Xinyang Liu
GDEPO: Group Dual-dynamic and Equal-right-advantage Policy Optimization with Enhanced Training Data Utilization for Sample-Constrained Reinforcement Learning
Jan 11, 2026Abstract:Automated Theorem Proving (ATP) represents a fundamental challenge in Artificial Intelligence (AI), requiring the construction of machine-verifiable proofs in formal languages such as Lean to evaluate AI reasoning capabilities. Reinforcement learning (RL), particularly the high-performance Group Relative Policy Optimization (GRPO) algorithm, has emerged as a mainstream approach for this task. However, in ATP scenarios, GRPO faces two critical issues: when composite rewards are used, its relative advantage estimation may conflict with the binary feedback from the formal verifier; meanwhile, its static sampling strategy may discard entire batches of data if no valid proof is found, resulting in zero contribution to model updates and significant data waste. To address these limitations, we propose Group Dual-dynamic and Equal-right-advantage Policy Optimization (GDEPO), a method incorporating three core mechanisms: 1) dynamic additional sampling, which resamples invalid batches until a valid proof is discovered; 2) equal-right advantage, decoupling the sign of the advantage function (based on correctness) from its magnitude (modulated by auxiliary rewards) to ensure stable and correct policy updates; and 3) dynamic additional iterations, applying extra gradient steps to initially failed but eventually successful samples to accelerate learning on challenging cases. Experiments conducted on three datasets of varying difficulty (MinF2F-test, MathOlympiadBench, PutnamBench) confirm the effectiveness of GDEPO, while ablation studies validate the necessity of its synergistic components. The proposed method enhances data utilization and optimization efficiency, offering a novel training paradigm for ATP.
Route Experts by Sequence, not by Token
Nov 09, 2025Abstract:Mixture-of-Experts (MoE) architectures scale large language models (LLMs) by activating only a subset of experts per token, but the standard TopK routing assigns the same fixed number of experts to all tokens, ignoring their varying complexity. Prior adaptive routing methods introduce additional modules and hyperparameters, often requiring costly retraining from scratch. We propose Sequence-level TopK (SeqTopK), a minimal modification that shifts the expert budget from the token level to the sequence level. By selecting the top $T \cdot K$ experts across all $T$ tokens, SeqTopK enables end-to-end learned dynamic allocation -- assigning more experts to difficult tokens and fewer to easy ones -- while preserving the same overall budget. SeqTopK requires only a few lines of code, adds less than 1% overhead, and remains fully compatible with pretrained MoE models. Experiments across math, coding, law, and writing show consistent improvements over TopK and prior parameter-free adaptive methods, with gains that become substantially larger under higher sparsity (up to 16.9%). These results highlight SeqTopK as a simple, efficient, and scalable routing strategy, particularly well-suited for the extreme sparsity regimes of next-generation LLMs. Code is available at https://github.com/Y-Research-SBU/SeqTopK.
Vision Transformer for Robust Occluded Person Reidentification in Complex Surveillance Scenes
Oct 31, 2025Abstract:Person re-identification (ReID) in surveillance is challenged by occlusion, viewpoint distortion, and poor image quality. Most existing methods rely on complex modules or perform well only on clear frontal images. We propose Sh-ViT (Shuffling Vision Transformer), a lightweight and robust model for occluded person ReID. Built on ViT-Base, Sh-ViT introduces three components: First, a Shuffle module in the final Transformer layer to break spatial correlations and enhance robustness to occlusion and blur; Second, scenario-adapted augmentation (geometric transforms, erasing, blur, and color adjustment) to simulate surveillance conditions; Third, DeiT-based knowledge distillation to improve learning with limited labels.To support real-world evaluation, we construct the MyTT dataset, containing over 10,000 pedestrians and 30,000+ images from base station inspections, with frequent equipment occlusion and camera variations. Experiments show that Sh-ViT achieves 83.2% Rank-1 and 80.1% mAP on MyTT, outperforming CNN and ViT baselines, and 94.6% Rank-1 and 87.5% mAP on Market1501, surpassing state-of-the-art methods.In summary, Sh-ViT improves robustness to occlusion and blur without external modules, offering a practical solution for surveillance-based personnel monitoring.
Conformal Sets in Multiple-Choice Question Answering under Black-Box Settings with Provable Coverage Guarantees
Aug 07, 2025Abstract:Large Language Models (LLMs) have shown remarkable progress in multiple-choice question answering (MCQA), but their inherent unreliability, such as hallucination and overconfidence, limits their application in high-risk domains. To address this, we propose a frequency-based uncertainty quantification method under black-box settings, leveraging conformal prediction (CP) to ensure provable coverage guarantees. Our approach involves multiple independent samplings of the model's output distribution for each input, with the most frequent sample serving as a reference to calculate predictive entropy (PE). Experimental evaluations across six LLMs and four datasets (MedMCQA, MedQA, MMLU, MMLU-Pro) demonstrate that frequency-based PE outperforms logit-based PE in distinguishing between correct and incorrect predictions, as measured by AUROC. Furthermore, the method effectively controls the empirical miscoverage rate under user-specified risk levels, validating that sampling frequency can serve as a viable substitute for logit-based probabilities in black-box scenarios. This work provides a distribution-free model-agnostic framework for reliable uncertainty quantification in MCQA with guaranteed coverage, enhancing the trustworthiness of LLMs in practical applications.
Route-and-Reason: Scaling Large Language Model Reasoning with Reinforced Model Router
Jun 06, 2025Abstract:Multi-step reasoning has proven essential for enhancing the problem-solving capabilities of Large Language Models (LLMs) by decomposing complex tasks into intermediate steps, either explicitly or implicitly. Extending the reasoning chain at test time through deeper thought processes or broader exploration, can furthur improve performance, but often incurs substantial costs due to the explosion in token usage. Yet, many reasoning steps are relatively simple and can be handled by more efficient smaller-scale language models (SLMs). This motivates hybrid approaches that allocate subtasks across models of varying capacities. However, realizing such collaboration requires accurate task decomposition and difficulty-aware subtask allocation, which is challenging. To address this, we propose R2-Reasoner, a novel framework that enables collaborative reasoning across heterogeneous LLMs by dynamically routing sub-tasks based on estimated complexity. At the core of our framework is a Reinforced Model Router, composed of a task decomposer and a subtask allocator. The task decomposer segments complex input queries into logically ordered subtasks, while the subtask allocator assigns each subtask to the most appropriate model, ranging from lightweight SLMs to powerful LLMs, balancing accuracy and efficiency. To train this router, we introduce a staged pipeline that combines supervised fine-tuning on task-specific datasets with Group Relative Policy Optimization algorithm, enabling self-supervised refinement through iterative reinforcement learning. Extensive experiments across four challenging benchmarks demonstrate that R2-Reasoner reduces API costs by 86.85% while maintaining or surpassing baseline accuracy. Our framework paves the way for more cost-effective and adaptive LLM reasoning. The code is open-source at https://anonymous.4open.science/r/R2_Reasoner .
Analysis of the MICCAI Brain Tumor Segmentation -- Metastases (BraTS-METS) 2025 Lighthouse Challenge: Brain Metastasis Segmentation on Pre- and Post-treatment MRI
Apr 16, 2025Abstract:Despite continuous advancements in cancer treatment, brain metastatic disease remains a significant complication of primary cancer and is associated with an unfavorable prognosis. One approach for improving diagnosis, management, and outcomes is to implement algorithms based on artificial intelligence for the automated segmentation of both pre- and post-treatment MRI brain images. Such algorithms rely on volumetric criteria for lesion identification and treatment response assessment, which are still not available in clinical practice. Therefore, it is critical to establish tools for rapid volumetric segmentations methods that can be translated to clinical practice and that are trained on high quality annotated data. The BraTS-METS 2025 Lighthouse Challenge aims to address this critical need by establishing inter-rater and intra-rater variability in dataset annotation by generating high quality annotated datasets from four individual instances of segmentation by neuroradiologists while being recorded on video (two instances doing "from scratch" and two instances after AI pre-segmentation). This high-quality annotated dataset will be used for testing phase in 2025 Lighthouse challenge and will be publicly released at the completion of the challenge. The 2025 Lighthouse challenge will also release the 2023 and 2024 segmented datasets that were annotated using an established pipeline of pre-segmentation, student annotation, two neuroradiologists checking, and one neuroradiologist finalizing the process. It builds upon its previous edition by including post-treatment cases in the dataset. Using these high-quality annotated datasets, the 2025 Lighthouse challenge plans to test benchmark algorithms for automated segmentation of pre-and post-treatment brain metastases (BM), trained on diverse and multi-institutional datasets of MRI images obtained from patients with brain metastases.
Beyond Matryoshka: Revisiting Sparse Coding for Adaptive Representation
Mar 05, 2025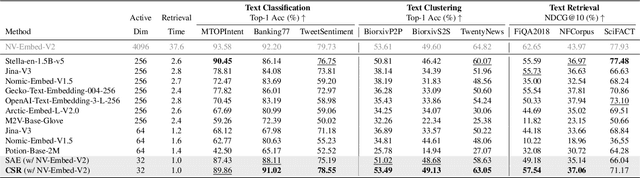
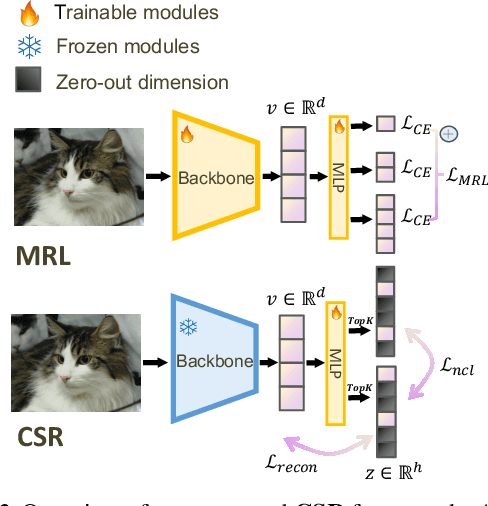
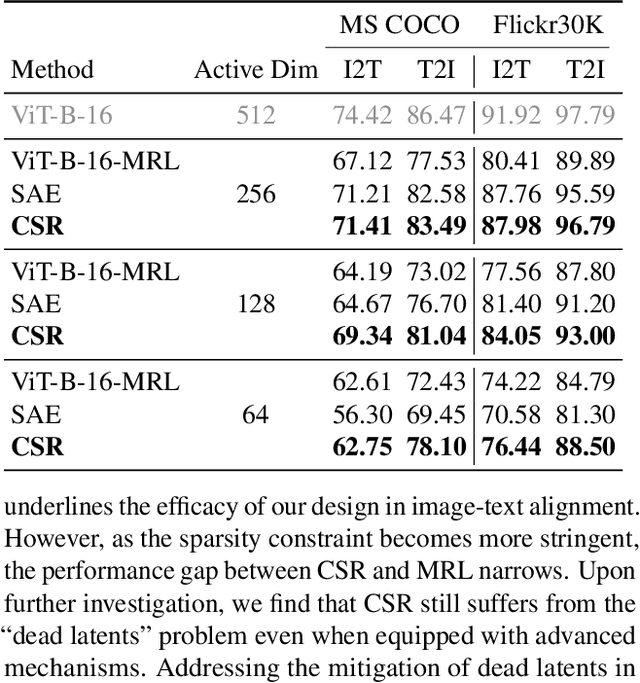
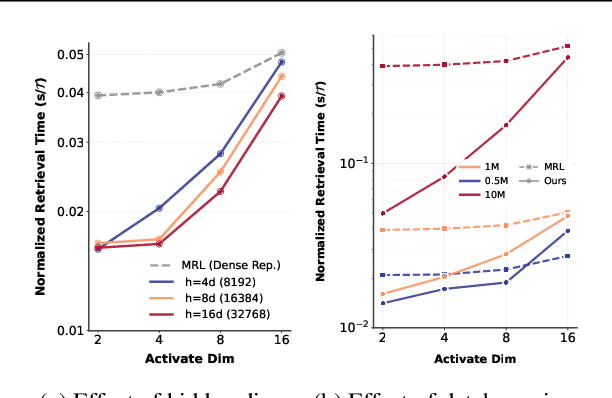
Abstract:Many large-scale systems rely on high-quality deep representations (embeddings) to facilitate tasks like retrieval, search, and generative modeling. Matryoshka Representation Learning (MRL) recently emerged as a solution for adaptive embedding lengths, but it requires full model retraining and suffers from noticeable performance degradations at short lengths. In this paper, we show that sparse coding offers a compelling alternative for achieving adaptive representation with minimal overhead and higher fidelity. We propose Contrastive Sparse Representation (CSR), a method that sparsifies pre-trained embeddings into a high-dimensional but selectively activated feature space. By leveraging lightweight autoencoding and task-aware contrastive objectives, CSR preserves semantic quality while allowing flexible, cost-effective inference at different sparsity levels. Extensive experiments on image, text, and multimodal benchmarks demonstrate that CSR consistently outperforms MRL in terms of both accuracy and retrieval speed-often by large margins-while also cutting training time to a fraction of that required by MRL. Our results establish sparse coding as a powerful paradigm for adaptive representation learning in real-world applications where efficiency and fidelity are both paramount. Code is available at https://github.com/neilwen987/CSR_Adaptive_Rep
Optimal Stochastic Trace Estimation in Generative Modeling
Feb 26, 2025



Abstract:Hutchinson estimators are widely employed in training divergence-based likelihoods for diffusion models to ensure optimal transport (OT) properties. However, this estimator often suffers from high variance and scalability concerns. To address these challenges, we investigate Hutch++, an optimal stochastic trace estimator for generative models, designed to minimize training variance while maintaining transport optimality. Hutch++ is particularly effective for handling ill-conditioned matrices with large condition numbers, which commonly arise when high-dimensional data exhibits a low-dimensional structure. To mitigate the need for frequent and costly QR decompositions, we propose practical schemes that balance frequency and accuracy, backed by theoretical guarantees. Our analysis demonstrates that Hutch++ leads to generations of higher quality. Furthermore, this method exhibits effective variance reduction in various applications, including simulations, conditional time series forecasts, and image generation.
SemiDFL: A Semi-Supervised Paradigm for Decentralized Federated Learning
Dec 18, 2024



Abstract:Decentralized federated learning (DFL) realizes cooperative model training among connected clients without relying on a central server, thereby mitigating communication bottlenecks and eliminating the single-point failure issue present in centralized federated learning (CFL). Most existing work on DFL focuses on supervised learning, assuming each client possesses sufficient labeled data for local training. However, in real-world applications, much of the data is unlabeled. We address this by considering a challenging yet practical semisupervised learning (SSL) scenario in DFL, where clients may have varying data sources: some with few labeled samples, some with purely unlabeled data, and others with both. In this work, we propose SemiDFL, the first semi-supervised DFL method that enhances DFL performance in SSL scenarios by establishing a consensus in both data and model spaces. Specifically, we utilize neighborhood information to improve the quality of pseudo-labeling, which is crucial for effectively leveraging unlabeled data. We then design a consensusbased diffusion model to generate synthesized data, which is used in combination with pseudo-labeled data to create mixed datasets. Additionally, we develop an adaptive aggregation method that leverages the model accuracy of synthesized data to further enhance SemiDFL performance. Through extensive experimentation, we demonstrate the remarkable performance superiority of the proposed DFL-Semi method over existing CFL and DFL schemes in both IID and non-IID SSL scenarios.
Adult Glioma Segmentation in Sub-Saharan Africa using Transfer Learning on Stratified Finetuning Data
Dec 05, 2024



Abstract:Gliomas, a kind of brain tumor characterized by high mortality, present substantial diagnostic challenges in low- and middle-income countries, particularly in Sub-Saharan Africa. This paper introduces a novel approach to glioma segmentation using transfer learning to address challenges in resource-limited regions with minimal and low-quality MRI data. We leverage pre-trained deep learning models, nnU-Net and MedNeXt, and apply a stratified fine-tuning strategy using the BraTS2023-Adult-Glioma and BraTS-Africa datasets. Our method exploits radiomic analysis to create stratified training folds, model training on a large brain tumor dataset, and transfer learning to the Sub-Saharan context. A weighted model ensembling strategy and adaptive post-processing are employed to enhance segmentation accuracy. The evaluation of our proposed method on unseen validation cases on the BraTS-Africa 2024 task resulted in lesion-wise mean Dice scores of 0.870, 0.865, and 0.926, for enhancing tumor, tumor core, and whole tumor regions and was ranked first for the challenge. Our approach highlights the ability of integrated machine-learning techniques to bridge the gap between the medical imaging capabilities of resource-limited countries and established developed regions. By tailoring our methods to a target population's specific needs and constraints, we aim to enhance diagnostic capabilities in isolated environments. Our findings underscore the importance of approaches like local data integration and stratification refinement to address healthcare disparities, ensure practical applicability, and enhance impact.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge