Xiaoran Zhang
Snap-and-tune: combining deep learning and test-time optimization for high-fidelity cardiovascular volumetric meshing
Jun 09, 2025Abstract:High-quality volumetric meshing from medical images is a key bottleneck for physics-based simulations in personalized medicine. For volumetric meshing of complex medical structures, recent studies have often utilized deep learning (DL)-based template deformation approaches to enable fast test-time generation with high spatial accuracy. However, these approaches still exhibit limitations, such as limited flexibility at high-curvature areas and unrealistic inter-part distances. In this study, we introduce a simple yet effective snap-and-tune strategy that sequentially applies DL and test-time optimization, which combines fast initial shape fitting with more detailed sample-specific mesh corrections. Our method provides significant improvements in both spatial accuracy and mesh quality, while being fully automated and requiring no additional training labels. Finally, we demonstrate the versatility and usefulness of our newly generated meshes via solid mechanics simulations in two different software platforms. Our code is available at https://github.com/danpak94/Deep-Cardiac-Volumetric-Mesh.
DLW-CI: A Dynamic Likelihood-Weighted Cooperative Infotaxis Approach for Multi-Source Search in Urban Environments Using Consumer Drone Networks
Apr 19, 2025Abstract:Consumer-grade drones equipped with low-cost sensors have emerged as a cornerstone of Autonomous Intelligent Systems (AISs) for environmental monitoring and hazardous substance detection in urban environments. However, existing research primarily addresses single-source search problems, overlooking the complexities of real-world urban scenarios where both the location and quantity of hazardous sources remain unknown. To address this issue, we propose the Dynamic Likelihood-Weighted Cooperative Infotaxis (DLW-CI) approach for consumer drone networks. Our approach enhances multi-drone collaboration in AISs by combining infotaxis (a cognitive search strategy) with optimized source term estimation and an innovative cooperative mechanism. Specifically, we introduce a novel source term estimation method that utilizes multiple parallel particle filters, with each filter dedicated to estimating the parameters of a potentially unknown source within the search scene. Furthermore, we develop a cooperative mechanism based on dynamic likelihood weights to prevent multiple drones from simultaneously estimating and searching for the same source, thus optimizing the energy efficiency and search coverage of the consumer AIS. Experimental results demonstrate that the DLW-CI approach significantly outperforms baseline methods regarding success rate, accuracy, and root mean square error, particularly in scenarios with relatively few sources, regardless of the presence of obstacles. Also, the effectiveness of the proposed approach is verified in a diffusion scenario generated by the computational fluid dynamics (CFD) model. Research findings indicate that our approach could improve source estimation accuracy and search efficiency by consumer drone-based AISs, making a valuable contribution to environmental safety monitoring applications within smart city infrastructure.
Adapting Vision Foundation Models for Real-time Ultrasound Image Segmentation
Mar 31, 2025Abstract:We propose a novel approach that adapts hierarchical vision foundation models for real-time ultrasound image segmentation. Existing ultrasound segmentation methods often struggle with adaptability to new tasks, relying on costly manual annotations, while real-time approaches generally fail to match state-of-the-art performance. To overcome these limitations, we introduce an adaptive framework that leverages the vision foundation model Hiera to extract multi-scale features, interleaved with DINOv2 representations to enhance visual expressiveness. These enriched features are then decoded to produce precise and robust segmentation. We conduct extensive evaluations on six public datasets and one in-house dataset, covering both cardiac and thyroid ultrasound segmentation. Experiments show that our approach outperforms state-of-the-art methods across multiple datasets and excels with limited supervision, surpassing nnUNet by over 20\% on average in the 1\% and 10\% data settings. Our method achieves $\sim$77 FPS inference speed with TensorRT on a single GPU, enabling real-time clinical applications.
Progressive Test Time Energy Adaptation for Medical Image Segmentation
Mar 20, 2025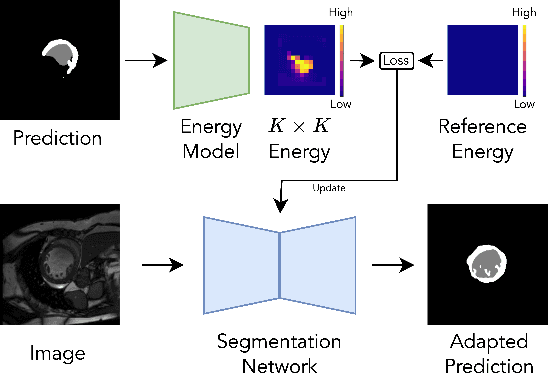
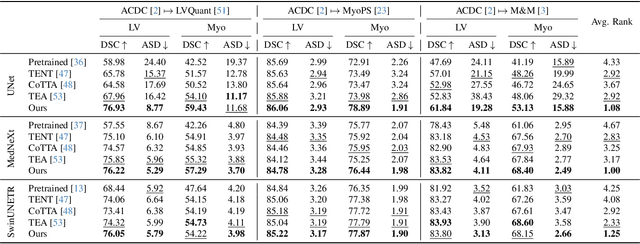
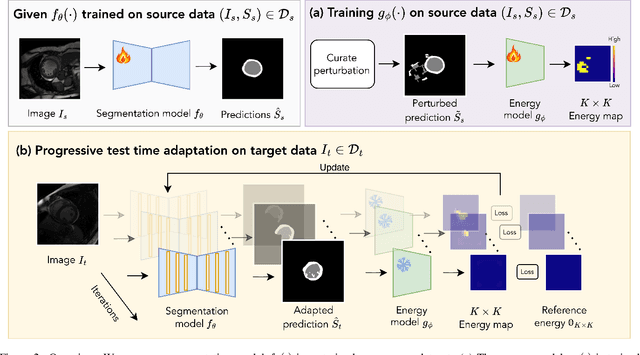
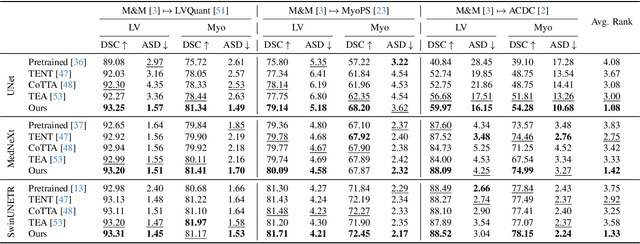
Abstract:We propose a model-agnostic, progressive test-time energy adaptation approach for medical image segmentation. Maintaining model performance across diverse medical datasets is challenging, as distribution shifts arise from inconsistent imaging protocols and patient variations. Unlike domain adaptation methods that require multiple passes through target data - impractical in clinical settings - our approach adapts pretrained models progressively as they process test data. Our method leverages a shape energy model trained on source data, which assigns an energy score at the patch level to segmentation maps: low energy represents in-distribution (accurate) shapes, while high energy signals out-of-distribution (erroneous) predictions. By minimizing this energy score at test time, we refine the segmentation model to align with the target distribution. To validate the effectiveness and adaptability, we evaluated our framework on eight public MRI (bSSFP, T1- and T2-weighted) and X-ray datasets spanning cardiac, spinal cord, and lung segmentation. We consistently outperform baselines both quantitatively and qualitatively.
L2RSI: Cross-view LiDAR-based Place Recognition for Large-scale Urban Scenes via Remote Sensing Imagery
Mar 14, 2025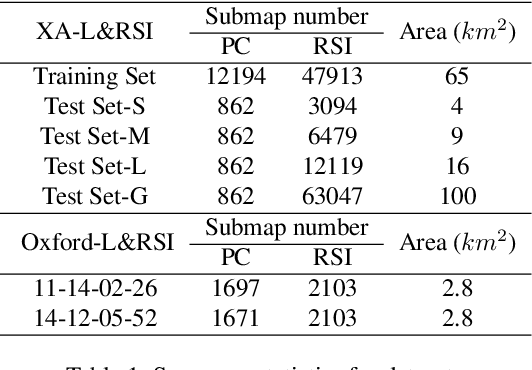
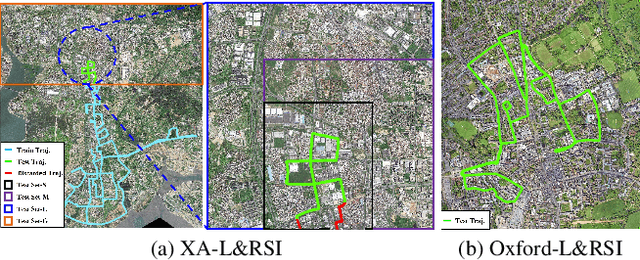
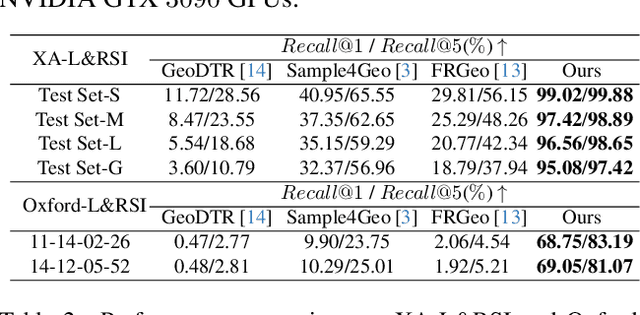
Abstract:We tackle the challenge of LiDAR-based place recognition, which traditionally depends on costly and time-consuming prior 3D maps. To overcome this, we first construct XA-L&RSI dataset, which encompasses approximately $110,000$ remote sensing submaps and $13,000$ LiDAR point cloud submaps captured in urban scenes, and propose a novel method, L2RSI, for cross-view LiDAR place recognition using high-resolution Remote Sensing Imagery. This approach enables large-scale localization capabilities at a reduced cost by leveraging readily available overhead images as map proxies. L2RSI addresses the dual challenges of cross-view and cross-modal place recognition by learning feature alignment between point cloud submaps and remote sensing submaps in the semantic domain. Additionally, we introduce a novel probability propagation method based on a dynamic Gaussian mixture model to refine position predictions, effectively leveraging temporal and spatial information. This approach enables large-scale retrieval and cross-scene generalization without fine-tuning. Extensive experiments on XA-L&RSI demonstrate that, within a $100km^2$ retrieval range, L2RSI accurately localizes $95.08\%$ of point cloud submaps within a $30m$ radius for top-$1$ retrieved location. We provide a video to more vividly display the place recognition results of L2RSI at https://shizw695.github.io/L2RSI/.
An Adaptive Correspondence Scoring Framework for Unsupervised Image Registration of Medical Images
Dec 01, 2023



Abstract:We propose an adaptive training scheme for unsupervised medical image registration. Existing methods rely on image reconstruction as the primary supervision signal. However, nuisance variables (e.g. noise and covisibility) often cause the loss of correspondence between medical images, violating the Lambertian assumption in physical waves (e.g. ultrasound) and consistent imaging acquisition. As the unsupervised learning scheme relies on intensity constancy to establish correspondence between images for reconstruction, this introduces spurious error residuals that are not modeled by the typical training objective. To mitigate this, we propose an adaptive framework that re-weights the error residuals with a correspondence scoring map during training, preventing the parametric displacement estimator from drifting away due to noisy gradients, which leads to performance degradations. To illustrate the versatility and effectiveness of our method, we tested our framework on three representative registration architectures across three medical image datasets along with other baselines. Our proposed adaptive framework consistently outperforms other methods both quantitatively and qualitatively. Paired t-tests show that our improvements are statistically significant. The code will be publicly available at \url{https://voldemort108x.github.io/AdaCS/}.
Heteroscedastic Uncertainty Estimation for Probabilistic Unsupervised Registration of Noisy Medical Images
Dec 01, 2023



Abstract:This paper proposes a heteroscedastic uncertainty estimation framework for unsupervised medical image registration. Existing methods rely on objectives (e.g. mean-squared error) that assume a uniform noise level across the image, disregarding the heteroscedastic and input-dependent characteristics of noise distribution in real-world medical images. This further introduces noisy gradients due to undesired penalization on outliers, causing unnatural deformation and performance degradation. To mitigate this, we propose an adaptive weighting scheme with a relative $\gamma$-exponentiated signal-to-noise ratio (SNR) for the displacement estimator after modeling the heteroscedastic noise using a separate variance estimator to prevent the model from being driven away by spurious gradients from error residuals, leading to more accurate displacement estimation. To illustrate the versatility and effectiveness of the proposed method, we tested our framework on two representative registration architectures across three medical image datasets. Our proposed framework consistently outperforms other baselines both quantitatively and qualitatively while also providing accurate and sensible uncertainty measures. Paired t-tests show that our improvements in registration accuracy are statistically significant. The code will be publicly available at \url{https://voldemort108x.github.io/hetero_uncertainty/}.
Rethinking Semi-Supervised Medical Image Segmentation: A Variance-Reduction Perspective
Feb 07, 2023



Abstract:For medical image segmentation, contrastive learning is the dominant practice to improve the quality of visual representations by contrasting semantically similar and dissimilar pairs of samples. This is enabled by the observation that without accessing ground truth label, negative examples with truly dissimilar anatomical features, if sampled, can significantly improve the performance. In reality, however, these samples may come from similar anatomical features and the models may struggle to distinguish the minority tail-class samples, making the tail classes more prone to misclassification, both of which typically lead to model collapse. In this paper, we propose ARCO, a semi-supervised contrastive learning (CL) framework with stratified group sampling theory in medical image segmentation. In particular, we first propose building ARCO through the concept of variance-reduced estimation, and show that certain variance-reduction techniques are particularly beneficial in medical image segmentation tasks with extremely limited labels. Furthermore, we theoretically prove these sampling techniques are universal in variance reduction. Finally, we experimentally validate our approaches on three benchmark datasets with different label settings, and our methods consistently outperform state-of-the-art semi- and fully-supervised methods. Additionally, we augment the CL frameworks with these sampling techniques and demonstrate significant gains over previous methods. We believe our work is an important step towards semi-supervised medical image segmentation by quantifying the limitation of current self-supervision objectives for accomplishing medical image analysis tasks.
Mine yOur owN Anatomy: Revisiting Medical Image Segmentation with Extremely Limited Labels
Sep 28, 2022
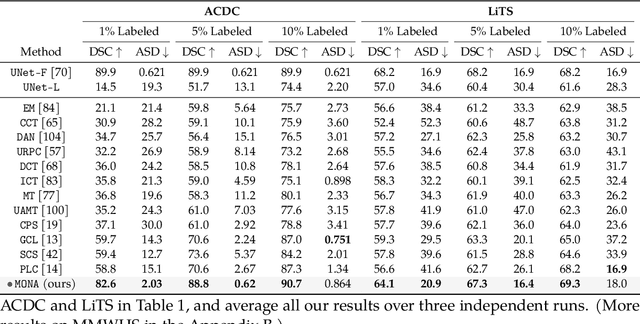
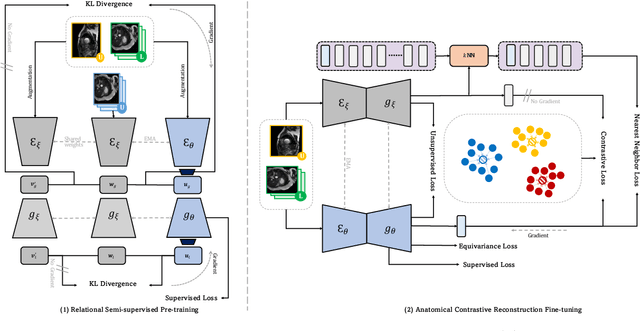
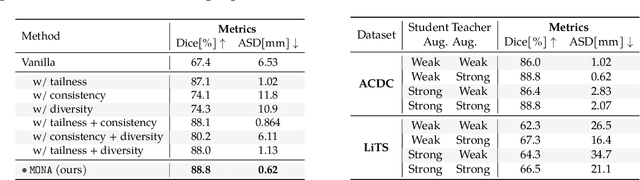
Abstract:Recent studies on contrastive learning have achieved remarkable performance solely by leveraging few labels in the context of medical image segmentation. Existing methods mainly focus on instance discrimination and invariant mapping. However, they face three common pitfalls: (1) tailness: medical image data usually follows an implicit long-tail class distribution. Blindly leveraging all pixels in training hence can lead to the data imbalance issues, and cause deteriorated performance; (2) consistency: it remains unclear whether a segmentation model has learned meaningful and yet consistent anatomical features due to the intra-class variations between different anatomical features; and (3) diversity: the intra-slice correlations within the entire dataset have received significantly less attention. This motivates us to seek a principled approach for strategically making use of the dataset itself to discover similar yet distinct samples from different anatomical views. In this paper, we introduce a novel semi-supervised medical image segmentation framework termed Mine yOur owN Anatomy (MONA), and make three contributions. First, prior work argues that every pixel equally matters to the model training; we observe empirically that this alone is unlikely to define meaningful anatomical features, mainly due to lacking the supervision signal. We show two simple solutions towards learning invariances - through the use of stronger data augmentations and nearest neighbors. Second, we construct a set of objectives that encourage the model to be capable of decomposing medical images into a collection of anatomical features in an unsupervised manner. Lastly, our extensive results on three benchmark datasets with different labeled settings validate the effectiveness of our proposed MONA which achieves new state-of-the-art under different labeled settings.
Learning correspondences of cardiac motion from images using biomechanics-informed modeling
Sep 01, 2022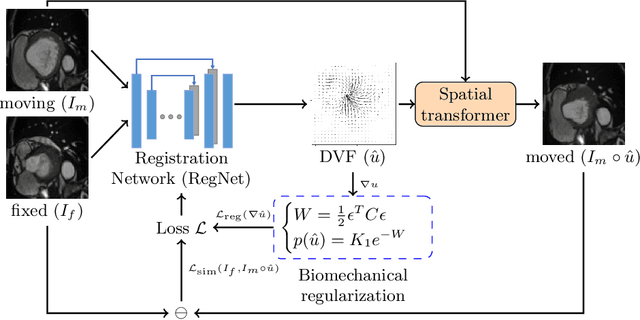
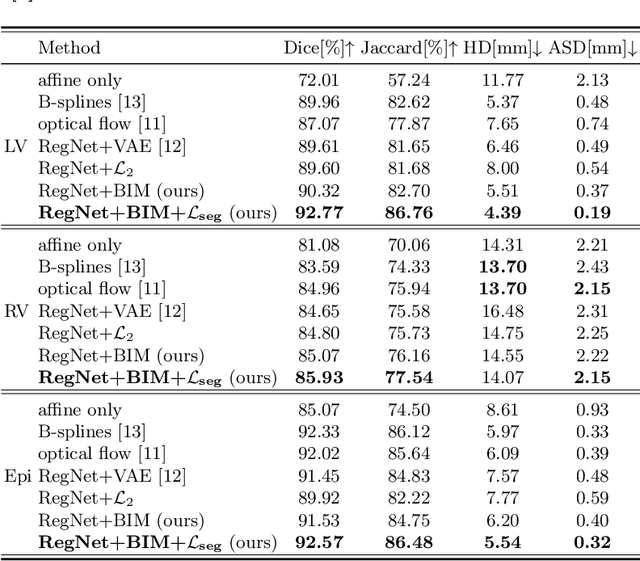
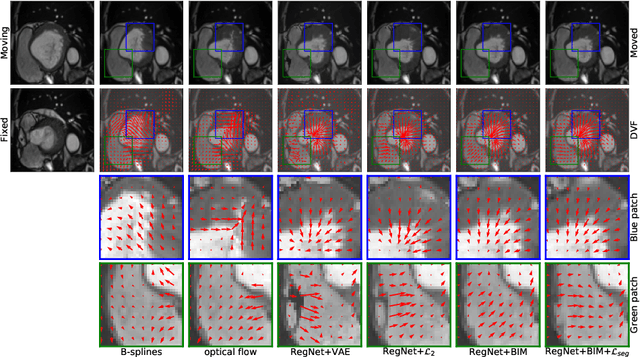
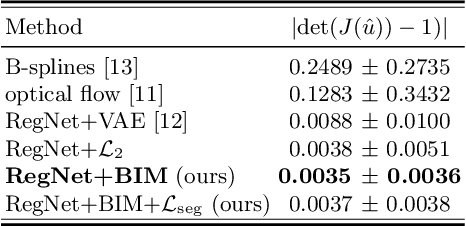
Abstract:Learning spatial-temporal correspondences in cardiac motion from images is important for understanding the underlying dynamics of cardiac anatomical structures. Many methods explicitly impose smoothness constraints such as the $\mathcal{L}_2$ norm on the displacement vector field (DVF), while usually ignoring biomechanical feasibility in the transformation. Other geometric constraints either regularize specific regions of interest such as imposing incompressibility on the myocardium or introduce additional steps such as training a separate network-based regularizer on physically simulated datasets. In this work, we propose an explicit biomechanics-informed prior as regularization on the predicted DVF in modeling a more generic biomechanically plausible transformation within all cardiac structures without introducing additional training complexity. We validate our methods on two publicly available datasets in the context of 2D MRI data and perform extensive experiments to illustrate the effectiveness and robustness of our proposed methods compared to other competing regularization schemes. Our proposed methods better preserve biomechanical properties by visual assessment and show advantages in segmentation performance using quantitative evaluation metrics. The code is publicly available at \url{https://github.com/Voldemort108X/bioinformed_reg}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge