Zhengwang Wu
Senior Member, IEEE
Early Autism Diagnosis based on Path Signature and Siamese Unsupervised Feature Compressor
Jul 12, 2023



Abstract:Autism Spectrum Disorder (ASD) has been emerging as a growing public health threat. Early diagnosis of ASD is crucial for timely, effective intervention and treatment. However, conventional diagnosis methods based on communications and behavioral patterns are unreliable for children younger than 2 years of age. Given evidences of neurodevelopmental abnormalities in ASD infants, we resort to a novel deep learning-based method to extract key features from the inherently scarce, class-imbalanced, and heterogeneous structural MR images for early autism diagnosis. Specifically, we propose a Siamese verification framework to extend the scarce data, and an unsupervised compressor to alleviate data imbalance by extracting key features. We also proposed weight constraints to cope with sample heterogeneity by giving different samples different voting weights during validation, and we used Path Signature to unravel meaningful developmental features from the two-time point data longitudinally. Extensive experiments have shown that our method performed well under practical scenarios, transcending existing machine learning methods.
Longitudinal Prediction of Postnatal Brain Magnetic Resonance Images via a Metamorphic Generative Adversarial Network
Aug 09, 2022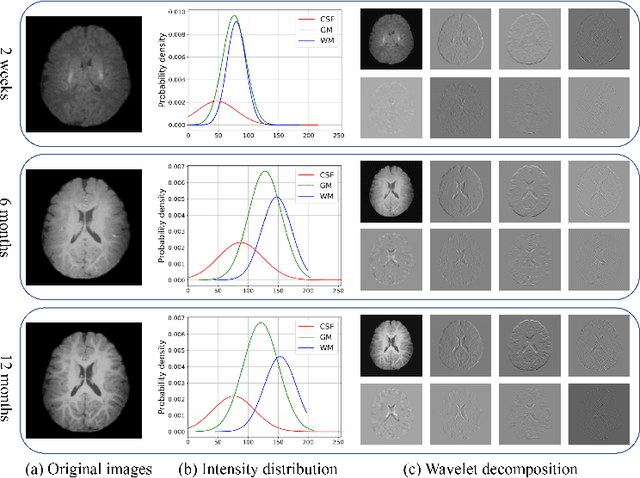

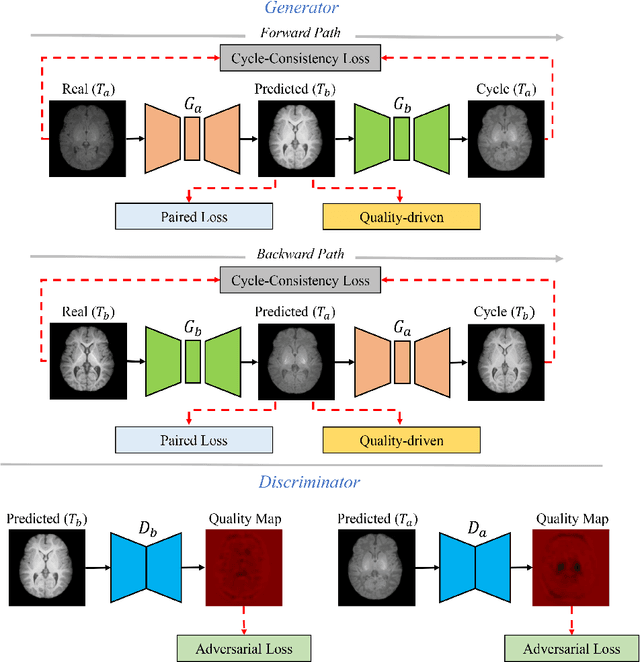
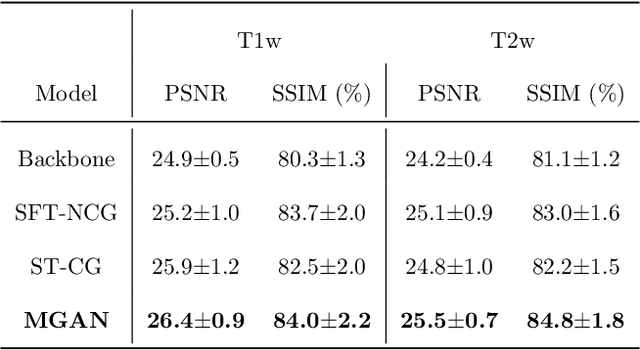
Abstract:Missing scans are inevitable in longitudinal studies due to either subject dropouts or failed scans. In this paper, we propose a deep learning framework to predict missing scans from acquired scans, catering to longitudinal infant studies. Prediction of infant brain MRI is challenging owing to the rapid contrast and structural changes particularly during the first year of life. We introduce a trustworthy metamorphic generative adversarial network (MGAN) for translating infant brain MRI from one time-point to another. MGAN has three key features: (i) Image translation leveraging spatial and frequency information for detail-preserving mapping; (ii) Quality-guided learning strategy that focuses attention on challenging regions. (iii) Multi-scale hybrid loss function that improves translation of tissue contrast and structural details. Experimental results indicate that MGAN outperforms existing GANs by accurately predicting both contrast and anatomical details.
Deep Modeling of Growth Trajectories for Longitudinal Prediction of Missing Infant Cortical Surfaces
Sep 12, 2020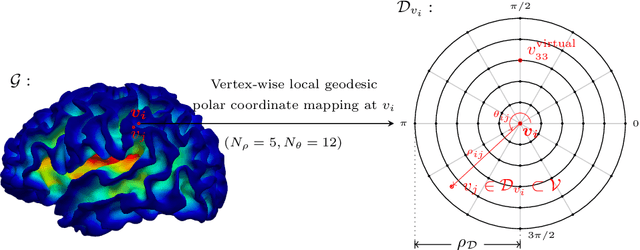
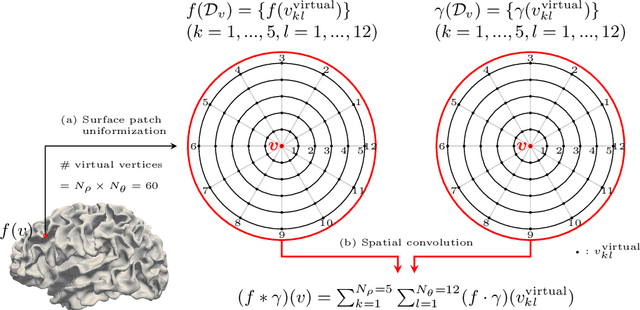
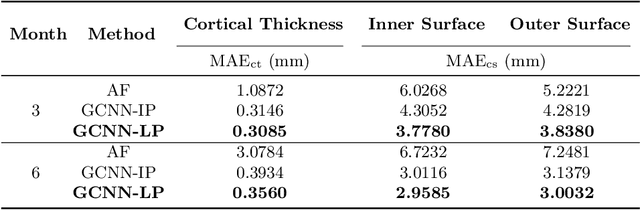
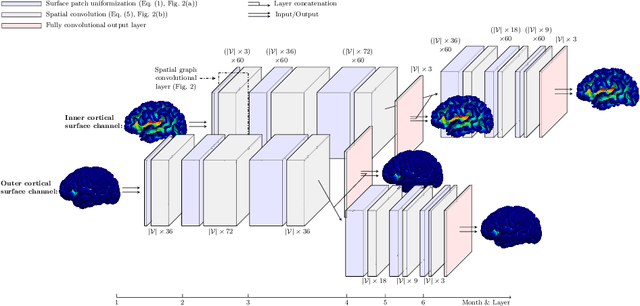
Abstract:Charting cortical growth trajectories is of paramount importance for understanding brain development. However, such analysis necessitates the collection of longitudinal data, which can be challenging due to subject dropouts and failed scans. In this paper, we will introduce a method for longitudinal prediction of cortical surfaces using a spatial graph convolutional neural network (GCNN), which extends conventional CNNs from Euclidean to curved manifolds. The proposed method is designed to model the cortical growth trajectories and jointly predict inner and outer cortical surfaces at multiple time points. Adopting a binary flag in loss calculation to deal with missing data, we fully utilize all available cortical surfaces for training our deep learning model, without requiring a complete collection of longitudinal data. Predicting the surfaces directly allows cortical attributes such as cortical thickness, curvature, and convexity to be computed for subsequent analysis. We will demonstrate with experimental results that our method is capable of capturing the nonlinearity of spatiotemporal cortical growth patterns and can predict cortical surfaces with improved accuracy.
Multi-Site Infant Brain Segmentation Algorithms: The iSeg-2019 Challenge
Jul 11, 2020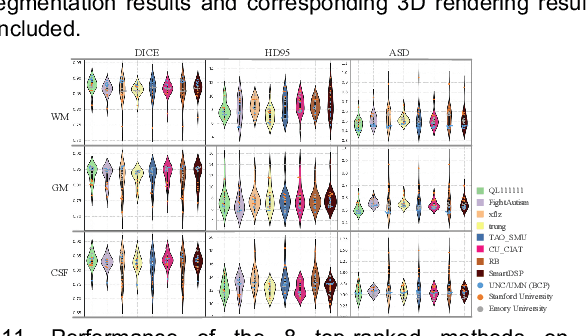
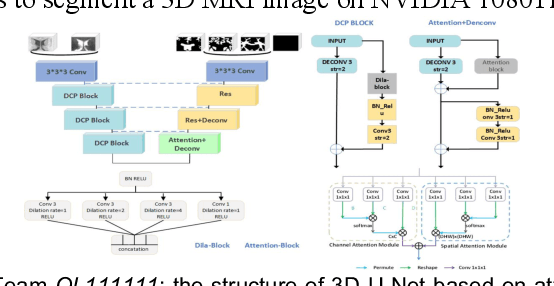
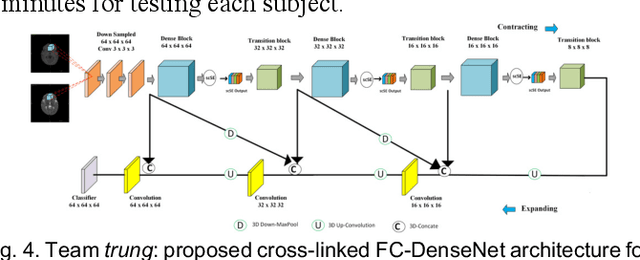
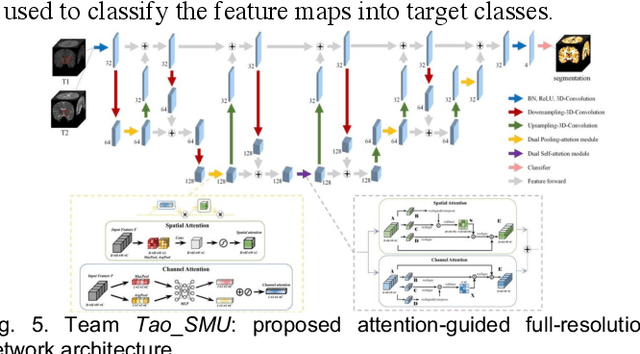
Abstract:To better understand early brain growth patterns in health and disorder, it is critical to accurately segment infant brain magnetic resonance (MR) images into white matter (WM), gray matter (GM), and cerebrospinal fluid (CSF). Deep learning-based methods have achieved state-of-the-art performance; however, one of major limitations is that the learning-based methods may suffer from the multi-site issue, that is, the models trained on a dataset from one site may not be applicable to the datasets acquired from other sites with different imaging protocols/scanners. To promote methodological development in the community, iSeg-2019 challenge (http://iseg2019.web.unc.edu) provides a set of 6-month infant subjects from multiple sites with different protocols/scanners for the participating methods. Training/validation subjects are from UNC (MAP) and testing subjects are from UNC/UMN (BCP), Stanford University, and Emory University. By the time of writing, there are 30 automatic segmentation methods participating in iSeg-2019. We review the 8 top-ranked teams by detailing their pipelines/implementations, presenting experimental results and evaluating performance in terms of the whole brain, regions of interest, and gyral landmark curves. We also discuss their limitations and possible future directions for the multi-site issue. We hope that the multi-site dataset in iSeg-2019 and this review article will attract more researchers on the multi-site issue.
Spherical U-Net on Cortical Surfaces: Methods and Applications
Apr 01, 2019


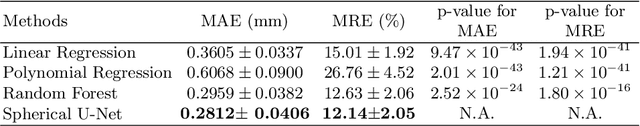
Abstract:Convolutional Neural Networks (CNNs) have been providing the state-of-the-art performance for learning-related problems involving 2D/3D images in Euclidean space. However, unlike in the Euclidean space, the shapes of many structures in medical imaging have a spherical topology in a manifold space, e.g., brain cortical or subcortical surfaces represented by triangular meshes, with large inter-subject and intrasubject variations in vertex number and local connectivity. Hence, there is no consistent neighborhood definition and thus no straightforward convolution/transposed convolution operations for cortical/subcortical surface data. In this paper, by leveraging the regular and consistent geometric structure of the resampled cortical surface mapped onto the spherical space, we propose a novel convolution filter analogous to the standard convolution on the image grid. Accordingly, we develop corresponding operations for convolution, pooling, and transposed convolution for spherical surface data and thus construct spherical CNNs. Specifically, we propose the Spherical U-Net architecture by replacing all operations in the standard U-Net with their spherical operation counterparts. We then apply the Spherical U-Net to two challenging and neuroscientifically important tasks in infant brains: cortical surface parcellation and cortical attribute map development prediction. Both applications demonstrate the competitive performance in the accuracy, computational efficiency, and effectiveness of our proposed Spherical U-Net, in comparison with the state-of-the-art methods.
Deep Morphological Simplification Network (MS-Net) for Guided Registration of Brain Magnetic Resonance Images
Feb 06, 2019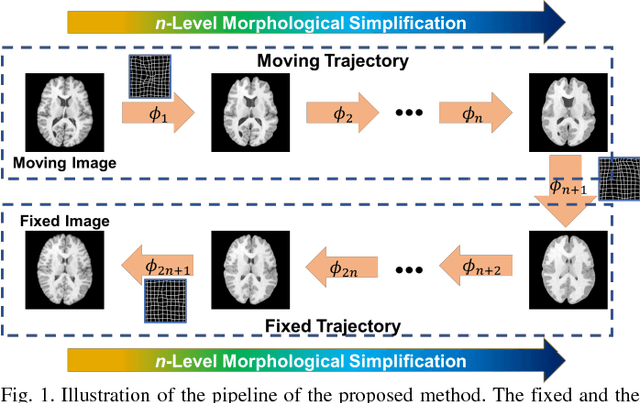
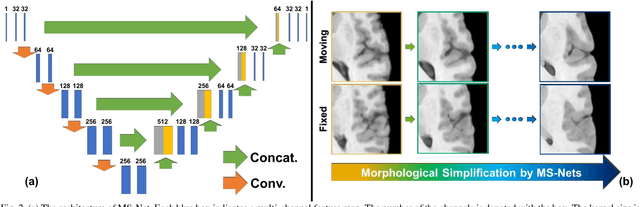
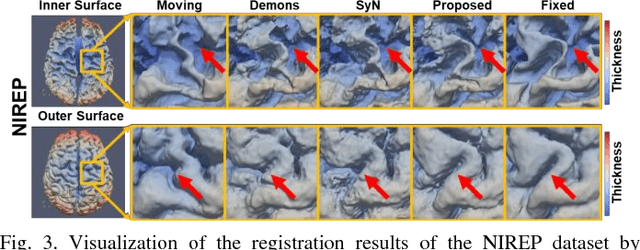
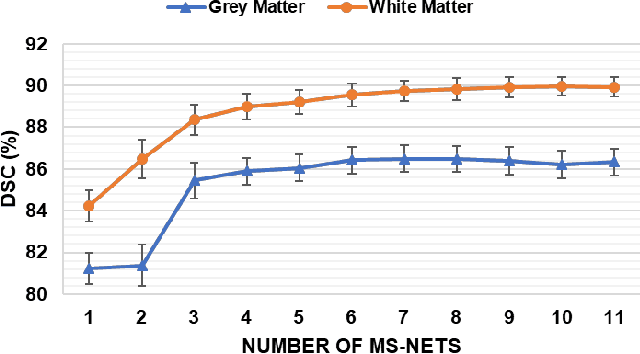
Abstract:Objective: Deformable brain MR image registration is challenging due to large inter-subject anatomical variation. For example, the highly complex cortical folding pattern makes it hard to accurately align corresponding cortical structures of individual images. In this paper, we propose a novel deep learning way to simplify the difficult registration problem of brain MR images. Methods: We train a morphological simplification network (MS-Net), which can generate a "simple" image with less anatomical details based on the "complex" input. With MS-Net, the complexity of the fixed image or the moving image under registration can be reduced gradually, thus building an individual (simplification) trajectory represented by MS-Net outputs. Since the generated images at the ends of the two trajectories (of the fixed and moving images) are so simple and very similar in appearance, they are easy to register. Thus, the two trajectories can act as a bridge to link the fixed and the moving images, and guide their registration. Results: Our experiments show that the proposed method can achieve highly accurate registration performance on different datasets (i.e., NIREP, LPBA, IBSR, CUMC, and MGH). Moreover, the method can be also easily transferred across diverse image datasets and obtain superior accuracy on surface alignment. Conclusion and Significance: We propose MS-Net as a powerful and flexible tool to simplify brain MR images and their registration. To our knowledge, this is the first work to simplify brain MR image registration by deep learning, instead of estimating deformation field directly.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge