Yunzhi Huang
Non-Adversarial Learning: Vector-Quantized Common Latent Space for Multi-Sequence MRI
Jul 03, 2024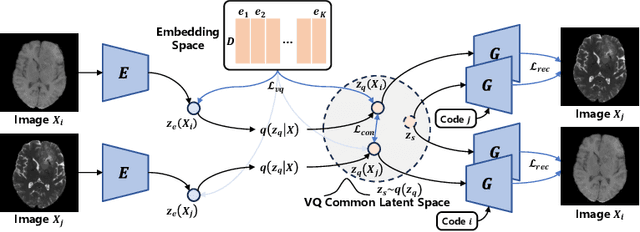
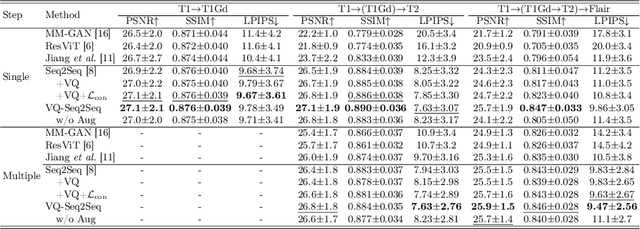
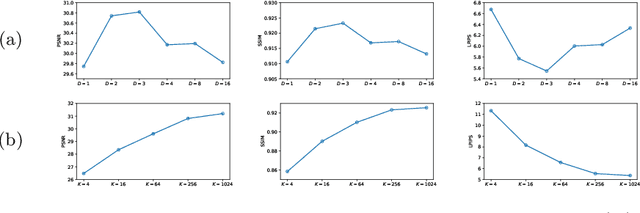

Abstract:Adversarial learning helps generative models translate MRI from source to target sequence when lacking paired samples. However, implementing MRI synthesis with adversarial learning in clinical settings is challenging due to training instability and mode collapse. To address this issue, we leverage intermediate sequences to estimate the common latent space among multi-sequence MRI, enabling the reconstruction of distinct sequences from the common latent space. We propose a generative model that compresses discrete representations of each sequence to estimate the Gaussian distribution of vector-quantized common (VQC) latent space between multiple sequences. Moreover, we improve the latent space consistency with contrastive learning and increase model stability by domain augmentation. Experiments using BraTS2021 dataset show that our non-adversarial model outperforms other GAN-based methods, and VQC latent space aids our model to achieve (1) anti-interference ability, which can eliminate the effects of noise, bias fields, and artifacts, and (2) solid semantic representation ability, with the potential of one-shot segmentation. Our code is publicly available.
An Explainable Deep Framework: Towards Task-Specific Fusion for Multi-to-One MRI Synthesis
Jul 03, 2023Abstract:Multi-sequence MRI is valuable in clinical settings for reliable diagnosis and treatment prognosis, but some sequences may be unusable or missing for various reasons. To address this issue, MRI synthesis is a potential solution. Recent deep learning-based methods have achieved good performance in combining multiple available sequences for missing sequence synthesis. Despite their success, these methods lack the ability to quantify the contributions of different input sequences and estimate the quality of generated images, making it hard to be practical. Hence, we propose an explainable task-specific synthesis network, which adapts weights automatically for specific sequence generation tasks and provides interpretability and reliability from two sides: (1) visualize the contribution of each input sequence in the fusion stage by a trainable task-specific weighted average module; (2) highlight the area the network tried to refine during synthesizing by a task-specific attention module. We conduct experiments on the BraTS2021 dataset of 1251 subjects, and results on arbitrary sequence synthesis indicate that the proposed method achieves better performance than the state-of-the-art methods. Our code is available at \url{https://github.com/fiy2W/mri_seq2seq}.
GSMorph: Gradient Surgery for cine-MRI Cardiac Deformable Registration
Jun 26, 2023Abstract:Deep learning-based deformable registration methods have been widely investigated in diverse medical applications. Learning-based deformable registration relies on weighted objective functions trading off registration accuracy and smoothness of the deformation field. Therefore, they inevitably require tuning the hyperparameter for optimal registration performance. Tuning the hyperparameters is highly computationally expensive and introduces undesired dependencies on domain knowledge. In this study, we construct a registration model based on the gradient surgery mechanism, named GSMorph, to achieve a hyperparameter-free balance on multiple losses. In GSMorph, we reformulate the optimization procedure by projecting the gradient of similarity loss orthogonally to the plane associated with the smoothness constraint, rather than additionally introducing a hyperparameter to balance these two competing terms. Furthermore, our method is model-agnostic and can be merged into any deep registration network without introducing extra parameters or slowing down inference. In this study, We compared our method with state-of-the-art (SOTA) deformable registration approaches over two publicly available cardiac MRI datasets. GSMorph proves superior to five SOTA learning-based registration models and two conventional registration techniques, SyN and Demons, on both registration accuracy and smoothness.
Synthesis-based Imaging-Differentiation Representation Learning for Multi-Sequence 3D/4D MRI
Feb 01, 2023



Abstract:Multi-sequence MRIs can be necessary for reliable diagnosis in clinical practice due to the complimentary information within sequences. However, redundant information exists across sequences, which interferes with mining efficient representations by modern machine learning or deep learning models. To handle various clinical scenarios, we propose a sequence-to-sequence generation framework (Seq2Seq) for imaging-differentiation representation learning. In this study, not only do we propose arbitrary 3D/4D sequence generation within one model to generate any specified target sequence, but also we are able to rank the importance of each sequence based on a new metric estimating the difficulty of a sequence being generated. Furthermore, we also exploit the generation inability of the model to extract regions that contain unique information for each sequence. We conduct extensive experiments using three datasets including a toy dataset of 20,000 simulated subjects, a brain MRI dataset of 1,251 subjects, and a breast MRI dataset of 2,101 subjects, to demonstrate that (1) our proposed Seq2Seq is efficient and lightweight for complex clinical datasets and can achieve excellent image quality; (2) top-ranking sequences can be used to replace complete sequences with non-inferior performance; (3) combining MRI with our imaging-differentiation map leads to better performance in clinical tasks such as glioblastoma MGMT promoter methylation status prediction and breast cancer pathological complete response status prediction. Our code is available at https://github.com/fiy2W/mri_seq2seq.
Unsupervised Cross-Modality Domain Adaptation for Vestibular Schwannoma Segmentation and Koos Grade Prediction based on Semi-Supervised Contrastive Learning
Oct 09, 2022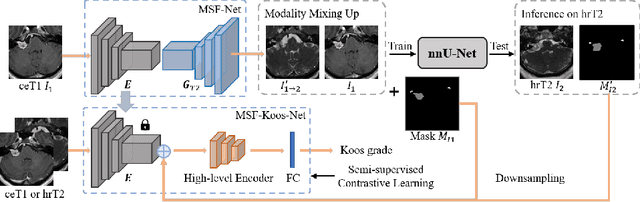
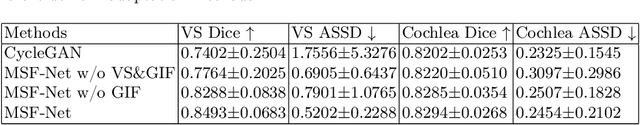

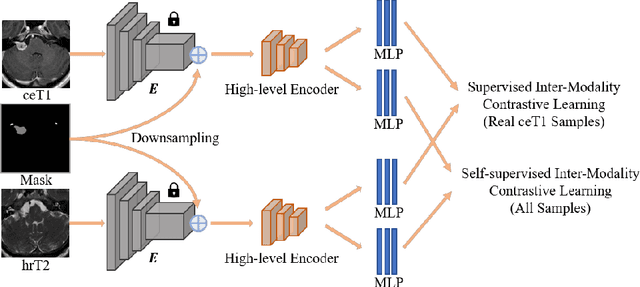
Abstract:Domain adaptation has been widely adopted to transfer styles across multi-vendors and multi-centers, as well as to complement the missing modalities. In this challenge, we proposed an unsupervised domain adaptation framework for cross-modality vestibular schwannoma (VS) and cochlea segmentation and Koos grade prediction. We learn the shared representation from both ceT1 and hrT2 images and recover another modality from the latent representation, and we also utilize proxy tasks of VS segmentation and brain parcellation to restrict the consistency of image structures in domain adaptation. After generating missing modalities, the nnU-Net model is utilized for VS and cochlea segmentation, while a semi-supervised contrastive learning pre-train approach is employed to improve the model performance for Koos grade prediction. On CrossMoDA validation phase Leaderboard, our method received rank 4 in task1 with a mean Dice score of 0.8394 and rank 2 in task2 with Macro-Average Mean Square Error of 0.3941. Our code is available at https://github.com/fiy2W/cmda2022.superpolymerization.
Longitudinal Prediction of Postnatal Brain Magnetic Resonance Images via a Metamorphic Generative Adversarial Network
Aug 09, 2022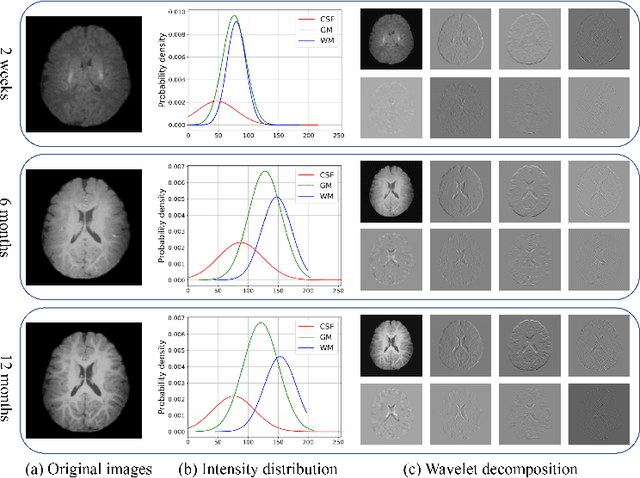

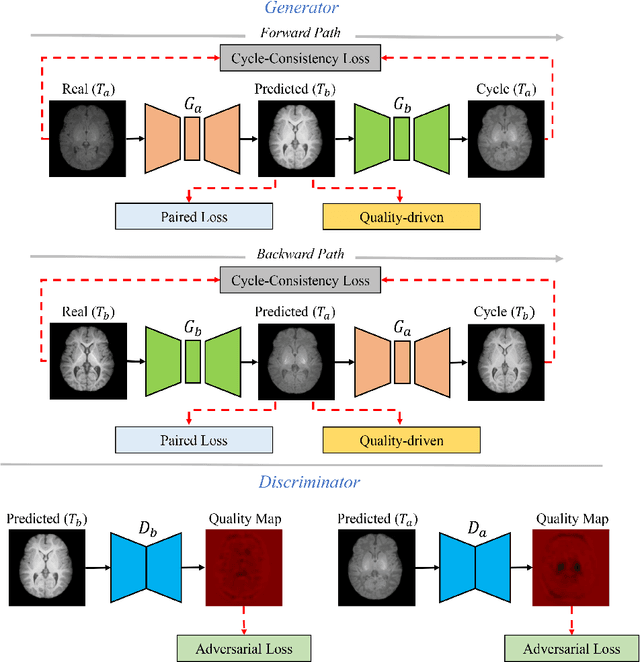
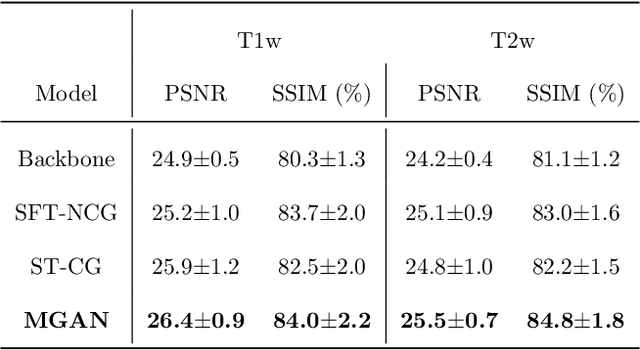
Abstract:Missing scans are inevitable in longitudinal studies due to either subject dropouts or failed scans. In this paper, we propose a deep learning framework to predict missing scans from acquired scans, catering to longitudinal infant studies. Prediction of infant brain MRI is challenging owing to the rapid contrast and structural changes particularly during the first year of life. We introduce a trustworthy metamorphic generative adversarial network (MGAN) for translating infant brain MRI from one time-point to another. MGAN has three key features: (i) Image translation leveraging spatial and frequency information for detail-preserving mapping; (ii) Quality-guided learning strategy that focuses attention on challenging regions. (iii) Multi-scale hybrid loss function that improves translation of tissue contrast and structural details. Experimental results indicate that MGAN outperforms existing GANs by accurately predicting both contrast and anatomical details.
Localizing the Recurrent Laryngeal Nerve via Ultrasound with a Bayesian Shape Framework
Jun 30, 2022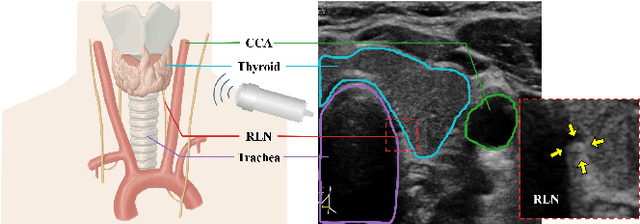
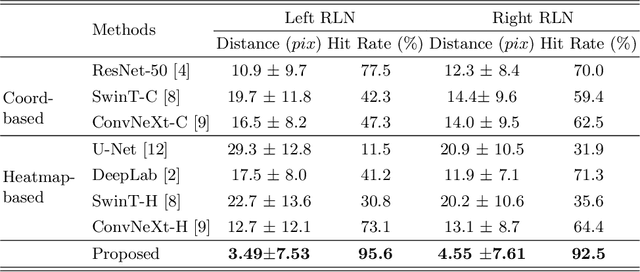
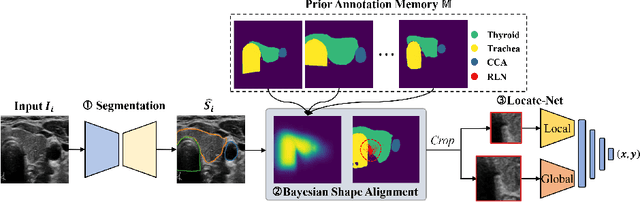
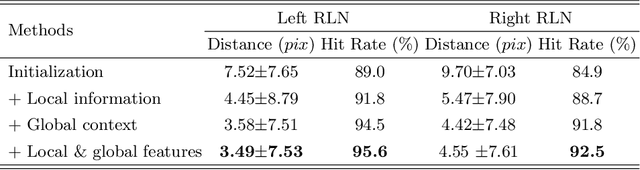
Abstract:Tumor infiltration of the recurrent laryngeal nerve (RLN) is a contraindication for robotic thyroidectomy and can be difficult to detect via standard laryngoscopy. Ultrasound (US) is a viable alternative for RLN detection due to its safety and ability to provide real-time feedback. However, the tininess of the RLN, with a diameter typically less than 3mm, poses significant challenges to the accurate localization of the RLN. In this work, we propose a knowledge-driven framework for RLN localization, mimicking the standard approach surgeons take to identify the RLN according to its surrounding organs. We construct a prior anatomical model based on the inherent relative spatial relationships between organs. Through Bayesian shape alignment (BSA), we obtain the candidate coordinates of the center of a region of interest (ROI) that encloses the RLN. The ROI allows a decreased field of view for determining the refined centroid of the RLN using a dual-path identification network, based on multi-scale semantic information. Experimental results indicate that the proposed method achieves superior hit rates and substantially smaller distance errors compared with state-of-the-art methods.
Learn2Reg: comprehensive multi-task medical image registration challenge, dataset and evaluation in the era of deep learning
Dec 23, 2021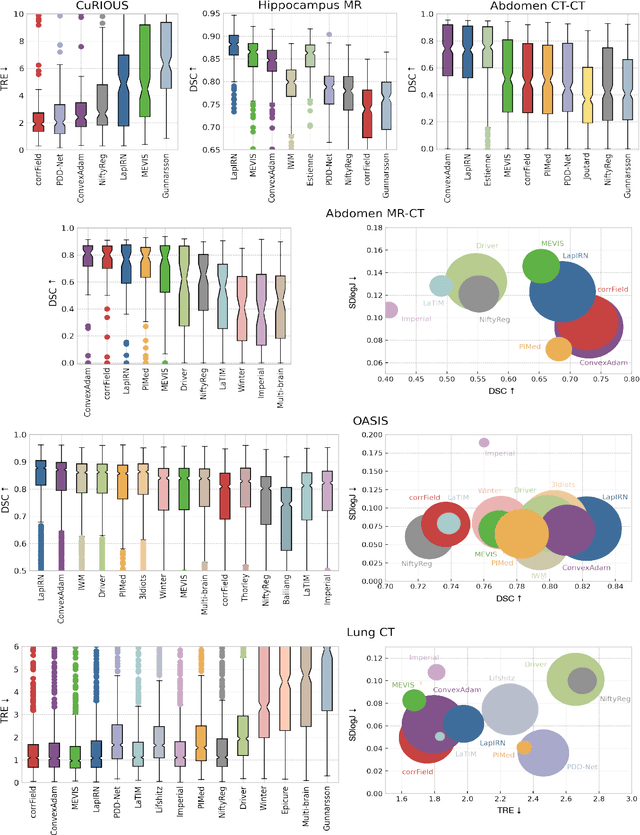
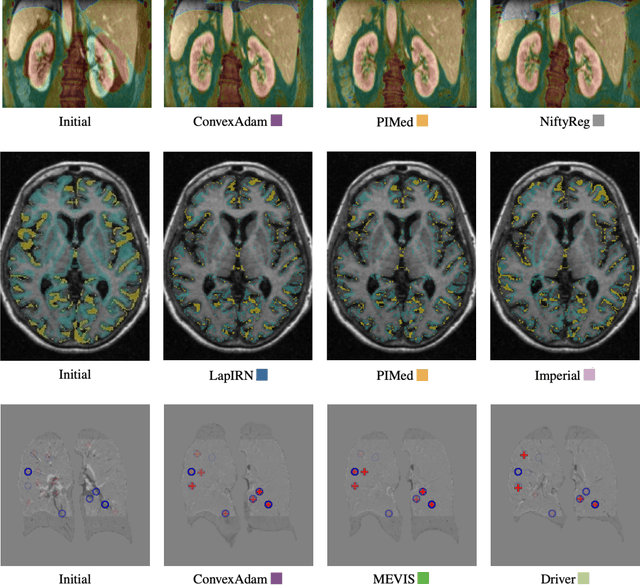
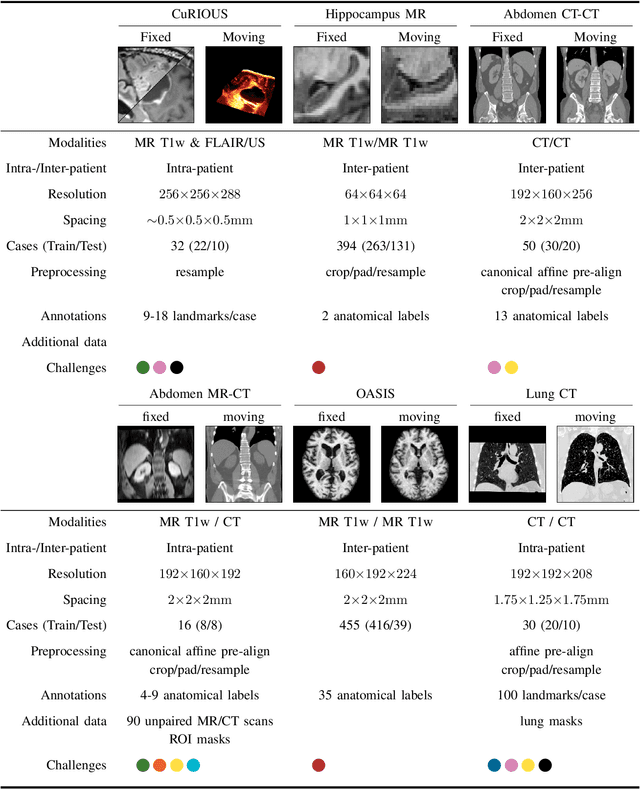
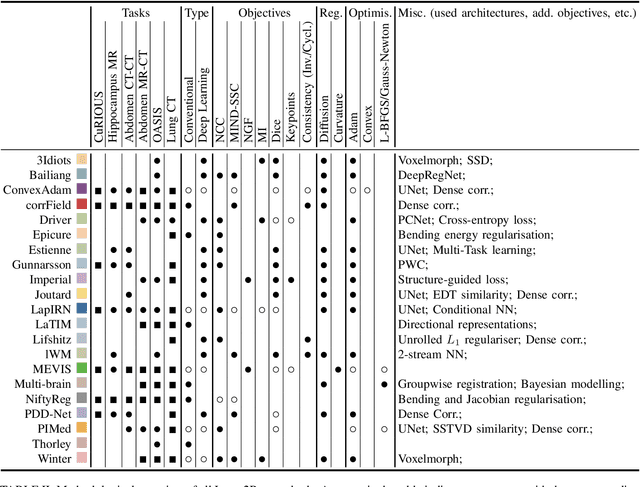
Abstract:Image registration is a fundamental medical image analysis task, and a wide variety of approaches have been proposed. However, only a few studies have comprehensively compared medical image registration approaches on a wide range of clinically relevant tasks, in part because of the lack of availability of such diverse data. This limits the development of registration methods, the adoption of research advances into practice, and a fair benchmark across competing approaches. The Learn2Reg challenge addresses these limitations by providing a multi-task medical image registration benchmark for comprehensive characterisation of deformable registration algorithms. A continuous evaluation will be possible at https://learn2reg.grand-challenge.org. Learn2Reg covers a wide range of anatomies (brain, abdomen, and thorax), modalities (ultrasound, CT, MR), availability of annotations, as well as intra- and inter-patient registration evaluation. We established an easily accessible framework for training and validation of 3D registration methods, which enabled the compilation of results of over 65 individual method submissions from more than 20 unique teams. We used a complementary set of metrics, including robustness, accuracy, plausibility, and runtime, enabling unique insight into the current state-of-the-art of medical image registration. This paper describes datasets, tasks, evaluation methods and results of the challenge, and the results of further analysis of transferability to new datasets, the importance of label supervision, and resulting bias.
Deformable Registration of Brain MR Images via a Hybrid Loss
Oct 28, 2021

Abstract:We learn a deformable registration model for T1-weighted MR images by considering multiple image characteristics via a hybrid loss. Our method registers the OASIS dataset with high accuracy while preserving deformation smoothness.
An Auto-Context Deformable Registration Network for Infant Brain MRI
May 19, 2020

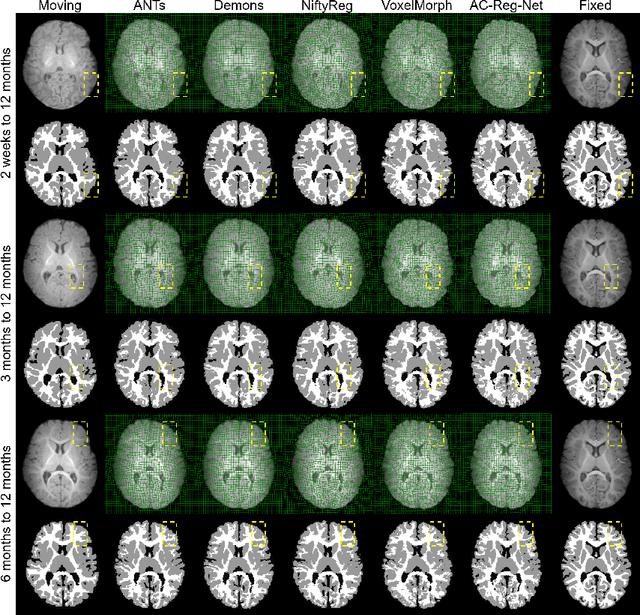
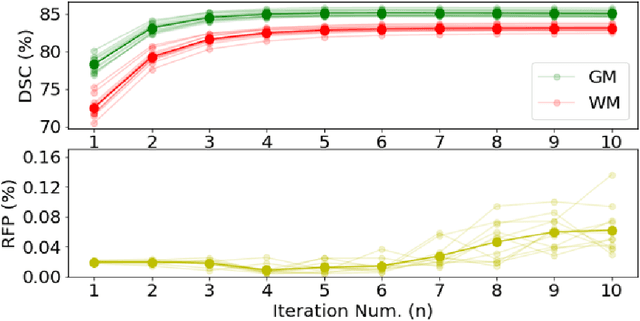
Abstract:Deformable image registration is fundamental to longitudinal and population analysis. Geometric alignment of the infant brain MR images is challenging, owing to rapid changes in image appearance in association with brain development. In this paper, we propose an infant-dedicated deep registration network that uses the auto-context strategy to gradually refine the deformation fields to obtain highly accurate correspondences. Instead of training multiple registration networks, our method estimates the deformation fields by invoking a single network multiple times for iterative deformation refinement. The final deformation field is obtained by the incremental composition of the deformation fields. Experimental results in comparison with state-of-the-art registration methods indicate that our method achieves higher accuracy while at the same time preserves the smoothness of the deformation fields. Our implementation is available online.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge