recently, numerous benchmarks have been developed to evaluate the logical reasoning abilities of large language models (LLMs). However, assessing the equally important creative capabilities of LLMs is challenging due to the subjective, diverse, and data-scarce nature of creativity, especially in multimodal scenarios. In this paper, we consider the comprehensive pipeline for evaluating the creativity of multimodal LLMs, with a focus on suitable evaluation platforms and methodologies. First, we find the Oogiri game, a creativity-driven task requiring humor, associative thinking, and the ability to produce unexpected responses to text, images, or both. This game aligns well with the input-output structure of modern multimodal LLMs and benefits from a rich repository of high-quality, human-annotated creative responses, making it an ideal platform for studying LLM creativity. Next, beyond using the Oogiri game for standard evaluations like ranking and selection, we propose LoTbench, an interactive, causality-aware evaluation framework, to further address some intrinsic risks in standard evaluations, such as information leakage and limited interpretability. The proposed LoTbench not only quantifies LLM creativity more effectively but also visualizes the underlying creative thought processes. Our results show that while most LLMs exhibit constrained creativity, the performance gap between LLMs and humans is not insurmountable. Furthermore, we observe a strong correlation between results from the multimodal cognition benchmark MMMU and LoTbench, but only a weak connection with traditional creativity metrics. This suggests that LoTbench better aligns with human cognitive theories, highlighting cognition as a critical foundation in the early stages of creativity and enabling the bridging of diverse concepts. https://lotbench.github.io
Knowledge Graph Based Agent for Complex, Knowledge-Intensive QA in Medicine
Oct 07, 2024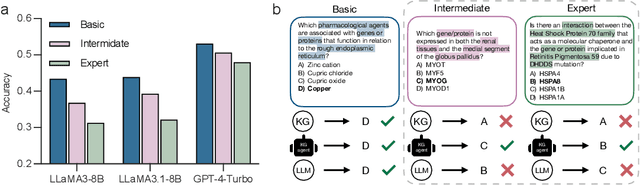
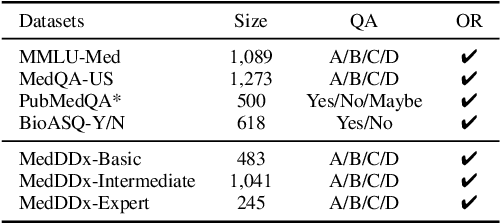
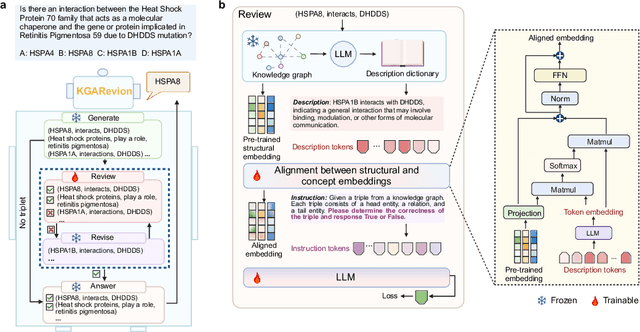
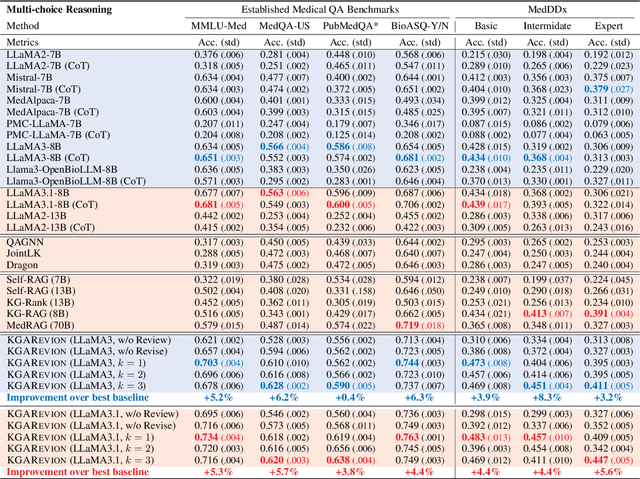
Abstract:Biomedical knowledge is uniquely complex and structured, requiring distinct reasoning strategies compared to other scientific disciplines like physics or chemistry. Biomedical scientists do not rely on a single approach to reasoning; instead, they use various strategies, including rule-based, prototype-based, and case-based reasoning. This diversity calls for flexible approaches that accommodate multiple reasoning strategies while leveraging in-domain knowledge. We introduce KGARevion, a knowledge graph (KG) based agent designed to address the complexity of knowledge-intensive medical queries. Upon receiving a query, KGARevion generates relevant triplets by using the knowledge base of the LLM. These triplets are then verified against a grounded KG to filter out erroneous information and ensure that only accurate, relevant data contribute to the final answer. Unlike RAG-based models, this multi-step process ensures robustness in reasoning while adapting to different models of medical reasoning. Evaluations on four gold-standard medical QA datasets show that KGARevion improves accuracy by over 5.2%, outperforming 15 models in handling complex medical questions. To test its capabilities, we curated three new medical QA datasets with varying levels of semantic complexity, where KGARevion achieved a 10.4% improvement in accuracy.
MoExtend: Tuning New Experts for Modality and Task Extension
Aug 07, 2024



Abstract:Large language models (LLMs) excel in various tasks but are primarily trained on text data, limiting their application scope. Expanding LLM capabilities to include vision-language understanding is vital, yet training them on multimodal data from scratch is challenging and costly. Existing instruction tuning methods, e.g., LLAVA, often connects a pretrained CLIP vision encoder and LLMs via fully fine-tuning LLMs to bridge the modality gap. However, full fine-tuning is plagued by catastrophic forgetting, i.e., forgetting previous knowledge, and high training costs particularly in the era of increasing tasks and modalities. To solve this issue, we introduce MoExtend, an effective framework designed to streamline the modality adaptation and extension of Mixture-of-Experts (MoE) models. MoExtend seamlessly integrates new experts into pre-trained MoE models, endowing them with novel knowledge without the need to tune pretrained models such as MoE and vision encoders. This approach enables rapid adaptation and extension to new modal data or tasks, effectively addressing the challenge of accommodating new modalities within LLMs. Furthermore, MoExtend avoids tuning pretrained models, thus mitigating the risk of catastrophic forgetting. Experimental results demonstrate the efficacy and efficiency of MoExtend in enhancing the multimodal capabilities of LLMs, contributing to advancements in multimodal AI research. Code: https://github.com/zhongshsh/MoExtend.
Composable Interventions for Language Models
Jul 09, 2024



Abstract:Test-time interventions for language models can enhance factual accuracy, mitigate harmful outputs, and improve model efficiency without costly retraining. But despite a flood of new methods, different types of interventions are largely developing independently. In practice, multiple interventions must be applied sequentially to the same model, yet we lack standardized ways to study how interventions interact. We fill this gap by introducing composable interventions, a framework to study the effects of using multiple interventions on the same language models, featuring new metrics and a unified codebase. Using our framework, we conduct extensive experiments and compose popular methods from three emerging intervention categories -- Knowledge Editing, Model Compression, and Machine Unlearning. Our results from 310 different compositions uncover meaningful interactions: compression hinders editing and unlearning, composing interventions hinges on their order of application, and popular general-purpose metrics are inadequate for assessing composability. Taken together, our findings showcase clear gaps in composability, suggesting a need for new multi-objective interventions. All of our code is public: https://github.com/hartvigsen-group/composable-interventions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge