Rodrigo Moreno
KTH Royal Institute of Technology, Department of Biomedical Engineering and Health Systems, Stockholm, Sweden
Scalable Low-Density Distributed Manipulation Using an Interconnected Actuator Array
Feb 23, 2026Abstract:Distributed Manipulator Systems, composed of arrays of robotic actuators necessitate dense actuator arrays to effectively manipulate small objects. This paper presents a system composed of modular 3-DoF robotic tiles interconnected by a compliant surface layer, forming a continuous, controllable manipulation surface. The compliant layer permits increased actuator spacing without compromising object manipulation capabilities, significantly reducing actuator density while maintaining robust control, even for smaller objects. We characterize the coupled workspace of the array and develop a manipulation strategy capable of translating objects to arbitrary positions within an N X N array. The approach is validated experimentally using a minimal 2 X 2 prototype, demonstrating the successful manipulation of objects with varied shapes and sizes.
Anatomy-Aware Lymphoma Lesion Detection in Whole-Body PET/CT
Nov 10, 2025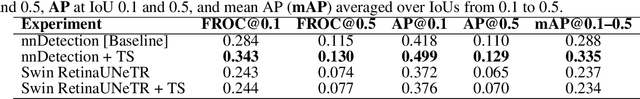
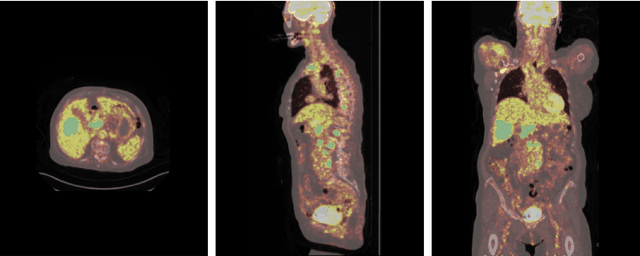
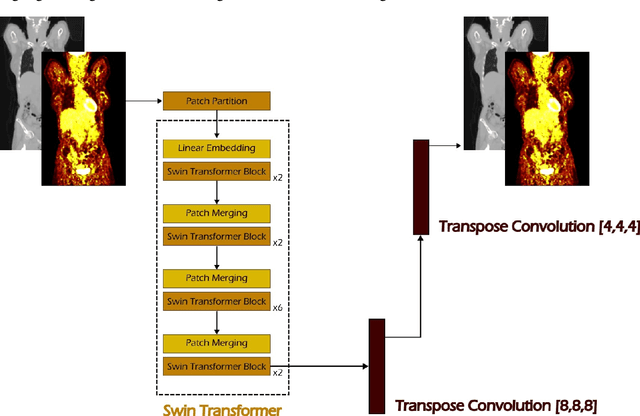
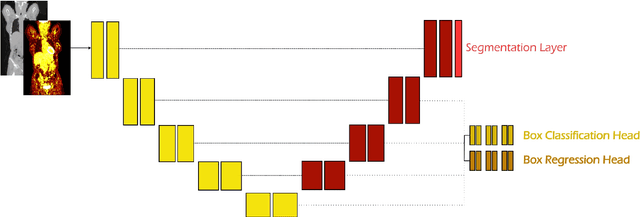
Abstract:Early cancer detection is crucial for improving patient outcomes, and 18F FDG PET/CT imaging plays a vital role by combining metabolic and anatomical information. Accurate lesion detection remains challenging due to the need to identify multiple lesions of varying sizes. In this study, we investigate the effect of adding anatomy prior information to deep learning-based lesion detection models. In particular, we add organ segmentation masks from the TotalSegmentator tool as auxiliary inputs to provide anatomical context to nnDetection, which is the state-of-the-art for lesion detection, and Swin Transformer. The latter is trained in two stages that combine self-supervised pre-training and supervised fine-tuning. The method is tested in the AutoPET and Karolinska lymphoma datasets. The results indicate that the inclusion of anatomical priors substantially improves the detection performance within the nnDetection framework, while it has almost no impact on the performance of the vision transformer. Moreover, we observe that Swin Transformer does not offer clear advantages over conventional convolutional neural network (CNN) encoders used in nnDetection. These findings highlight the critical role of the anatomical context in cancer lesion detection, especially in CNN-based models.
Flexible and Foldable: Workspace Analysis and Object Manipulation Using a Soft, Interconnected, Origami-Inspired Actuator Array
Sep 17, 2025Abstract:Object manipulation is a fundamental challenge in robotics, where systems must balance trade-offs among manipulation capabilities, system complexity, and throughput. Distributed manipulator systems (DMS) use the coordinated motion of actuator arrays to perform complex object manipulation tasks, seeing widespread exploration within the literature and in industry. However, existing DMS designs typically rely on high actuator densities and impose constraints on object-to-actuator scale ratios, limiting their adaptability. We present a novel DMS design utilizing an array of 3-DoF, origami-inspired robotic tiles interconnected by a compliant surface layer. Unlike conventional DMS, our approach enables manipulation not only at the actuator end effectors but also across a flexible surface connecting all actuators; creating a continuous, controllable manipulation surface. We analyse the combined workspace of such a system, derive simple motion primitives, and demonstrate its capabilities to translate simple geometric objects across an array of tiles. By leveraging the inter-tile connective material, our approach significantly reduces actuator density, increasing the area over which an object can be manipulated by x1.84 without an increase in the number of actuators. This design offers a lower cost and complexity alternative to traditional high-density arrays, and introduces new opportunities for manipulation strategies that leverage the flexibility of the interconnected surface.
Autonomous nanoparticle synthesis by design
May 19, 2025Abstract:Controlled synthesis of materials with specified atomic structures underpins technological advances yet remains reliant on iterative, trial-and-error approaches. Nanoparticles (NPs), whose atomic arrangement dictates their emergent properties, are particularly challenging to synthesise due to numerous tunable parameters. Here, we introduce an autonomous approach explicitly targeting synthesis of atomic-scale structures. Our method autonomously designs synthesis protocols by matching real time experimental total scattering (TS) and pair distribution function (PDF) data to simulated target patterns, without requiring prior synthesis knowledge. We demonstrate this capability at a synchrotron, successfully synthesising two structurally distinct gold NPs: 5 nm decahedral and 10 nm face-centred cubic structures. Ultimately, specifying a simulated target scattering pattern, thus representing a bespoke atomic structure, and obtaining both the synthesised material and its reproducible synthesis protocol on demand may revolutionise materials design. Thus, ScatterLab provides a generalisable blueprint for autonomous, atomic structure-targeted synthesis across diverse systems and applications.
Synthesizing Individualized Aging Brains in Health and Disease with Generative Models and Parallel Transport
Feb 28, 2025



Abstract:Simulating prospective magnetic resonance imaging (MRI) scans from a given individual brain image is challenging, as it requires accounting for canonical changes in aging and/or disease progression while also considering the individual brain's current status and unique characteristics. While current deep generative models can produce high-resolution anatomically accurate templates for population-wide studies, their ability to predict future aging trajectories for individuals remains limited, particularly in capturing subject-specific neuroanatomical variations over time. In this study, we introduce Individualized Brain Synthesis (InBrainSyn), a framework for synthesizing high-resolution subject-specific longitudinal MRI scans that simulate neurodegeneration in both Alzheimer's disease (AD) and normal aging. InBrainSyn uses a parallel transport algorithm to adapt the population-level aging trajectories learned by a generative deep template network, enabling individualized aging synthesis. As InBrainSyn uses diffeomorphic transformations to simulate aging, the synthesized images are topologically consistent with the original anatomy by design. We evaluated InBrainSyn both quantitatively and qualitatively on AD and healthy control cohorts from the Open Access Series of Imaging Studies - version 3 dataset. Experimentally, InBrainSyn can also model neuroanatomical transitions between normal aging and AD. An evaluation of an external set supports its generalizability. Overall, with only a single baseline scan, InBrainSyn synthesizes realistic 3D spatiotemporal T1w MRI scans, producing personalized longitudinal aging trajectories. The code for InBrainSyn is available at: https://github.com/Fjr9516/InBrainSyn.
Neural Cellular Automata for Decentralized Sensing using a Soft Inductive Sensor Array for Distributed Manipulator Systems
Feb 03, 2025Abstract:In Distributed Manipulator Systems (DMS), decentralization is a highly desirable property as it promotes robustness and facilitates scalability by distributing computational burden and eliminating singular points of failure. However, current DMS typically utilize a centralized approach to sensing, such as single-camera computer vision systems. This centralization poses a risk to system reliability and offers a significant limiting factor to system size. In this work, we introduce a decentralized approach for sensing and in a Distributed Manipulator Systems using Neural Cellular Automata (NCA). Demonstrating a decentralized sensing in a hardware implementation, we present a novel inductive sensor board designed for distributed sensing and evaluate its ability to estimate global object properties, such as the geometric center, through local interactions and computations. Experiments demonstrate that NCA-based sensing networks accurately estimate object position at 0.24 times the inter sensor distance. They maintain resilience under sensor faults and noise, and scale seamlessly across varying network sizes. These findings underscore the potential of local, decentralized computations to enable scalable, fault-tolerant, and noise-resilient object property estimation in DMS
Unsupervised Domain Adaptation for Pediatric Brain Tumor Segmentation
Jun 24, 2024



Abstract:Significant advances have been made toward building accurate automatic segmentation models for adult gliomas. However, the performance of these models often degrades when applied to pediatric glioma due to their imaging and clinical differences (domain shift). Obtaining sufficient annotated data for pediatric glioma is typically difficult because of its rare nature. Also, manual annotations are scarce and expensive. In this work, we propose Domain-Adapted nnU-Net (DA-nnUNet) to perform unsupervised domain adaptation from adult glioma (source domain) to pediatric glioma (target domain). Specifically, we add a domain classifier connected with a gradient reversal layer (GRL) to a backbone nnU-Net. Once the classifier reaches a very high accuracy, the GRL is activated with the goal of transferring domain-invariant features from the classifier to the segmentation model while preserving segmentation accuracy on the source domain. The accuracy of the classifier slowly degrades to chance levels. No annotations are used in the target domain. The method is compared to 8 different supervised models using BraTS-Adult glioma (N=1251) and BraTS-PED glioma data (N=99). The proposed method shows notable performance enhancements in the tumor core (TC) region compared to the model that only uses adult data: ~32% better Dice scores and ~20 better 95th percentile Hausdorff distances. Moreover, our unsupervised approach shows no statistically significant difference compared to the practical upper bound model using manual annotations from both datasets in TC region. The code is shared at https://github.com/Fjr9516/DA_nnUNet.
A deformation-based morphometry framework for disentangling Alzheimer's disease from normal aging using learned normal aging templates
Nov 14, 2023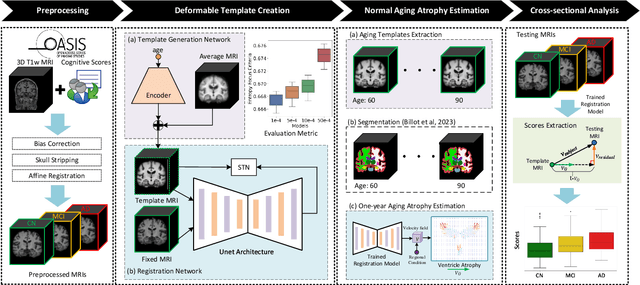
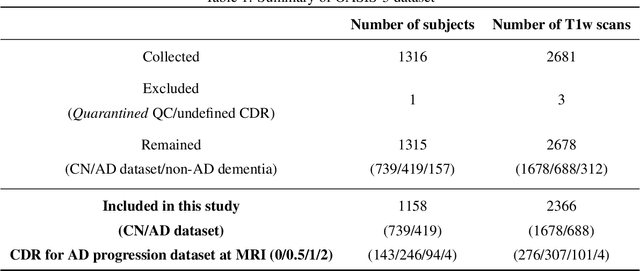
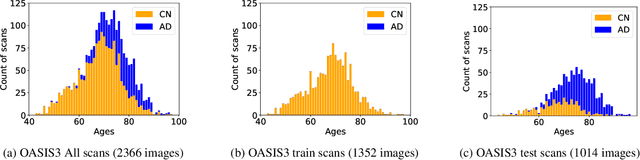
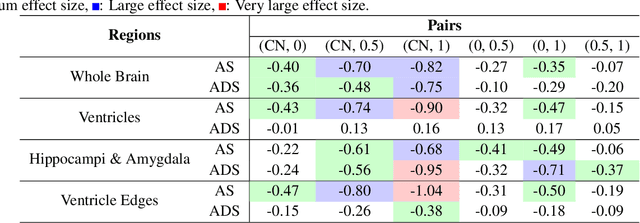
Abstract:Alzheimer's Disease and normal aging are both characterized by brain atrophy. The question of whether AD-related brain atrophy represents accelerated aging or a neurodegeneration process distinct from that in normal aging remains unresolved. Moreover, precisely disentangling AD-related brain atrophy from normal aging in a clinical context is complex. In this study, we propose a deformation-based morphometry framework to estimate normal aging and AD-specific atrophy patterns of subjects from morphological MRI scans. We first leverage deep-learning-based methods to create age-dependent templates of cognitively normal (CN) subjects. These templates model the normal aging atrophy patterns in a CN population. Then, we use the learned diffeomorphic registration to estimate the one-year normal aging pattern at the voxel level. We register the testing image to the 60-year-old CN template in the second step. Finally, normal aging and AD-specific scores are estimated by measuring the alignment of this registration with the one-year normal aging pattern. The methodology was developed and evaluated on the OASIS3 dataset with 1,014 T1-weighted MRI scans. Of these, 326 scans were from CN subjects, and 688 scans were from individuals clinically diagnosed with AD at different stages of clinical severity defined by clinical dementia rating (CDR) scores. The results show that ventricles predominantly follow an accelerated normal aging pattern in subjects with AD. In turn, hippocampi and amygdala regions were affected by both normal aging and AD-specific factors. Interestingly, hippocampi and amygdala regions showed more of an accelerated normal aging pattern for subjects during the early clinical stages of the disease, while the AD-specific score increases in later clinical stages. Our code is freely available at https://github.com/Fjr9516/DBM_with_DL.
Merging multiple input descriptors and supervisors in a deep neural network for tractogram filtering
Jul 11, 2023Abstract:One of the main issues of the current tractography methods is their high false-positive rate. Tractogram filtering is an option to remove false-positive streamlines from tractography data in a post-processing step. In this paper, we train a deep neural network for filtering tractography data in which every streamline of a tractogram is classified as {\em plausible, implausible}, or {\em inconclusive}. For this, we use four different tractogram filtering strategies as supervisors: TractQuerier, RecobundlesX, TractSeg, and an anatomy-inspired filter. Their outputs are combined to obtain the classification labels for the streamlines. We assessed the importance of different types of information along the streamlines for performing this classification task, including the coordinates of the streamlines, diffusion data, landmarks, T1-weighted information, and a brain parcellation. We found that the streamline coordinates are the most relevant followed by the diffusion data in this particular classification task.
Generative Aging of Brain Images with Diffeomorphic Registration
May 31, 2022



Abstract:Analyzing and predicting brain aging is essential for early prognosis and accurate diagnosis of cognitive diseases. The technique of neuroimaging, such as Magnetic Resonance Imaging (MRI), provides a noninvasive means of observing the aging process within the brain. With longitudinal image data collection, data-intensive Artificial Intelligence (AI) algorithms have been used to examine brain aging. However, existing state-of-the-art algorithms tend to be restricted to group-level predictions and suffer from unreal predictions. This paper proposes a methodology for generating longitudinal MRI scans that capture subject-specific neurodegeneration and retain anatomical plausibility in aging. The proposed methodology is developed within the framework of diffeomorphic registration and relies on three key novel technological advances to generate subject-level anatomically plausible predictions: i) a computationally efficient and individualized generative framework based on registration; ii) an aging generative module based on biological linear aging progression; iii) a quality control module to fit registration for generation task. Our methodology was evaluated on 2662 T1-weighted (T1-w) MRI scans from 796 participants from three different cohorts. First, we applied 6 commonly used criteria to demonstrate the aging simulation ability of the proposed methodology; Secondly, we evaluated the quality of the synthetic images using quantitative measurements and qualitative assessment by a neuroradiologist. Overall, the experimental results show that the proposed method can produce anatomically plausible predictions that can be used to enhance longitudinal datasets, in turn enabling data-hungry AI-driven healthcare tools.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge