Wenjun Liao
University of Electronic Science and Technology of China, Chengdu, China, Department of Radiation Oncology, Sichuan Cancer Hospital & Institute, Sichuan Cancer Center, Chengdu, China
SegRap2025: A Benchmark of Gross Tumor Volume and Lymph Node Clinical Target Volume Segmentation for Radiotherapy Planning of Nasopharyngeal Carcinoma
Jan 28, 2026Abstract:Accurate delineation of Gross Tumor Volume (GTV), Lymph Node Clinical Target Volume (LN CTV), and Organ-at-Risk (OAR) from Computed Tomography (CT) scans is essential for precise radiotherapy planning in Nasopharyngeal Carcinoma (NPC). Building upon SegRap2023, which focused on OAR and GTV segmentation using single-center paired non-contrast CT (ncCT) and contrast-enhanced CT (ceCT) scans, the SegRap2025 challenge aims to enhance the generalizability and robustness of segmentation models across imaging centers and modalities. SegRap2025 comprises two tasks: Task01 addresses GTV segmentation using paired CT from the SegRap2023 dataset, with an additional external testing set to evaluate cross-center generalization, and Task02 focuses on LN CTV segmentation using multi-center training data and an unseen external testing set, where each case contains paired CT scans or a single modality, emphasizing both cross-center and cross-modality robustness. This paper presents the challenge setup and provides a comprehensive analysis of the solutions submitted by ten participating teams. For GTV segmentation task, the top-performing models achieved average Dice Similarity Coefficient (DSC) of 74.61% and 56.79% on the internal and external testing cohorts, respectively. For LN CTV segmentation task, the highest average DSC values reached 60.24%, 60.50%, and 57.23% on paired CT, ceCT-only, and ncCT-only subsets, respectively. SegRap2025 establishes a large-scale multi-center, multi-modality benchmark for evaluating the generalization and robustness in radiotherapy target segmentation, providing valuable insights toward clinically applicable automated radiotherapy planning systems. The benchmark is available at: https://hilab-git.github.io/SegRap2025_Challenge.
CLISC: Bridging clip and sam by enhanced cam for unsupervised brain tumor segmentation
Jan 27, 2025



Abstract:Brain tumor segmentation is important for diagnosis of the tumor, and current deep-learning methods rely on a large set of annotated images for training, with high annotation costs. Unsupervised segmentation is promising to avoid human annotations while the performance is often limited. In this study, we present a novel unsupervised segmentation approach that leverages the capabilities of foundation models, and it consists of three main steps: (1) A vision-language model (i.e., CLIP) is employed to obtain image-level pseudo-labels for training a classification network. Class Activation Mapping (CAM) is then employed to extract Regions of Interest (ROIs), where an adaptive masking-based data augmentation is used to enhance ROI identification.(2) The ROIs are used to generate bounding box and point prompts for the Segment Anything Model (SAM) to obtain segmentation pseudo-labels. (3) A 3D segmentation network is trained with the SAM-derived pseudo-labels, where low-quality pseudo-labels are filtered out in a self-learning process based on the similarity between the SAM's output and the network's prediction. Evaluation on the BraTS2020 dataset demonstrates that our approach obtained an average Dice Similarity Score (DSC) of 85.60%, outperforming five state-of-the-art unsupervised segmentation methods by more than 10 percentage points. Besides, our approach outperforms directly using SAM for zero-shot inference, and its performance is close to fully supervised learning.
Head and Neck Tumor Segmentation of MRI from Pre- and Mid-radiotherapy with Pre-training, Data Augmentation and Dual Flow UNet
Dec 19, 2024



Abstract:Head and neck tumors and metastatic lymph nodes are crucial for treatment planning and prognostic analysis. Accurate segmentation and quantitative analysis of these structures require pixel-level annotation, making automated segmentation techniques essential for the diagnosis and treatment of head and neck cancer. In this study, we investigated the effects of multiple strategies on the segmentation of pre-radiotherapy (pre-RT) and mid-radiotherapy (mid-RT) images. For the segmentation of pre-RT images, we utilized: 1) a fully supervised learning approach, and 2) the same approach enhanced with pre-trained weights and the MixUp data augmentation technique. For mid-RT images, we introduced a novel computational-friendly network architecture that features separate encoders for mid-RT images and registered pre-RT images with their labels. The mid-RT encoder branch integrates information from pre-RT images and labels progressively during the forward propagation. We selected the highest-performing model from each fold and used their predictions to create an ensemble average for inference. In the final test, our models achieved a segmentation performance of 82.38% for pre-RT and 72.53% for mid-RT on aggregated Dice Similarity Coefficient (DSC) as HiLab. Our code is available at https://github.com/WltyBY/HNTS-MRG2024_train_code.
Weakly Supervised Lymph Nodes Segmentation Based on Partial Instance Annotations with Pre-trained Dual-branch Network and Pseudo Label Learning
Aug 18, 2024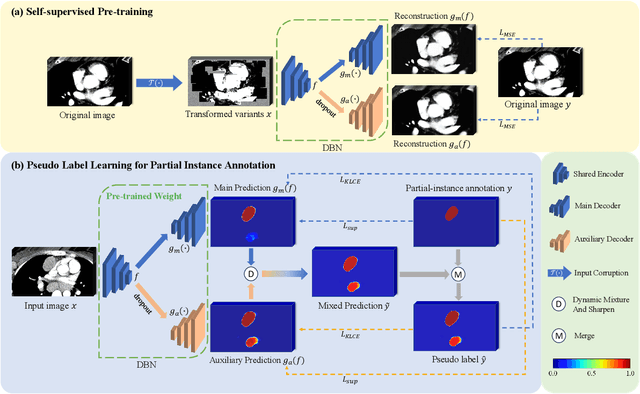
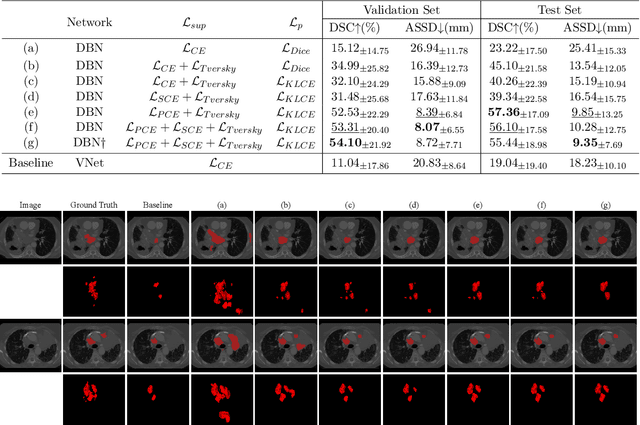
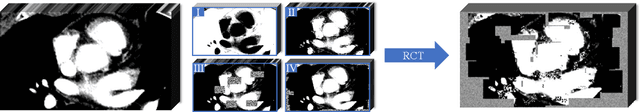
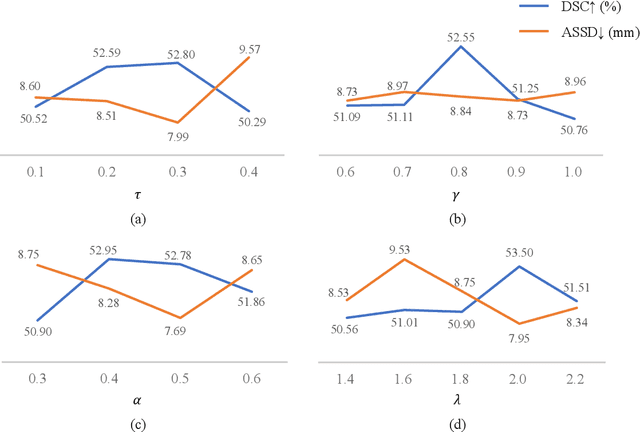
Abstract:Assessing the presence of potentially malignant lymph nodes aids in estimating cancer progression, and identifying surrounding benign lymph nodes can assist in determining potential metastatic pathways for cancer. For quantitative analysis, automatic segmentation of lymph nodes is crucial. However, due to the labor-intensive and time-consuming manual annotation process required for a large number of lymph nodes, it is more practical to annotate only a subset of the lymph node instances to reduce annotation costs. In this study, we propose a pre-trained Dual-Branch network with Dynamically Mixed Pseudo label (DBDMP) to learn from partial instance annotations for lymph nodes segmentation. To obtain reliable pseudo labels for lymph nodes that are not annotated, we employ a dual-decoder network to generate different outputs that are then dynamically mixed. We integrate the original weak partial annotations with the mixed pseudo labels to supervise the network. To further leverage the extensive amount of unannotated voxels, we apply a self-supervised pre-training strategy to enhance the model's feature extraction capability. Experiments on the mediastinal Lymph Node Quantification (LNQ) dataset demonstrate that our method, compared to directly learning from partial instance annotations, significantly improves the Dice Similarity Coefficient (DSC) from 11.04% to 54.10% and reduces the Average Symmetric Surface Distance (ASSD) from 20.83 $mm$ to 8.72 $mm$. The code is available at https://github.com/WltyBY/LNQ2023_training_code.git
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2024:013
Rethinking Abdominal Organ Segmentation (RAOS) in the clinical scenario: A robustness evaluation benchmark with challenging cases
Jun 19, 2024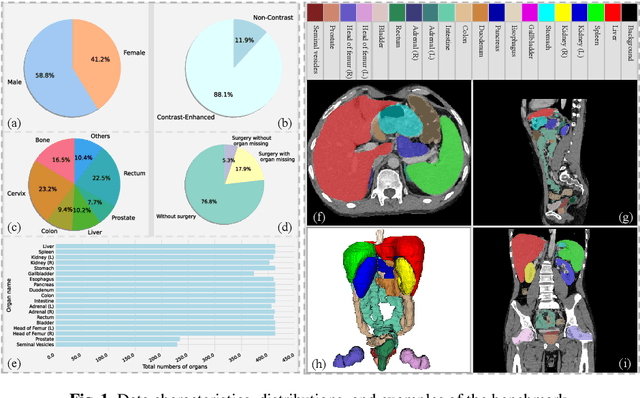
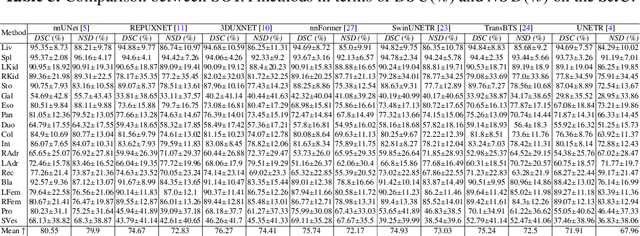
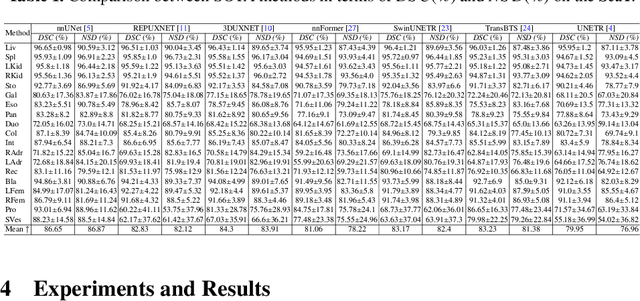
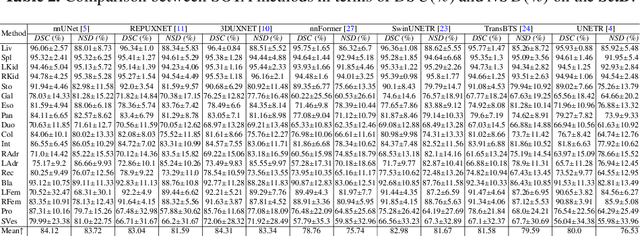

Abstract:Deep learning has enabled great strides in abdominal multi-organ segmentation, even surpassing junior oncologists on common cases or organs. However, robustness on corner cases and complex organs remains a challenging open problem for clinical adoption. To investigate model robustness, we collected and annotated the RAOS dataset comprising 413 CT scans ($\sim$80k 2D images, $\sim$8k 3D organ annotations) from 413 patients each with 17 (female) or 19 (male) labelled organs, manually delineated by oncologists. We grouped scans based on clinical information into 1) diagnosis/radiotherapy (317 volumes), 2) partial excision without the whole organ missing (22 volumes), and 3) excision with the whole organ missing (74 volumes). RAOS provides a potential benchmark for evaluating model robustness including organ hallucination. It also includes some organs that can be very hard to access on public datasets like the rectum, colon, intestine, prostate and seminal vesicles. We benchmarked several state-of-the-art methods in these three clinical groups to evaluate performance and robustness. We also assessed cross-generalization between RAOS and three public datasets. This dataset and comprehensive analysis establish a potential baseline for future robustness research: \url{https://github.com/Luoxd1996/RAOS}.
Diversified and Personalized Multi-rater Medical Image Segmentation
Mar 20, 2024Abstract:Annotation ambiguity due to inherent data uncertainties such as blurred boundaries in medical scans and different observer expertise and preferences has become a major obstacle for training deep-learning based medical image segmentation models. To address it, the common practice is to gather multiple annotations from different experts, leading to the setting of multi-rater medical image segmentation. Existing works aim to either merge different annotations into the "groundtruth" that is often unattainable in numerous medical contexts, or generate diverse results, or produce personalized results corresponding to individual expert raters. Here, we bring up a more ambitious goal for multi-rater medical image segmentation, i.e., obtaining both diversified and personalized results. Specifically, we propose a two-stage framework named D-Persona (first Diversification and then Personalization). In Stage I, we exploit multiple given annotations to train a Probabilistic U-Net model, with a bound-constrained loss to improve the prediction diversity. In this way, a common latent space is constructed in Stage I, where different latent codes denote diversified expert opinions. Then, in Stage II, we design multiple attention-based projection heads to adaptively query the corresponding expert prompts from the shared latent space, and then perform the personalized medical image segmentation. We evaluated the proposed model on our in-house Nasopharyngeal Carcinoma dataset and the public lung nodule dataset (i.e., LIDC-IDRI). Extensive experiments demonstrated our D-Persona can provide diversified and personalized results at the same time, achieving new SOTA performance for multi-rater medical image segmentation. Our code will be released at https://github.com/ycwu1997/D-Persona.
SegRap2023: A Benchmark of Organs-at-Risk and Gross Tumor Volume Segmentation for Radiotherapy Planning of Nasopharyngeal Carcinoma
Dec 15, 2023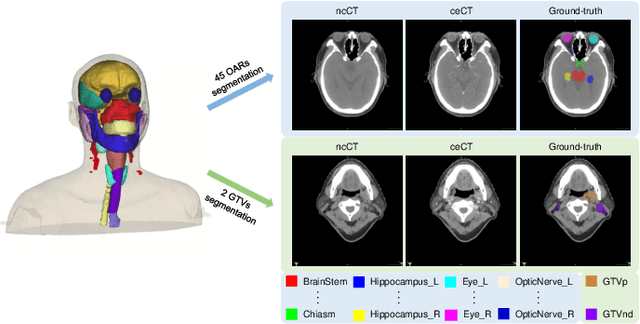
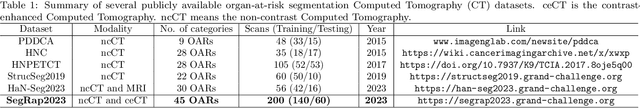
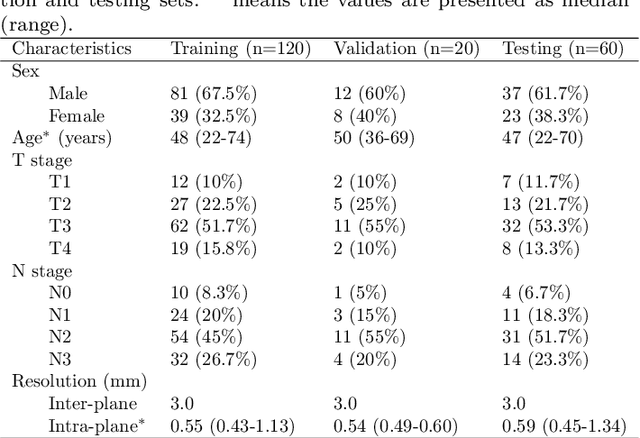
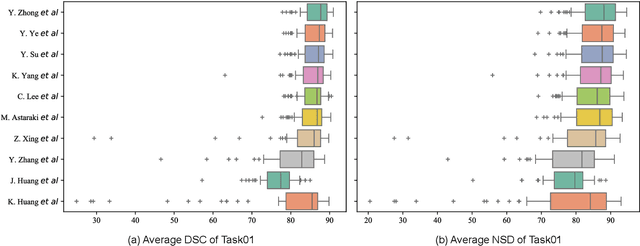
Abstract:Radiation therapy is a primary and effective NasoPharyngeal Carcinoma (NPC) treatment strategy. The precise delineation of Gross Tumor Volumes (GTVs) and Organs-At-Risk (OARs) is crucial in radiation treatment, directly impacting patient prognosis. Previously, the delineation of GTVs and OARs was performed by experienced radiation oncologists. Recently, deep learning has achieved promising results in many medical image segmentation tasks. However, for NPC OARs and GTVs segmentation, few public datasets are available for model development and evaluation. To alleviate this problem, the SegRap2023 challenge was organized in conjunction with MICCAI2023 and presented a large-scale benchmark for OAR and GTV segmentation with 400 Computed Tomography (CT) scans from 200 NPC patients, each with a pair of pre-aligned non-contrast and contrast-enhanced CT scans. The challenge's goal was to segment 45 OARs and 2 GTVs from the paired CT scans. In this paper, we detail the challenge and analyze the solutions of all participants. The average Dice similarity coefficient scores for all submissions ranged from 76.68\% to 86.70\%, and 70.42\% to 73.44\% for OARs and GTVs, respectively. We conclude that the segmentation of large-size OARs is well-addressed, and more efforts are needed for GTVs and small-size or thin-structure OARs. The benchmark will remain publicly available here: https://segrap2023.grand-challenge.org
Dual-Reference Source-Free Active Domain Adaptation for Nasopharyngeal Carcinoma Tumor Segmentation across Multiple Hospitals
Sep 23, 2023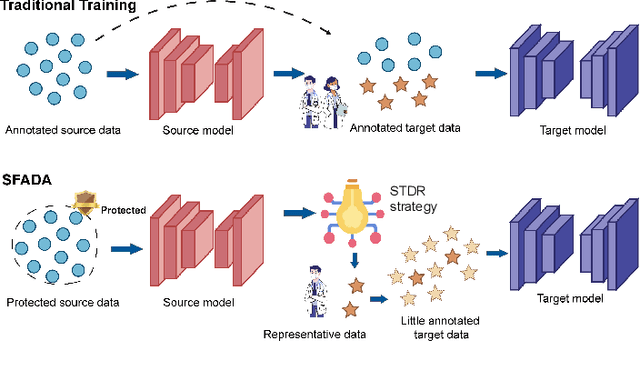
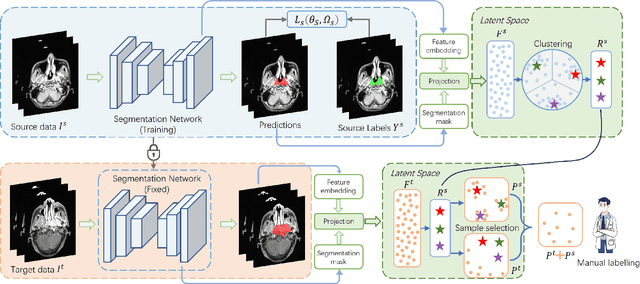
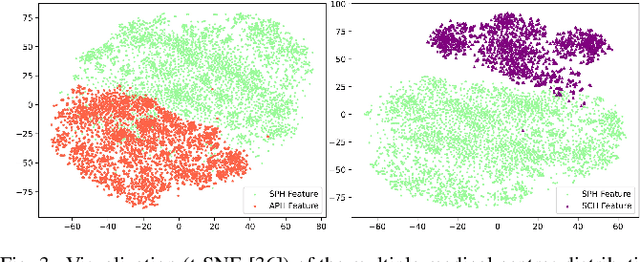
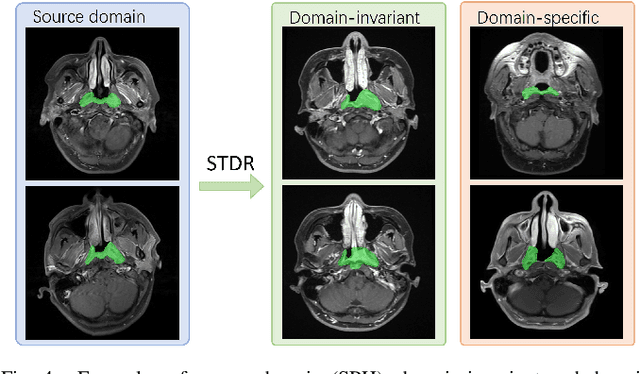
Abstract:Nasopharyngeal carcinoma (NPC) is a prevalent and clinically significant malignancy that predominantly impacts the head and neck area. Precise delineation of the Gross Tumor Volume (GTV) plays a pivotal role in ensuring effective radiotherapy for NPC. Despite recent methods that have achieved promising results on GTV segmentation, they are still limited by lacking carefully-annotated data and hard-to-access data from multiple hospitals in clinical practice. Although some unsupervised domain adaptation (UDA) has been proposed to alleviate this problem, unconditionally mapping the distribution distorts the underlying structural information, leading to inferior performance. To address this challenge, we devise a novel Sourece-Free Active Domain Adaptation (SFADA) framework to facilitate domain adaptation for the GTV segmentation task. Specifically, we design a dual reference strategy to select domain-invariant and domain-specific representative samples from a specific target domain for annotation and model fine-tuning without relying on source-domain data. Our approach not only ensures data privacy but also reduces the workload for oncologists as it just requires annotating a few representative samples from the target domain and does not need to access the source data. We collect a large-scale clinical dataset comprising 1057 NPC patients from five hospitals to validate our approach. Experimental results show that our method outperforms the UDA methods and achieves comparable results to the fully supervised upper bound, even with few annotations, highlighting the significant medical utility of our approach. In addition, there is no public dataset about multi-center NPC segmentation, we will release code and dataset for future research.
Scribble-based 3D Multiple Abdominal Organ Segmentation via Triple-branch Multi-dilated Network with Pixel- and Class-wise Consistency
Sep 18, 2023

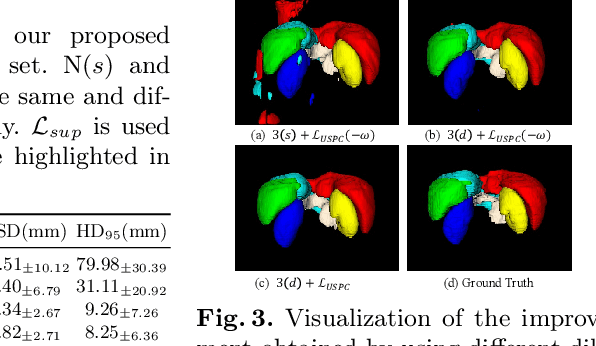
Abstract:Multi-organ segmentation in abdominal Computed Tomography (CT) images is of great importance for diagnosis of abdominal lesions and subsequent treatment planning. Though deep learning based methods have attained high performance, they rely heavily on large-scale pixel-level annotations that are time-consuming and labor-intensive to obtain. Due to its low dependency on annotation, weakly supervised segmentation has attracted great attention. However, there is still a large performance gap between current weakly-supervised methods and fully supervised learning, leaving room for exploration. In this work, we propose a novel 3D framework with two consistency constraints for scribble-supervised multiple abdominal organ segmentation from CT. Specifically, we employ a Triple-branch multi-Dilated network (TDNet) with one encoder and three decoders using different dilation rates to capture features from different receptive fields that are complementary to each other to generate high-quality soft pseudo labels. For more stable unsupervised learning, we use voxel-wise uncertainty to rectify the soft pseudo labels and then supervise the outputs of each decoder. To further regularize the network, class relationship information is exploited by encouraging the generated class affinity matrices to be consistent across different decoders under multi-view projection. Experiments on the public WORD dataset show that our method outperforms five existing scribble-supervised methods.
CDDSA: Contrastive Domain Disentanglement and Style Augmentation for Generalizable Medical Image Segmentation
Nov 22, 2022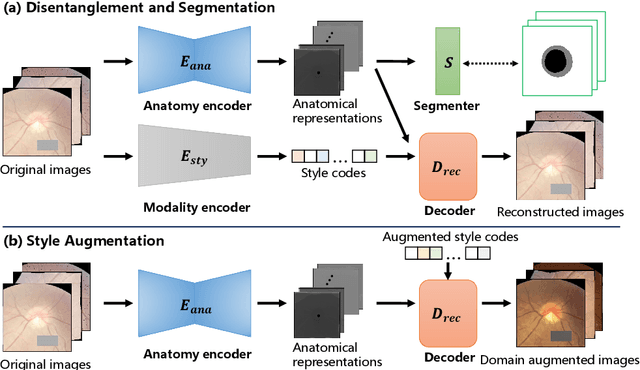
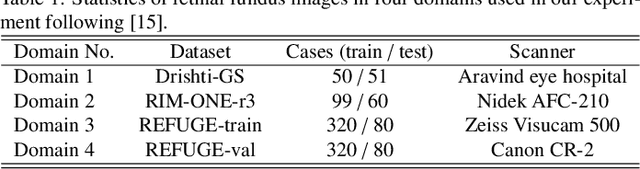
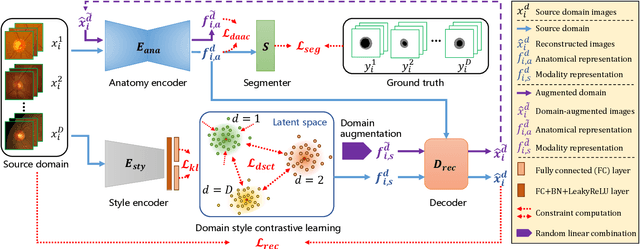

Abstract:Generalization to previously unseen images with potential domain shifts and different styles is essential for clinically applicable medical image segmentation, and the ability to disentangle domain-specific and domain-invariant features is key for achieving Domain Generalization (DG). However, existing DG methods can hardly achieve effective disentanglement to get high generalizability. To deal with this problem, we propose an efficient Contrastive Domain Disentanglement and Style Augmentation (CDDSA) framework for generalizable medical image segmentation. First, a disentangle network is proposed to decompose an image into a domain-invariant anatomical representation and a domain-specific style code, where the former is sent to a segmentation model that is not affected by the domain shift, and the disentangle network is regularized by a decoder that combines the anatomical and style codes to reconstruct the input image. Second, to achieve better disentanglement, a contrastive loss is proposed to encourage the style codes from the same domain and different domains to be compact and divergent, respectively. Thirdly, to further improve generalizability, we propose a style augmentation method based on the disentanglement representation to synthesize images in various unseen styles with shared anatomical structures. Our method was validated on a public multi-site fundus image dataset for optic cup and disc segmentation and an in-house multi-site Nasopharyngeal Carcinoma Magnetic Resonance Image (NPC-MRI) dataset for nasopharynx Gross Tumor Volume (GTVnx) segmentation. Experimental results showed that the proposed CDDSA achieved remarkable generalizability across different domains, and it outperformed several state-of-the-art methods in domain-generalizable segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge