Simone Bendazzoli
Anatomy-Aware Lymphoma Lesion Detection in Whole-Body PET/CT
Nov 10, 2025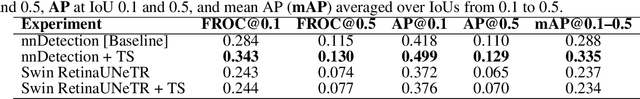
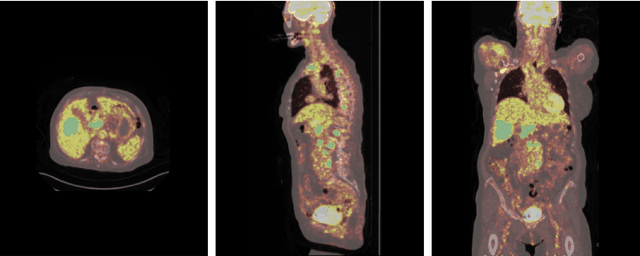
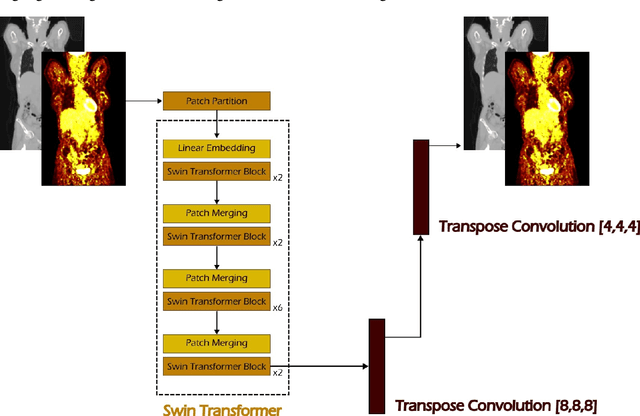
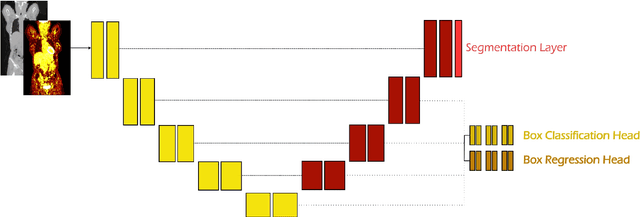
Abstract:Early cancer detection is crucial for improving patient outcomes, and 18F FDG PET/CT imaging plays a vital role by combining metabolic and anatomical information. Accurate lesion detection remains challenging due to the need to identify multiple lesions of varying sizes. In this study, we investigate the effect of adding anatomy prior information to deep learning-based lesion detection models. In particular, we add organ segmentation masks from the TotalSegmentator tool as auxiliary inputs to provide anatomical context to nnDetection, which is the state-of-the-art for lesion detection, and Swin Transformer. The latter is trained in two stages that combine self-supervised pre-training and supervised fine-tuning. The method is tested in the AutoPET and Karolinska lymphoma datasets. The results indicate that the inclusion of anatomical priors substantially improves the detection performance within the nnDetection framework, while it has almost no impact on the performance of the vision transformer. Moreover, we observe that Swin Transformer does not offer clear advantages over conventional convolutional neural network (CNN) encoders used in nnDetection. These findings highlight the critical role of the anatomical context in cancer lesion detection, especially in CNN-based models.
Lesion Segmentation in Whole-Body Multi-Tracer PET-CT Images; a Contribution to AutoPET 2024 Challenge
Sep 22, 2024



Abstract:The automatic segmentation of pathological regions within whole-body PET-CT volumes has the potential to streamline various clinical applications such as diagno-sis, prognosis, and treatment planning. This study aims to address this challenge by contributing to the AutoPET MICCAI 2024 challenge through a proposed workflow that incorporates image preprocessing, tracer classification, and lesion segmentation steps. The implementation of this pipeline led to a significant enhancement in the segmentation accuracy of the models. This improvement is evidenced by an average overall Dice score of 0.548 across 1611 training subjects, 0.631 and 0.559 for classi-fied FDG and PSMA subjects of the training set, and 0.792 on the preliminary testing phase dataset.
Unsupervised Domain Adaptation for Pediatric Brain Tumor Segmentation
Jun 24, 2024



Abstract:Significant advances have been made toward building accurate automatic segmentation models for adult gliomas. However, the performance of these models often degrades when applied to pediatric glioma due to their imaging and clinical differences (domain shift). Obtaining sufficient annotated data for pediatric glioma is typically difficult because of its rare nature. Also, manual annotations are scarce and expensive. In this work, we propose Domain-Adapted nnU-Net (DA-nnUNet) to perform unsupervised domain adaptation from adult glioma (source domain) to pediatric glioma (target domain). Specifically, we add a domain classifier connected with a gradient reversal layer (GRL) to a backbone nnU-Net. Once the classifier reaches a very high accuracy, the GRL is activated with the goal of transferring domain-invariant features from the classifier to the segmentation model while preserving segmentation accuracy on the source domain. The accuracy of the classifier slowly degrades to chance levels. No annotations are used in the target domain. The method is compared to 8 different supervised models using BraTS-Adult glioma (N=1251) and BraTS-PED glioma data (N=99). The proposed method shows notable performance enhancements in the tumor core (TC) region compared to the model that only uses adult data: ~32% better Dice scores and ~20 better 95th percentile Hausdorff distances. Moreover, our unsupervised approach shows no statistically significant difference compared to the practical upper bound model using manual annotations from both datasets in TC region. The code is shared at https://github.com/Fjr9516/DA_nnUNet.
SegRap2023: A Benchmark of Organs-at-Risk and Gross Tumor Volume Segmentation for Radiotherapy Planning of Nasopharyngeal Carcinoma
Dec 15, 2023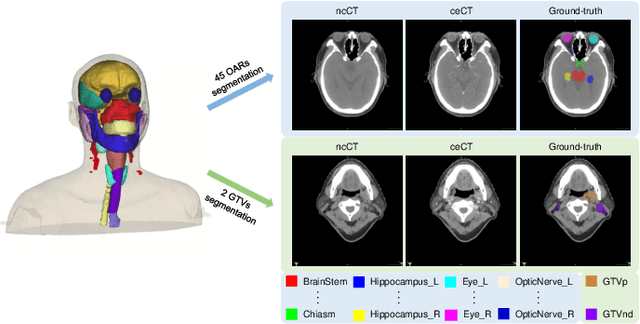
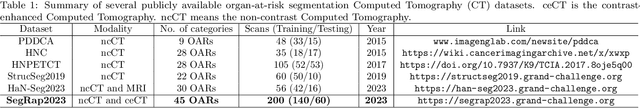
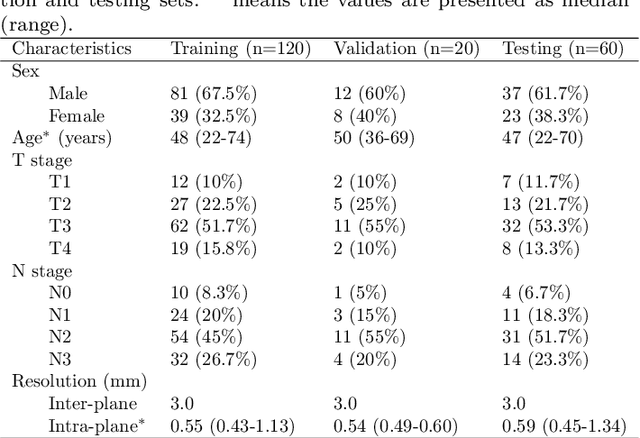
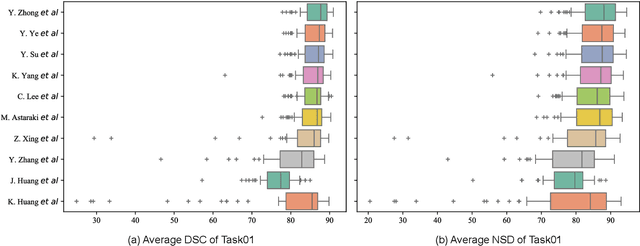
Abstract:Radiation therapy is a primary and effective NasoPharyngeal Carcinoma (NPC) treatment strategy. The precise delineation of Gross Tumor Volumes (GTVs) and Organs-At-Risk (OARs) is crucial in radiation treatment, directly impacting patient prognosis. Previously, the delineation of GTVs and OARs was performed by experienced radiation oncologists. Recently, deep learning has achieved promising results in many medical image segmentation tasks. However, for NPC OARs and GTVs segmentation, few public datasets are available for model development and evaluation. To alleviate this problem, the SegRap2023 challenge was organized in conjunction with MICCAI2023 and presented a large-scale benchmark for OAR and GTV segmentation with 400 Computed Tomography (CT) scans from 200 NPC patients, each with a pair of pre-aligned non-contrast and contrast-enhanced CT scans. The challenge's goal was to segment 45 OARs and 2 GTVs from the paired CT scans. In this paper, we detail the challenge and analyze the solutions of all participants. The average Dice similarity coefficient scores for all submissions ranged from 76.68\% to 86.70\%, and 70.42\% to 73.44\% for OARs and GTVs, respectively. We conclude that the segmentation of large-size OARs is well-addressed, and more efforts are needed for GTVs and small-size or thin-structure OARs. The benchmark will remain publicly available here: https://segrap2023.grand-challenge.org
Fully Automatic Segmentation of Gross Target Volume and Organs-at-Risk for Radiotherapy Planning of Nasopharyngeal Carcinoma
Oct 04, 2023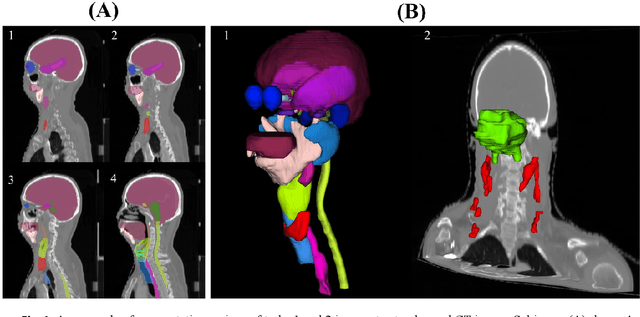


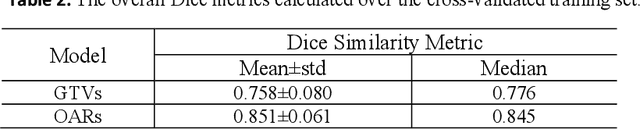
Abstract:Target segmentation in CT images of Head&Neck (H&N) region is challenging due to low contrast between adjacent soft tissue. The SegRap 2023 challenge has been focused on benchmarking the segmentation algorithms of Nasopharyngeal Carcinoma (NPC) which would be employed as auto-contouring tools for radiation treatment planning purposes. We propose a fully-automatic framework and develop two models for a) segmentation of 45 Organs at Risk (OARs) and b) two Gross Tumor Volumes (GTVs). To this end, we preprocess the image volumes by harmonizing the intensity distributions and then automatically cropping the volumes around the target regions. The preprocessed volumes were employed to train a standard 3D U-Net model for each task, separately. Our method took second place for each of the tasks in the validation phase of the challenge. The proposed framework is available at https://github.com/Astarakee/segrap2023
PriorNet: lesion segmentation in PET-CT including prior tumor appearance information
Oct 05, 2022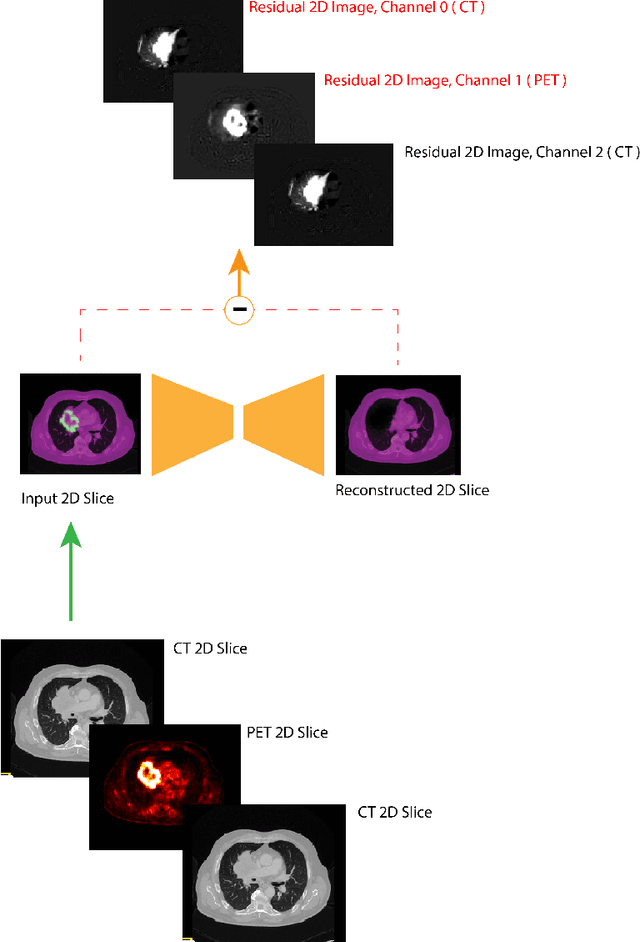
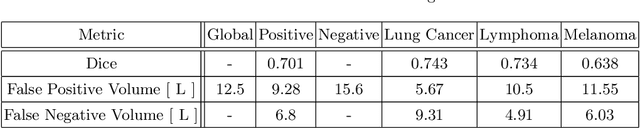

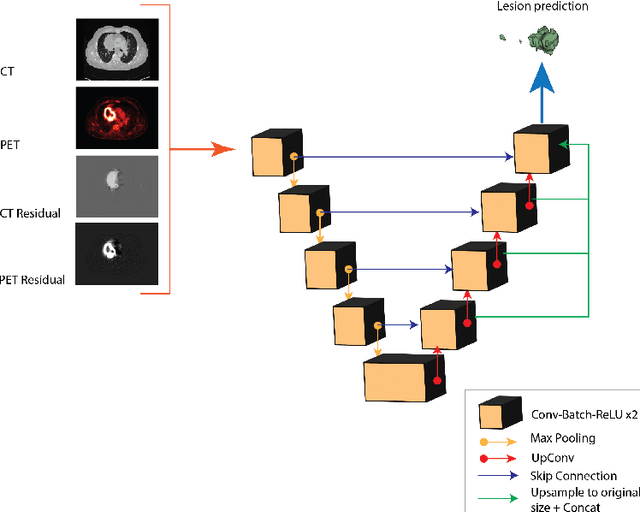
Abstract:Tumor segmentation in PET-CT images is challenging due to the dual nature of the acquired information: low metabolic information in CT and low spatial resolution in PET. U-Net architecture is the most common and widely recognized approach when developing a fully automatic image segmentation method in the medical field. We proposed a two-step approach, aiming to refine and improve the segmentation performances of tumoral lesions in PET-CT. The first step generates a prior tumor appearance map from the PET-CT volumes, regarded as prior tumor information. The second step, consisting of a standard U-Net, receives the prior tumor appearance map and PET-CT images to generate the lesion mask. We evaluated the method on the 1014 cases available for the AutoPET 2022 challenge, and the results showed an average Dice score of 0.701 on the positive cases.
Development and evaluation of a 3D annotation software for interactive COVID-19 lesion segmentation in chest CT
Jan 13, 2021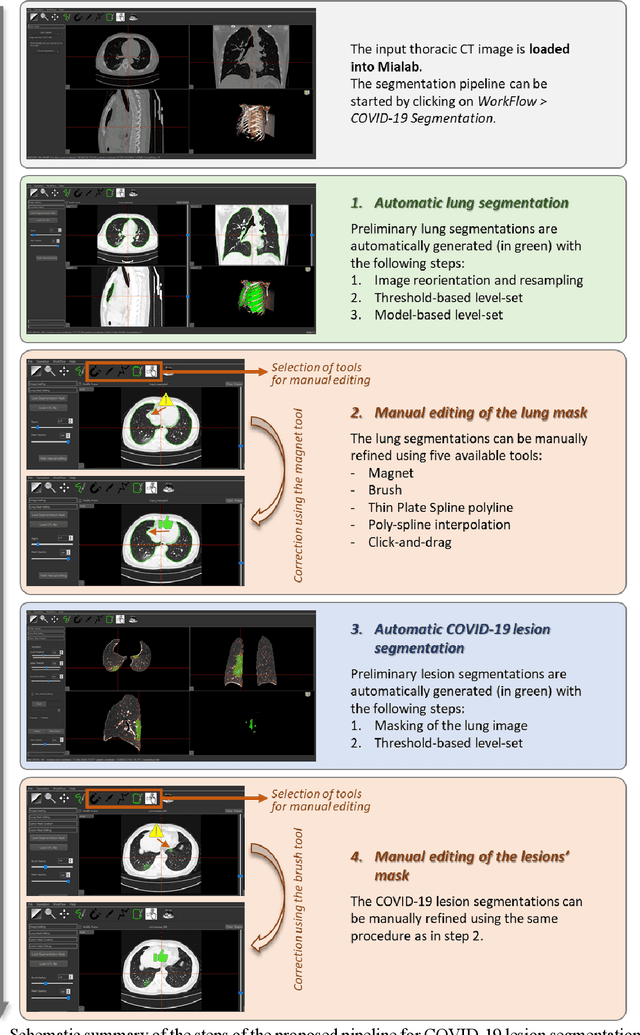
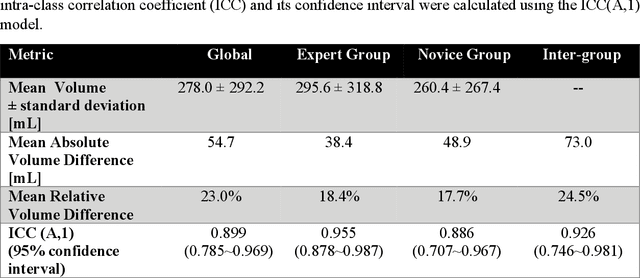
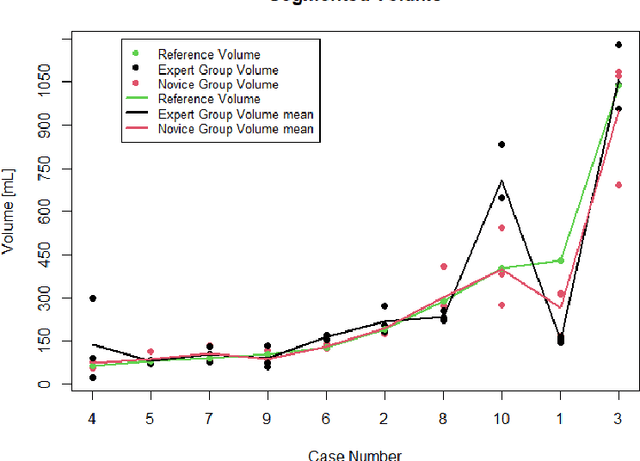
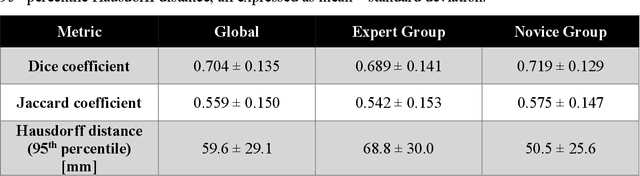
Abstract:Segmentation of COVID-19 lesions from chest CT scans is of great importance for better diagnosing the disease and investigating its extent. However, manual segmentation can be very time consuming and subjective, given the lesions' large variation in shape, size and position. On the other hand, we still lack large manually segmented datasets that could be used for training machine learning-based models for fully automatic segmentation. In this work, we propose a new interactive and user-friendly tool for COVID-19 lesion segmentation, which works by alternating automatic steps (based on level-set segmentation and statistical shape modeling) with manual correction steps. The present software was tested by two different expertise groups: one group of three radiologists and one of three users with an engineering background. Promising segmentation results were obtained by both groups, which achieved satisfactory agreement both between- and within-group. Moreover, our interactive tool was shown to significantly speed up the lesion segmentation process, when compared to fully manual segmentation. Finally, we investigated inter-observer variability and how it is strongly influenced by several subjective factors, showing the importance for AI researchers and clinical doctors to be aware of the uncertainty in lesion segmentation results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge