Kelly Payette
Center for MR Research, University Children's Hospital Zurich, University of Zurich, Zurich, Switzerland, Neuroscience Center Zurich, University of Zurich, Zurich, Switzerland
Advances in Automated Fetal Brain MRI Segmentation and Biometry: Insights from the FeTA 2024 Challenge
May 05, 2025



Abstract:Accurate fetal brain tissue segmentation and biometric analysis are essential for studying brain development in utero. The FeTA Challenge 2024 advanced automated fetal brain MRI analysis by introducing biometry prediction as a new task alongside tissue segmentation. For the first time, our diverse multi-centric test set included data from a new low-field (0.55T) MRI dataset. Evaluation metrics were also expanded to include the topology-specific Euler characteristic difference (ED). Sixteen teams submitted segmentation methods, most of which performed consistently across both high- and low-field scans. However, longitudinal trends indicate that segmentation accuracy may be reaching a plateau, with results now approaching inter-rater variability. The ED metric uncovered topological differences that were missed by conventional metrics, while the low-field dataset achieved the highest segmentation scores, highlighting the potential of affordable imaging systems when paired with high-quality reconstruction. Seven teams participated in the biometry task, but most methods failed to outperform a simple baseline that predicted measurements based solely on gestational age, underscoring the challenge of extracting reliable biometric estimates from image data alone. Domain shift analysis identified image quality as the most significant factor affecting model generalization, with super-resolution pipelines also playing a substantial role. Other factors, such as gestational age, pathology, and acquisition site, had smaller, though still measurable, effects. Overall, FeTA 2024 offers a comprehensive benchmark for multi-class segmentation and biometry estimation in fetal brain MRI, underscoring the need for data-centric approaches, improved topological evaluation, and greater dataset diversity to enable clinically robust and generalizable AI tools.
Towards contrast- and pathology-agnostic clinical fetal brain MRI segmentation using SynthSeg
Apr 14, 2025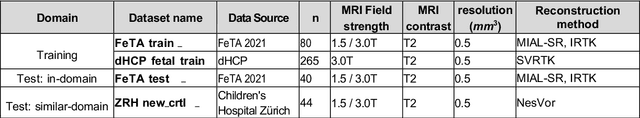



Abstract:Magnetic resonance imaging (MRI) has played a crucial role in fetal neurodevelopmental research. Structural annotations of MR images are an important step for quantitative analysis of the developing human brain, with Deep learning providing an automated alternative for this otherwise tedious manual process. However, segmentation performances of Convolutional Neural Networks often suffer from domain shift, where the network fails when applied to subjects that deviate from the distribution with which it is trained on. In this work, we aim to train networks capable of automatically segmenting fetal brain MRIs with a wide range of domain shifts pertaining to differences in subject physiology and acquisition environments, in particular shape-based differences commonly observed in pathological cases. We introduce a novel data-driven train-time sampling strategy that seeks to fully exploit the diversity of a given training dataset to enhance the domain generalizability of the trained networks. We adapted our sampler, together with other existing data augmentation techniques, to the SynthSeg framework, a generator that utilizes domain randomization to generate diverse training data, and ran thorough experimentations and ablation studies on a wide range of training/testing data to test the validity of the approaches. Our networks achieved notable improvements in the segmentation quality on testing subjects with intense anatomical abnormalities (p < 1e-4), though at the cost of a slighter decrease in performance in cases with fewer abnormalities. Our work also lays the foundation for future works on creating and adapting data-driven sampling strategies for other training pipelines.
Pathological MRI Segmentation by Synthetic Pathological Data Generation in Fetuses and Neonates
Jan 31, 2025



Abstract:Developing new methods for the automated analysis of clinical fetal and neonatal MRI data is limited by the scarcity of annotated pathological datasets and privacy concerns that often restrict data sharing, hindering the effectiveness of deep learning models. We address this in two ways. First, we introduce Fetal&Neonatal-DDPM, a novel diffusion model framework designed to generate high-quality synthetic pathological fetal and neonatal MRIs from semantic label images. Second, we enhance training data by modifying healthy label images through morphological alterations to simulate conditions such as ventriculomegaly, cerebellar and pontocerebellar hypoplasia, and microcephaly. By leveraging Fetal&Neonatal-DDPM, we synthesize realistic pathological MRIs from these modified pathological label images. Radiologists rated the synthetic MRIs as significantly (p < 0.05) superior in quality and diagnostic value compared to real MRIs, demonstrating features such as blood vessels and choroid plexus, and improved alignment with label annotations. Synthetic pathological data enhanced state-of-the-art nnUNet segmentation performance, particularly for severe ventriculomegaly cases, with the greatest improvements achieved in ventricle segmentation (Dice scores: 0.9253 vs. 0.7317). This study underscores the potential of generative AI as transformative tool for data augmentation, offering improved segmentation performance in pathological cases. This development represents a significant step towards improving analysis and segmentation accuracy in prenatal imaging, and also offers new ways for data anonymization through the generation of pathologic image data.
Multi-Center Fetal Brain Tissue Annotation (FeTA) Challenge 2022 Results
Feb 08, 2024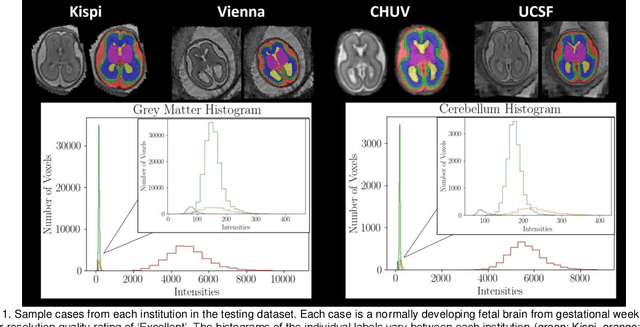
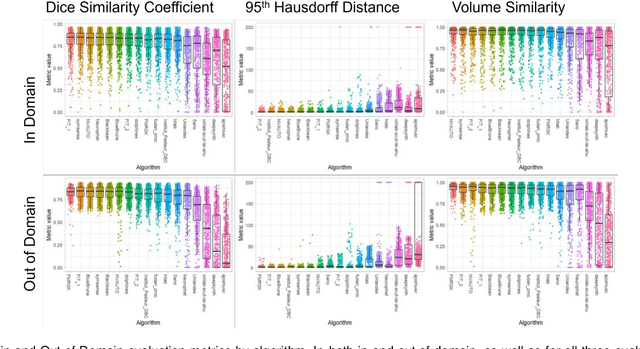
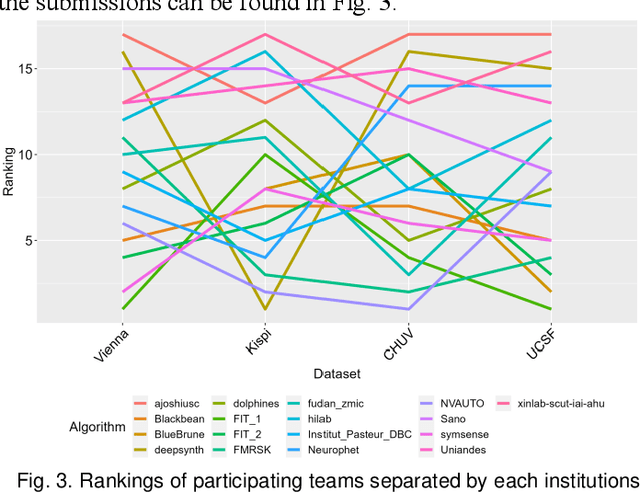
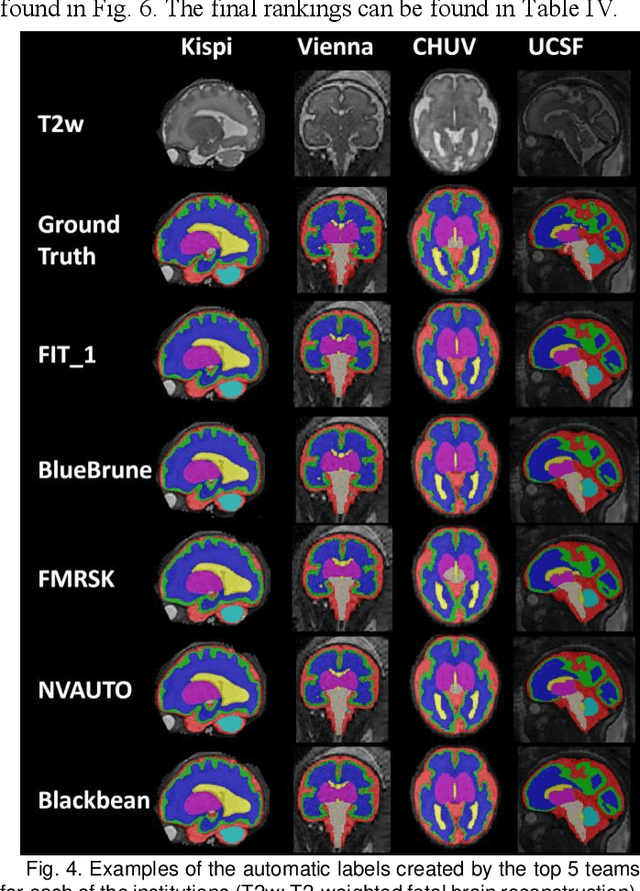
Abstract:Segmentation is a critical step in analyzing the developing human fetal brain. There have been vast improvements in automatic segmentation methods in the past several years, and the Fetal Brain Tissue Annotation (FeTA) Challenge 2021 helped to establish an excellent standard of fetal brain segmentation. However, FeTA 2021 was a single center study, and the generalizability of algorithms across different imaging centers remains unsolved, limiting real-world clinical applicability. The multi-center FeTA Challenge 2022 focuses on advancing the generalizability of fetal brain segmentation algorithms for magnetic resonance imaging (MRI). In FeTA 2022, the training dataset contained images and corresponding manually annotated multi-class labels from two imaging centers, and the testing data contained images from these two imaging centers as well as two additional unseen centers. The data from different centers varied in many aspects, including scanners used, imaging parameters, and fetal brain super-resolution algorithms applied. 16 teams participated in the challenge, and 17 algorithms were evaluated. Here, a detailed overview and analysis of the challenge results are provided, focusing on the generalizability of the submissions. Both in- and out of domain, the white matter and ventricles were segmented with the highest accuracy, while the most challenging structure remains the cerebral cortex due to anatomical complexity. The FeTA Challenge 2022 was able to successfully evaluate and advance generalizability of multi-class fetal brain tissue segmentation algorithms for MRI and it continues to benchmark new algorithms. The resulting new methods contribute to improving the analysis of brain development in utero.
An automated pipeline for quantitative T2* fetal body MRI and segmentation at low field
Aug 09, 2023Abstract:Fetal Magnetic Resonance Imaging at low field strengths is emerging as an exciting direction in perinatal health. Clinical low field (0.55T) scanners are beneficial for fetal imaging due to their reduced susceptibility-induced artefacts, increased T2* values, and wider bore (widening access for the increasingly obese pregnant population). However, the lack of standard automated image processing tools such as segmentation and reconstruction hampers wider clinical use. In this study, we introduce a semi-automatic pipeline using quantitative MRI for the fetal body at low field strength resulting in fast and detailed quantitative T2* relaxometry analysis of all major fetal body organs. Multi-echo dynamic sequences of the fetal body were acquired and reconstructed into a single high-resolution volume using deformable slice-to-volume reconstruction, generating both structural and quantitative T2* 3D volumes. A neural network trained using a semi-supervised approach was created to automatically segment these fetal body 3D volumes into ten different organs (resulting in dice values > 0.74 for 8 out of 10 organs). The T2* values revealed a strong relationship with GA in the lungs, liver, and kidney parenchyma (R^2>0.5). This pipeline was used successfully for a wide range of GAs (17-40 weeks), and is robust to motion artefacts. Low field fetal MRI can be used to perform advanced MRI analysis, and is a viable option for clinical scanning.
Why is the winner the best?
Mar 30, 2023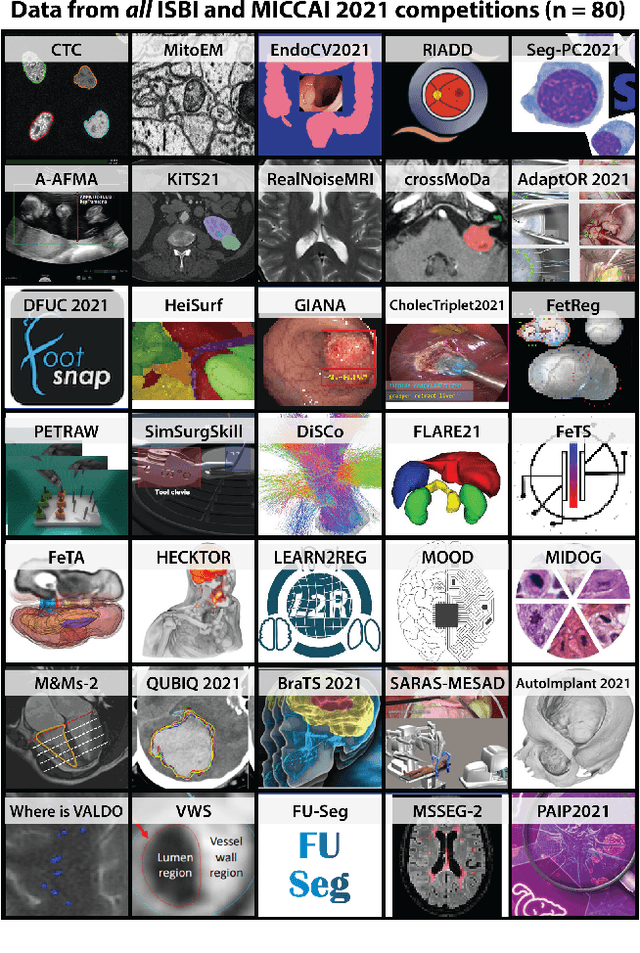
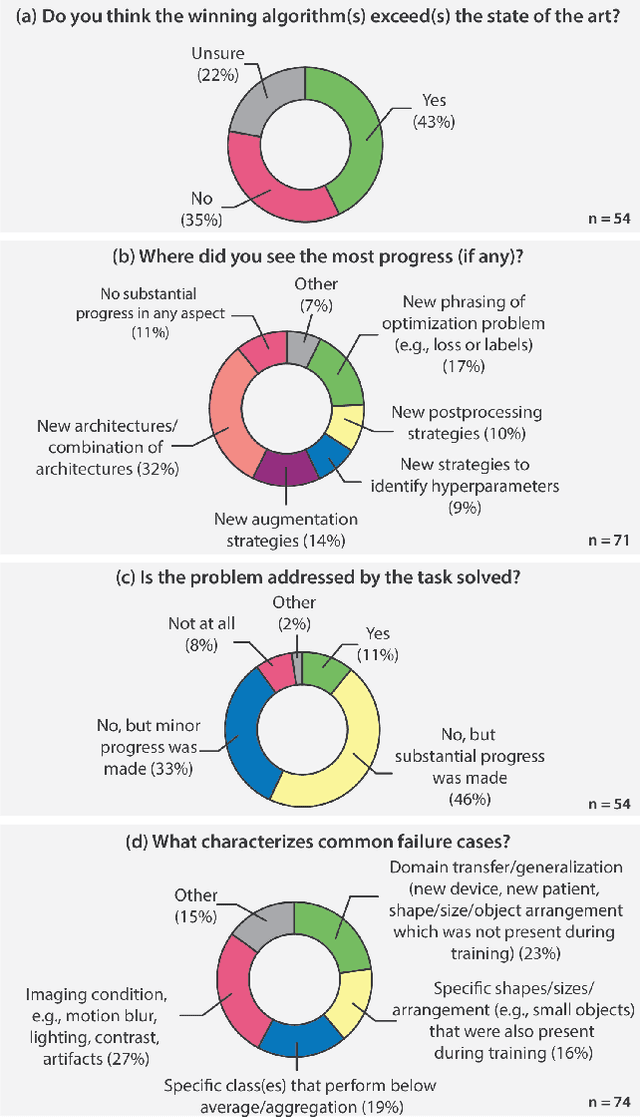
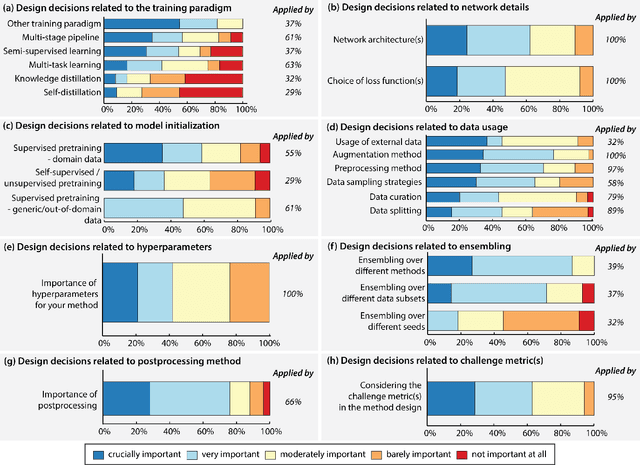
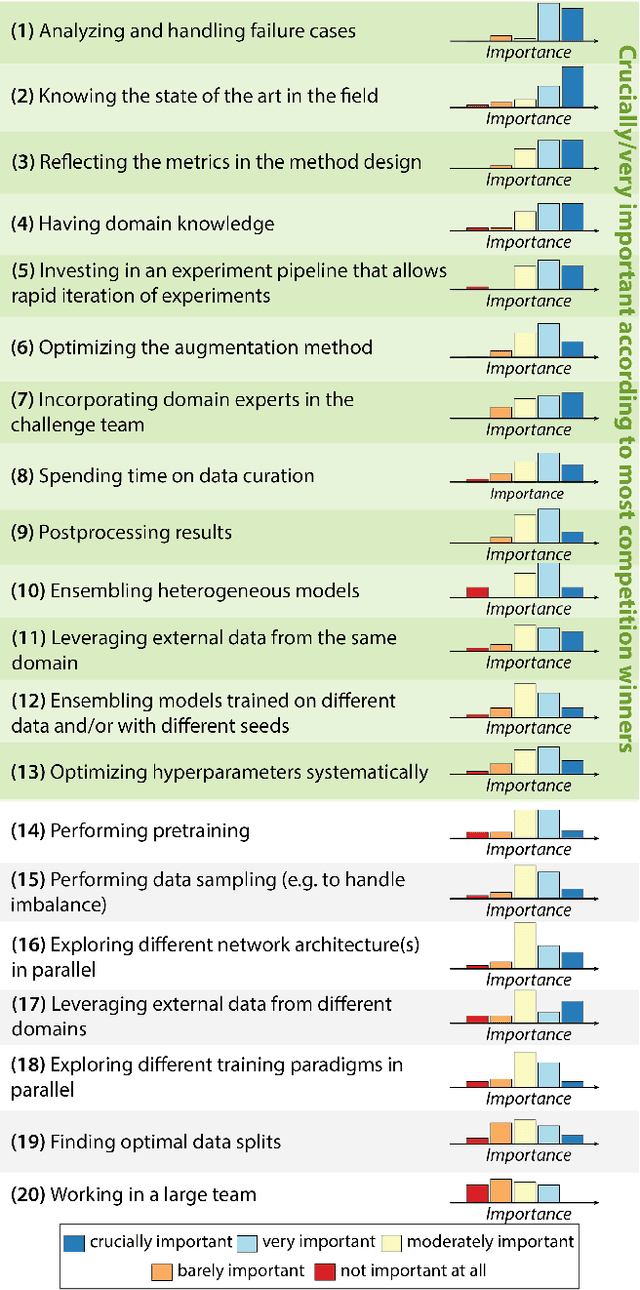
Abstract:International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Synthesis of realistic fetal MRI with conditional Generative Adversarial Networks
Sep 20, 2022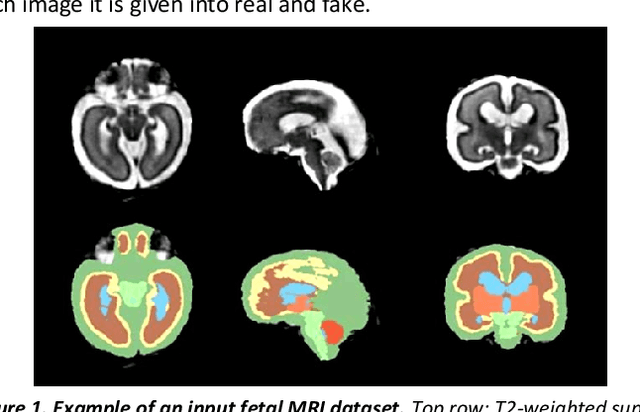
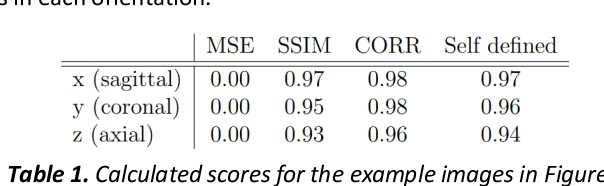
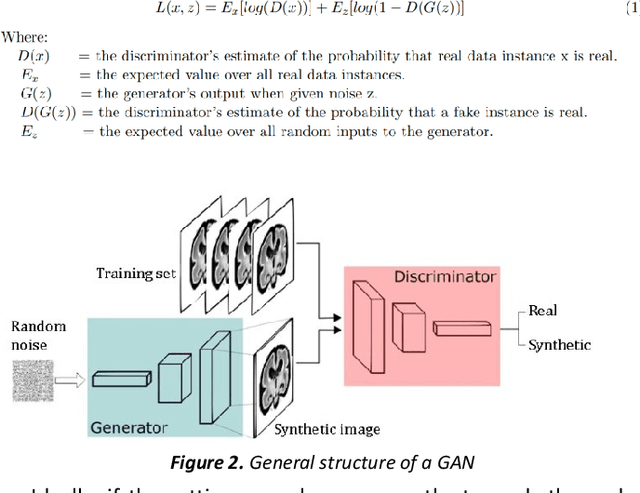
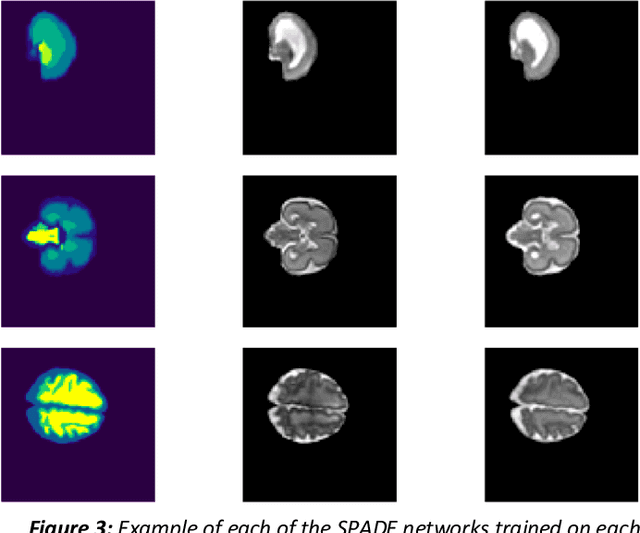
Abstract:Fetal brain magnetic resonance imaging serves as an emerging modality for prenatal counseling and diagnosis in disorders affecting the brain. Machine learning based segmentation plays an important role in the quantification of brain development. However, a limiting factor is the lack of sufficiently large, labeled training data. Our study explored the application of SPADE, a conditional general adversarial network (cGAN), which learns the mapping from the label to the image space. The input to the network was super-resolution T2-weighted cerebral MRI data of 120 fetuses (gestational age range: 20-35 weeks, normal and pathological), which were annotated for 7 different tissue categories. SPADE networks were trained on 256*256 2D slices of the reconstructed volumes (image and label pairs) in each orthogonal orientation. To combine the generated volumes from each orientation into one image, a simple mean of the outputs of the three networks was taken. Based on the label maps only, we synthesized highly realistic images. However, some finer details, like small vessels were not synthesized. A structural similarity index (SSIM) of 0.972+-0.016 and correlation coefficient of 0.974+-0.008 were achieved. To demonstrate the capacity of the cGAN to create new anatomical variants, we artificially dilated the ventricles in the segmentation map and created synthetic MRI of different degrees of fetal hydrocephalus. cGANs, such as the SPADE algorithm, allow the generation of hypothetically unseen scenarios and anatomical configurations in the label space, which data in turn can be utilized for training various machine learning algorithms. In the future, this algorithm would be used for generating large, synthetic datasets representing fetal brain development. These datasets would potentially improve the performance of currently available segmentation networks.
Fetal Brain Tissue Annotation and Segmentation Challenge Results
Apr 20, 2022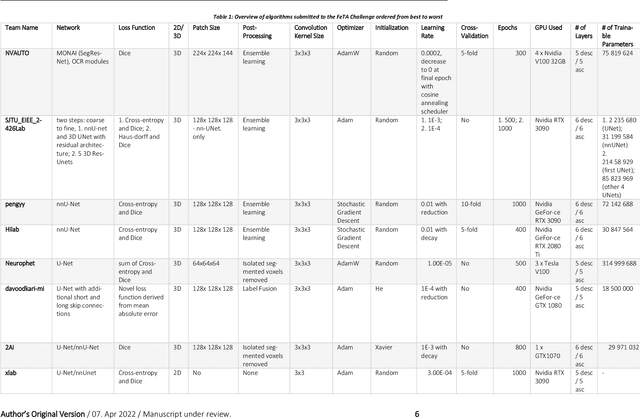
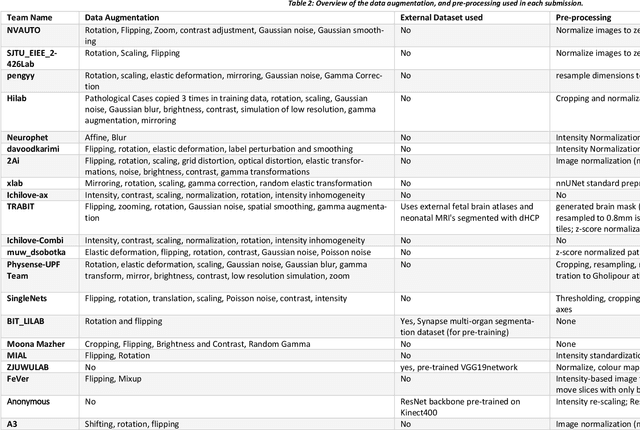
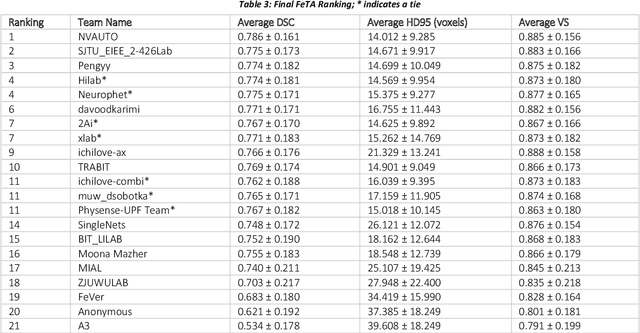
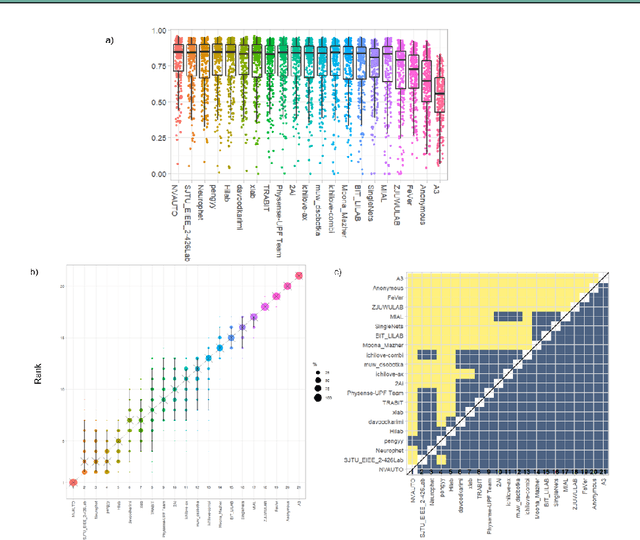
Abstract:In-utero fetal MRI is emerging as an important tool in the diagnosis and analysis of the developing human brain. Automatic segmentation of the developing fetal brain is a vital step in the quantitative analysis of prenatal neurodevelopment both in the research and clinical context. However, manual segmentation of cerebral structures is time-consuming and prone to error and inter-observer variability. Therefore, we organized the Fetal Tissue Annotation (FeTA) Challenge in 2021 in order to encourage the development of automatic segmentation algorithms on an international level. The challenge utilized FeTA Dataset, an open dataset of fetal brain MRI reconstructions segmented into seven different tissues (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, brainstem, deep grey matter). 20 international teams participated in this challenge, submitting a total of 21 algorithms for evaluation. In this paper, we provide a detailed analysis of the results from both a technical and clinical perspective. All participants relied on deep learning methods, mainly U-Nets, with some variability present in the network architecture, optimization, and image pre- and post-processing. The majority of teams used existing medical imaging deep learning frameworks. The main differences between the submissions were the fine tuning done during training, and the specific pre- and post-processing steps performed. The challenge results showed that almost all submissions performed similarly. Four of the top five teams used ensemble learning methods. However, one team's algorithm performed significantly superior to the other submissions, and consisted of an asymmetrical U-Net network architecture. This paper provides a first of its kind benchmark for future automatic multi-tissue segmentation algorithms for the developing human brain in utero.
Synthetic magnetic resonance images for domain adaptation: Application to fetal brain tissue segmentation
Nov 08, 2021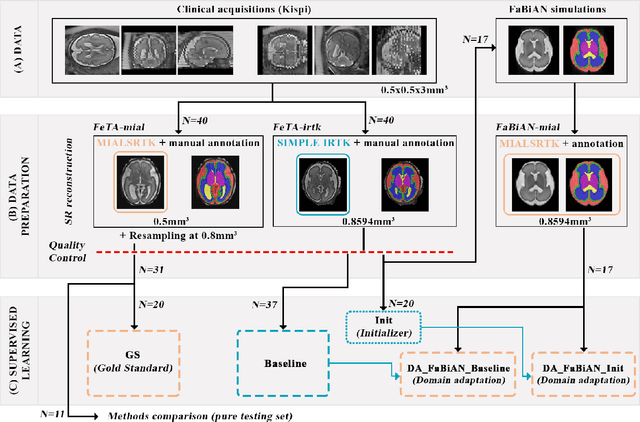
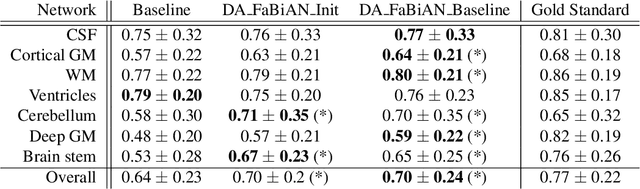

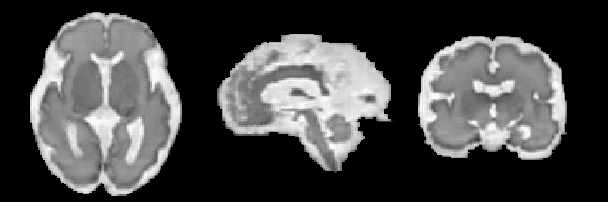
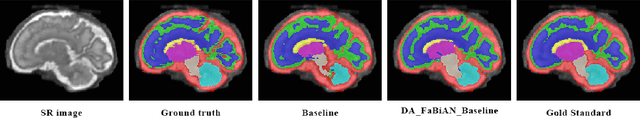
Abstract:The quantitative assessment of the developing human brain in utero is crucial to fully understand neurodevelopment. Thus, automated multi-tissue fetal brain segmentation algorithms are being developed, which in turn require annotated data to be trained. However, the available annotated fetal brain datasets are limited in number and heterogeneity, hampering domain adaptation strategies for robust segmentation. In this context, we use FaBiAN, a Fetal Brain magnetic resonance Acquisition Numerical phantom, to simulate various realistic magnetic resonance images of the fetal brain along with its class labels. We demonstrate that these multiple synthetic annotated data, generated at no cost and further reconstructed using the target super-resolution technique, can be successfully used for domain adaptation of a deep learning method that segments seven brain tissues. Overall, the accuracy of the segmentation is significantly enhanced, especially in the cortical gray matter, the white matter, the cerebellum, the deep gray matter and the brain stem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge