Jiangshan Lu
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
CDDSA: Contrastive Domain Disentanglement and Style Augmentation for Generalizable Medical Image Segmentation
Nov 22, 2022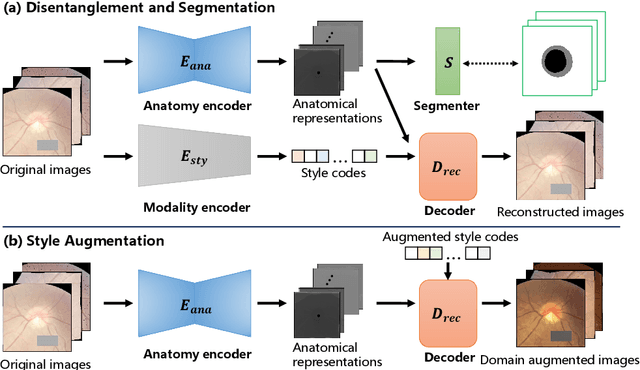
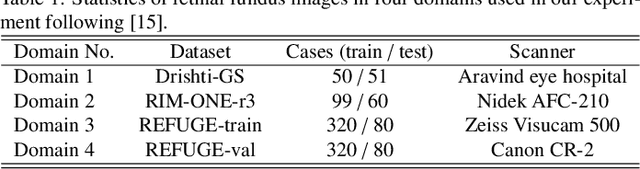
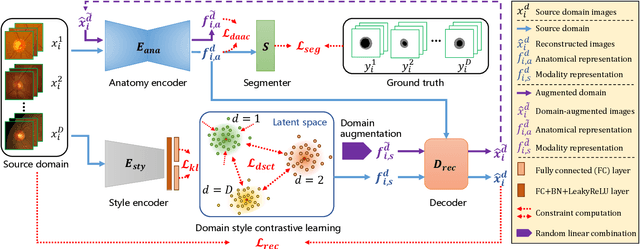

Abstract:Generalization to previously unseen images with potential domain shifts and different styles is essential for clinically applicable medical image segmentation, and the ability to disentangle domain-specific and domain-invariant features is key for achieving Domain Generalization (DG). However, existing DG methods can hardly achieve effective disentanglement to get high generalizability. To deal with this problem, we propose an efficient Contrastive Domain Disentanglement and Style Augmentation (CDDSA) framework for generalizable medical image segmentation. First, a disentangle network is proposed to decompose an image into a domain-invariant anatomical representation and a domain-specific style code, where the former is sent to a segmentation model that is not affected by the domain shift, and the disentangle network is regularized by a decoder that combines the anatomical and style codes to reconstruct the input image. Second, to achieve better disentanglement, a contrastive loss is proposed to encourage the style codes from the same domain and different domains to be compact and divergent, respectively. Thirdly, to further improve generalizability, we propose a style augmentation method based on the disentanglement representation to synthesize images in various unseen styles with shared anatomical structures. Our method was validated on a public multi-site fundus image dataset for optic cup and disc segmentation and an in-house multi-site Nasopharyngeal Carcinoma Magnetic Resonance Image (NPC-MRI) dataset for nasopharynx Gross Tumor Volume (GTVnx) segmentation. Experimental results showed that the proposed CDDSA achieved remarkable generalizability across different domains, and it outperformed several state-of-the-art methods in domain-generalizable segmentation.
Contrastive Domain Disentanglement for Generalizable Medical Image Segmentation
May 13, 2022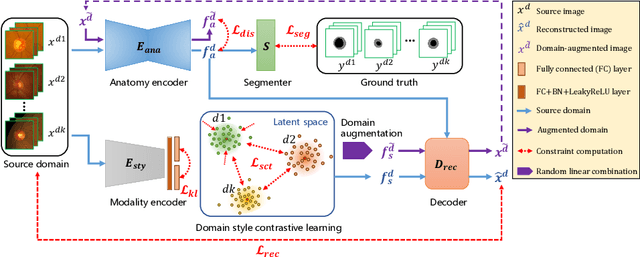
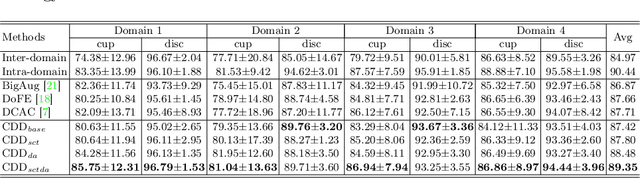
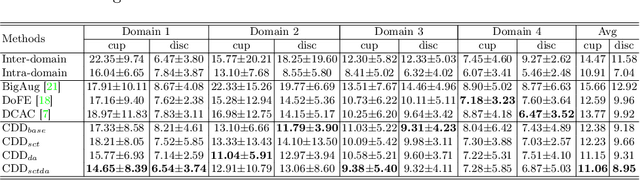
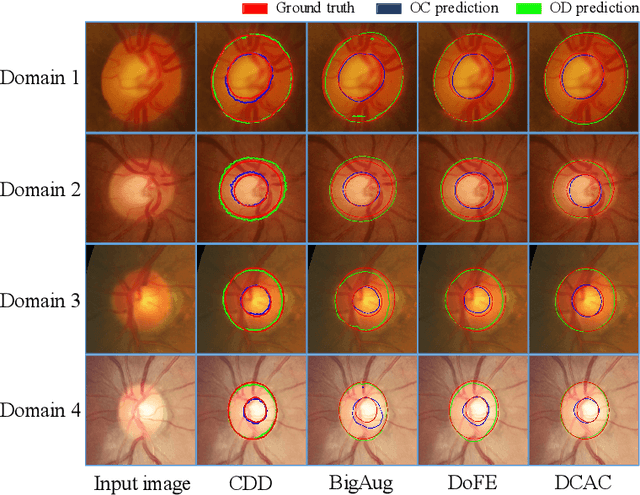
Abstract:Efficiently utilizing discriminative features is crucial for convolutional neural networks to achieve remarkable performance in medical image segmentation and is also important for model generalization across multiple domains, where letting model recognize domain-specific and domain-invariant information among multi-site datasets is a reasonable strategy for domain generalization. Unfortunately, most of the recent disentangle networks are not directly adaptable to unseen-domain datasets because of the limitations of offered data distribution. To tackle this deficiency, we propose Contrastive Domain Disentangle (CDD) network for generalizable medical image segmentation. We first introduce a disentangle network to decompose medical images into an anatomical representation factor and a modality representation factor. Then, a style contrastive loss is proposed to encourage the modality representations from the same domain to distribute as close as possible while different domains are estranged from each other. Finally, we propose a domain augmentation strategy that can randomly generate new domains for model generalization training. Experimental results on multi-site fundus image datasets for optic cup and disc segmentation show that the CDD has good model generalization. Our proposed CDD outperforms several state-of-the-art methods in domain generalizable segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge