Sahar Ahmad
Reconstruction of Cortical Surfaces with Spherical Topology from Infant Brain MRI via Recurrent Deformation Learning
Dec 10, 2023
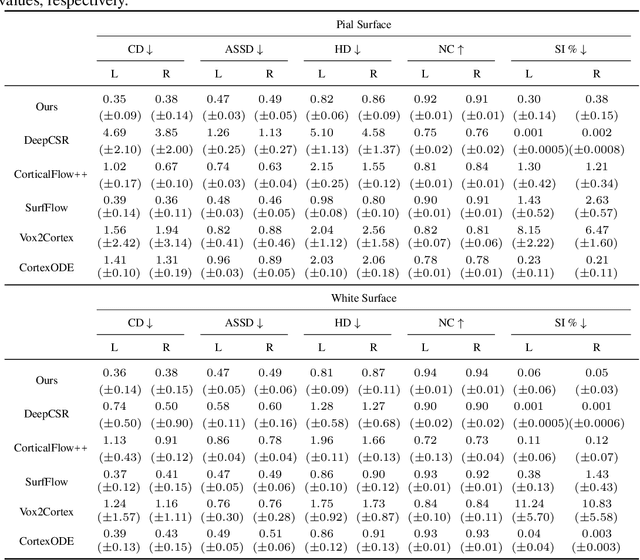
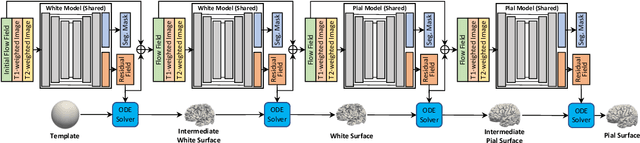
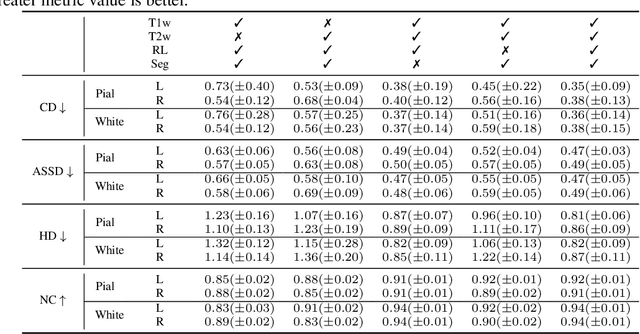
Abstract:Cortical surface reconstruction (CSR) from MRI is key to investigating brain structure and function. While recent deep learning approaches have significantly improved the speed of CSR, a substantial amount of runtime is still needed to map the cortex to a topologically-correct spherical manifold to facilitate downstream geometric analyses. Moreover, this mapping is possible only if the topology of the surface mesh is homotopic to a sphere. Here, we present a method for simultaneous CSR and spherical mapping efficiently within seconds. Our approach seamlessly connects two sub-networks for white and pial surface generation. Residual diffeomorphic deformations are learned iteratively to gradually warp a spherical template mesh to the white and pial surfaces while preserving mesh topology and uniformity. The one-to-one vertex correspondence between the template sphere and the cortical surfaces allows easy and direct mapping of geometric features like convexity and curvature to the sphere for visualization and downstream processing. We demonstrate the efficacy of our approach on infant brain MRI, which poses significant challenges to CSR due to tissue contrast changes associated with rapid brain development during the first postnatal year. Performance evaluation based on a dataset of infants from 0 to 12 months demonstrates that our method substantially enhances mesh regularity and reduces geometric errors, outperforming state-of-the-art deep learning approaches, all while maintaining high computational efficiency.
Brain Tissue Segmentation Across the Human Lifespan via Supervised Contrastive Learning
Jan 03, 2023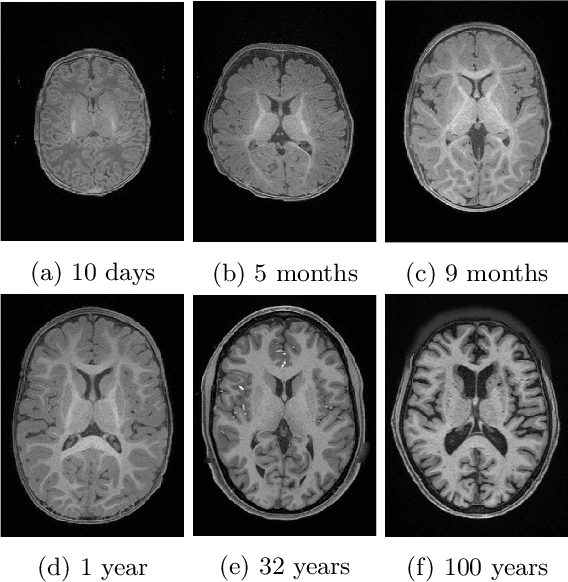
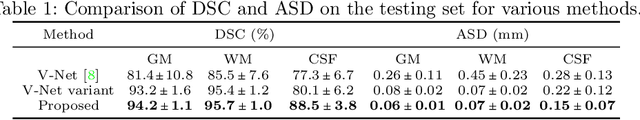
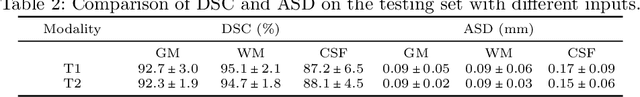
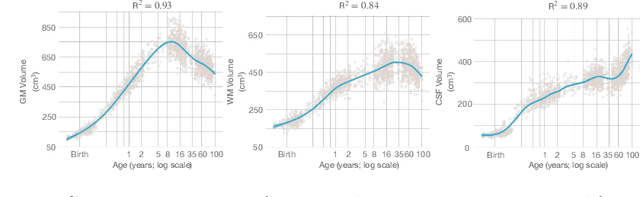
Abstract:Automatic segmentation of brain MR images into white matter (WM), gray matter (GM), and cerebrospinal fluid (CSF) is critical for tissue volumetric analysis and cortical surface reconstruction. Due to dramatic structural and appearance changes associated with developmental and aging processes, existing brain tissue segmentation methods are only viable for specific age groups. Consequently, methods developed for one age group may fail for another. In this paper, we make the first attempt to segment brain tissues across the entire human lifespan (0-100 years of age) using a unified deep learning model. To overcome the challenges related to structural variability underpinned by biological processes, intensity inhomogeneity, motion artifacts, scanner-induced differences, and acquisition protocols, we propose to use contrastive learning to improve the quality of feature representations in a latent space for effective lifespan tissue segmentation. We compared our approach with commonly used segmentation methods on a large-scale dataset of 2,464 MR images. Experimental results show that our model accurately segments brain tissues across the lifespan and outperforms existing methods.
Longitudinal Prediction of Postnatal Brain Magnetic Resonance Images via a Metamorphic Generative Adversarial Network
Aug 09, 2022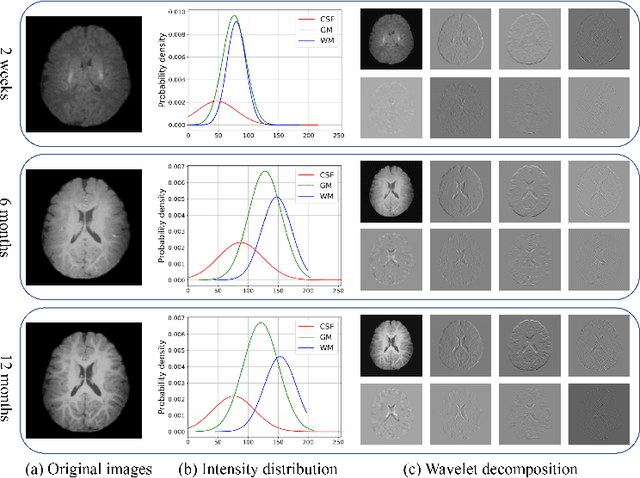

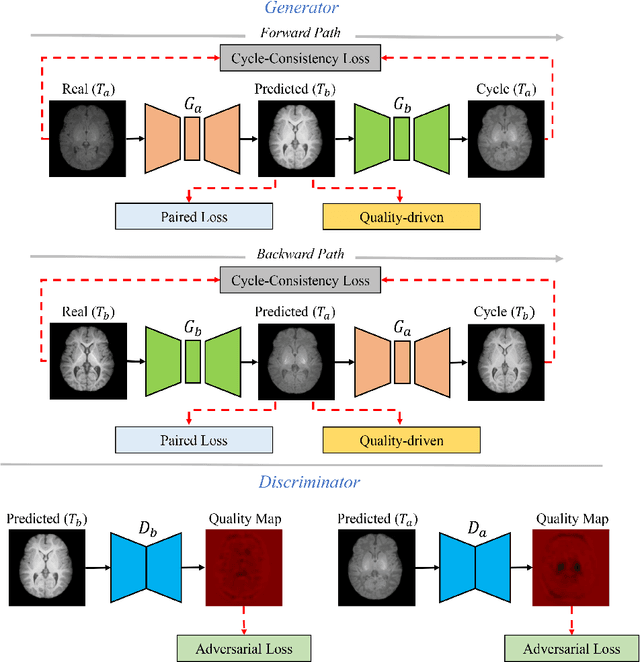
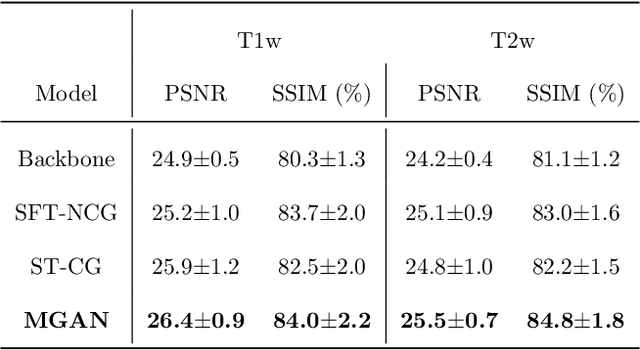
Abstract:Missing scans are inevitable in longitudinal studies due to either subject dropouts or failed scans. In this paper, we propose a deep learning framework to predict missing scans from acquired scans, catering to longitudinal infant studies. Prediction of infant brain MRI is challenging owing to the rapid contrast and structural changes particularly during the first year of life. We introduce a trustworthy metamorphic generative adversarial network (MGAN) for translating infant brain MRI from one time-point to another. MGAN has three key features: (i) Image translation leveraging spatial and frequency information for detail-preserving mapping; (ii) Quality-guided learning strategy that focuses attention on challenging regions. (iii) Multi-scale hybrid loss function that improves translation of tissue contrast and structural details. Experimental results indicate that MGAN outperforms existing GANs by accurately predicting both contrast and anatomical details.
An Auto-Context Deformable Registration Network for Infant Brain MRI
May 19, 2020

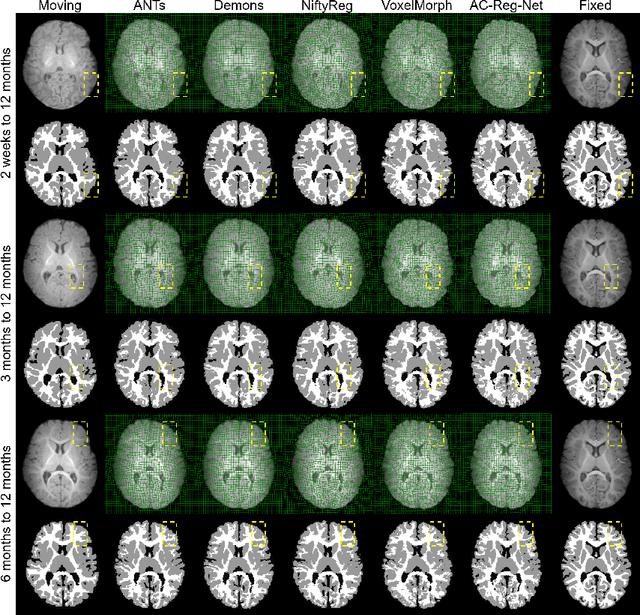
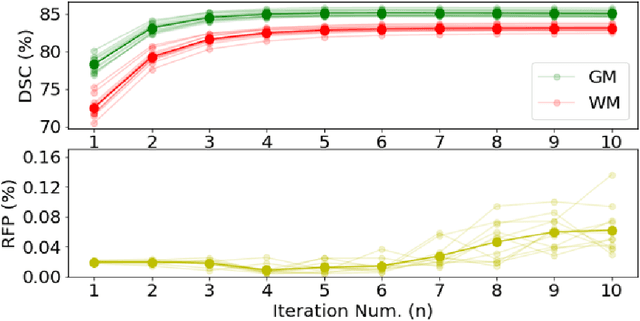
Abstract:Deformable image registration is fundamental to longitudinal and population analysis. Geometric alignment of the infant brain MR images is challenging, owing to rapid changes in image appearance in association with brain development. In this paper, we propose an infant-dedicated deep registration network that uses the auto-context strategy to gradually refine the deformation fields to obtain highly accurate correspondences. Instead of training multiple registration networks, our method estimates the deformation fields by invoking a single network multiple times for iterative deformation refinement. The final deformation field is obtained by the incremental composition of the deformation fields. Experimental results in comparison with state-of-the-art registration methods indicate that our method achieves higher accuracy while at the same time preserves the smoothness of the deformation fields. Our implementation is available online.
Synthesis and Inpainting-Based MR-CT Registration for Image-Guided Thermal Ablation of Liver Tumors
Jul 30, 2019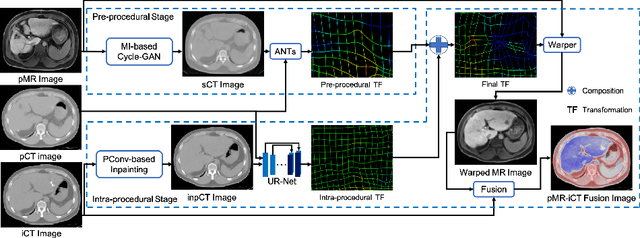
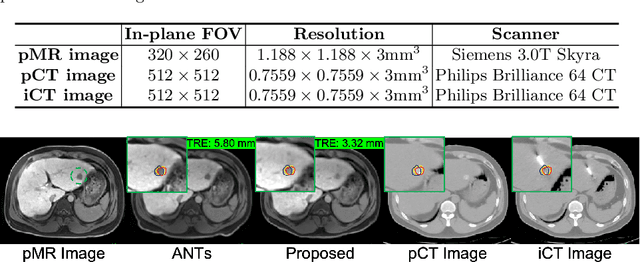
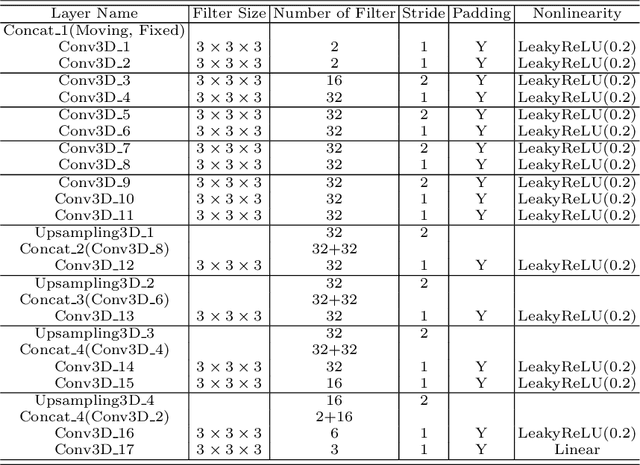
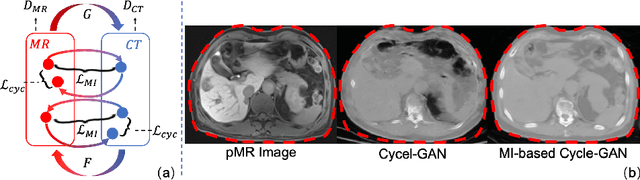
Abstract:Thermal ablation is a minimally invasive procedure for treat-ing small or unresectable tumors. Although CT is widely used for guiding ablation procedures, the contrast of tumors against surrounding normal tissues in CT images is often poor, aggravating the difficulty in accurate thermal ablation. In this paper, we propose a fast MR-CT image registration method to overlay a pre-procedural MR (pMR) image onto an intra-procedural CT (iCT) image for guiding the thermal ablation of liver tumors. By first using a Cycle-GAN model with mutual information constraint to generate synthesized CT (sCT) image from the cor-responding pMR, pre-procedural MR-CT image registration is carried out through traditional mono-modality CT-CT image registration. At the intra-procedural stage, a partial-convolution-based network is first used to inpaint the probe and its artifacts in the iCT image. Then, an unsupervised registration network is used to efficiently align the pre-procedural CT (pCT) with the inpainted iCT (inpCT) image. The final transformation from pMR to iCT is obtained by combining the two estimated transformations,i.e., (1) from the pMR image space to the pCT image space (through sCT) and (2) from the pCT image space to the iCT image space (through inpCT). Experimental results confirm that the proposed method achieves high registration accuracy with a very fast computational speed.
Task Decomposition and Synchronization for Semantic Biomedical Image Segmentation
May 21, 2019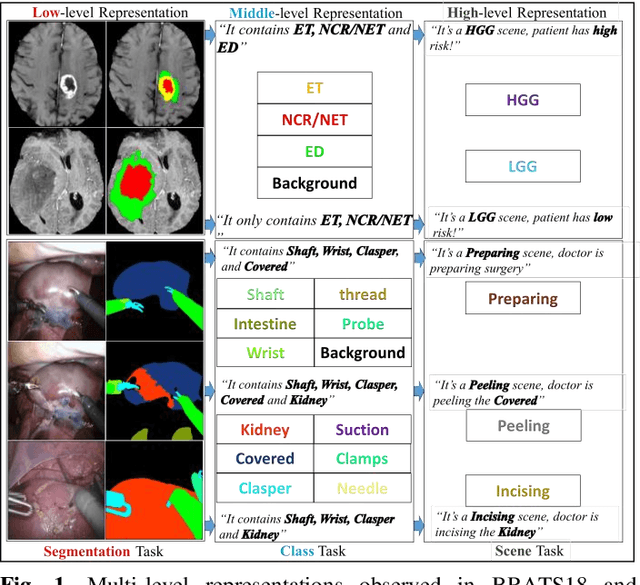
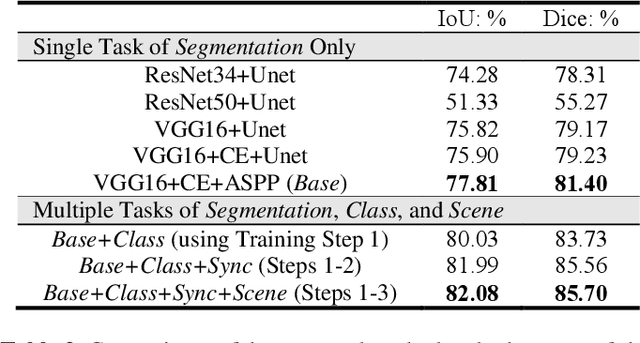

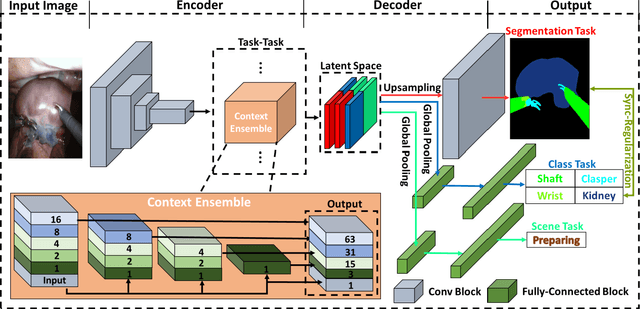
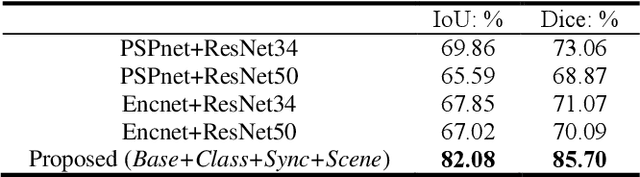
Abstract:Semantic segmentation is essentially important to biomedical image analysis. Many recent works mainly focus on integrating the Fully Convolutional Network (FCN) architecture with sophisticated convolution implementation and deep supervision. In this paper, we propose to decompose the single segmentation task into three subsequent sub-tasks, including (1) pixel-wise image segmentation, (2) prediction of the class labels of the objects within the image, and (3) classification of the scene the image belonging to. While these three sub-tasks are trained to optimize their individual loss functions of different perceptual levels, we propose to let them interact by the task-task context ensemble. Moreover, we propose a novel sync-regularization to penalize the deviation between the outputs of the pixel-wise segmentation and the class prediction tasks. These effective regularizations help FCN utilize context information comprehensively and attain accurate semantic segmentation, even though the number of the images for training may be limited in many biomedical applications. We have successfully applied our framework to three diverse 2D/3D medical image datasets, including Robotic Scene Segmentation Challenge 18 (ROBOT18), Brain Tumor Segmentation Challenge 18 (BRATS18), and Retinal Fundus Glaucoma Challenge (REFUGE18). We have achieved top-tier performance in all three challenges.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge