Md Mahfuzur Rahman Siddiquee
AnoFPDM: Anomaly Segmentation with Forward Process of Diffusion Models for Brain MRI
Apr 24, 2024

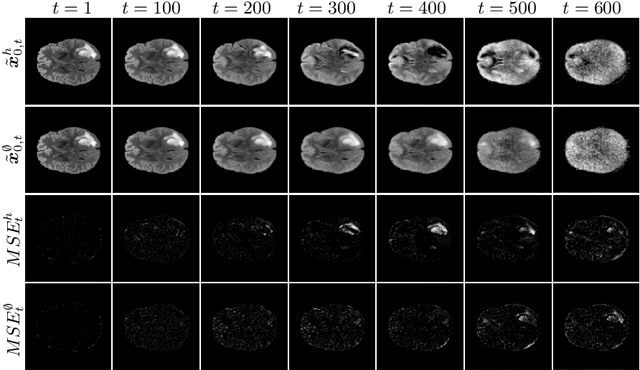

Abstract:Weakly-supervised diffusion models (DM) in anomaly segmentation, leveraging image-level labels, have attracted significant attention for their superior performance compared to unsupervised methods. It eliminates the need for pixel-level labels in training, offering a more cost-effective alternative to supervised methods. However, existing methods are not fully weakly-supervised because they heavily rely on costly pixel-level labels for hyperparameter tuning in inference. To tackle this challenge, we introduce Anomaly Segmentation with Forward Process of Diffusion Models (AnoFPDM), a fully weakly-supervised framework that operates without the need for pixel-level labels. Leveraging the unguided forward process as a reference, we identify suitable hyperparameters, i.e., noise scale and threshold, for each input image. We aggregate anomaly maps from each step in the forward process, enhancing the signal strength of anomalous regions. Remarkably, our proposed method outperforms recent state-of-the-art weakly-supervised approaches, even without utilizing pixel-level labels.
A Robust Ensemble Algorithm for Ischemic Stroke Lesion Segmentation: Generalizability and Clinical Utility Beyond the ISLES Challenge
Apr 03, 2024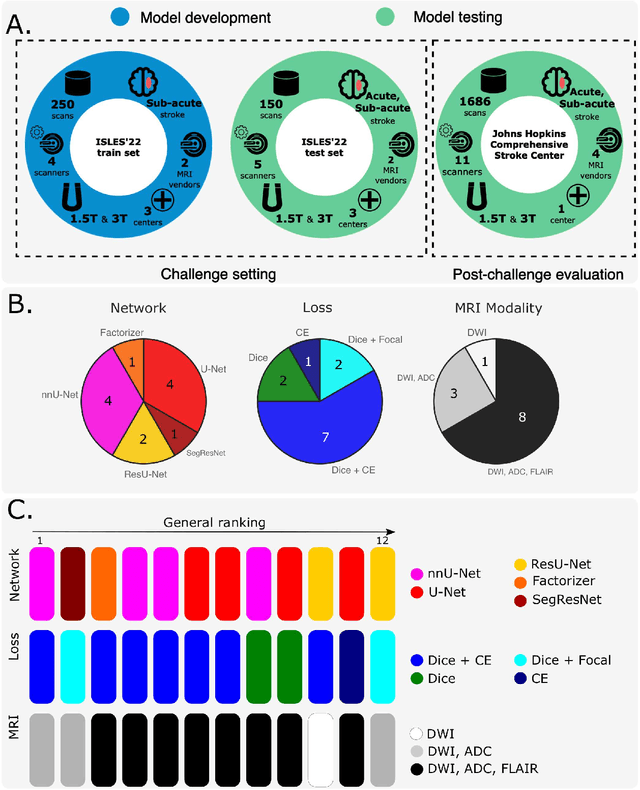
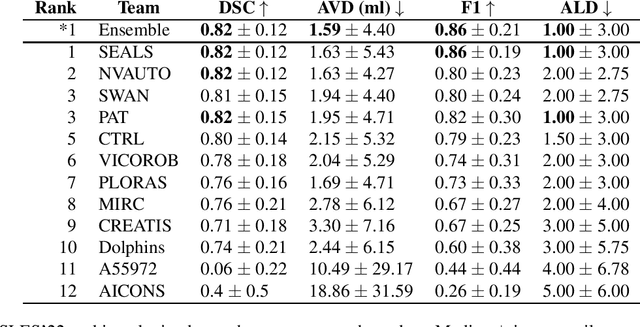
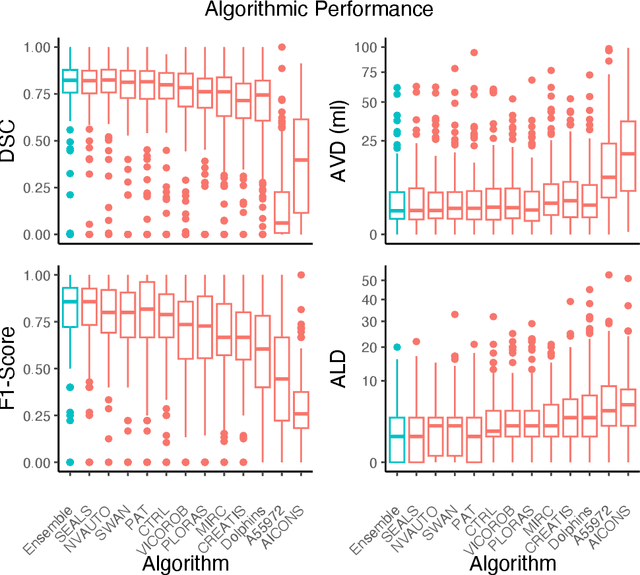
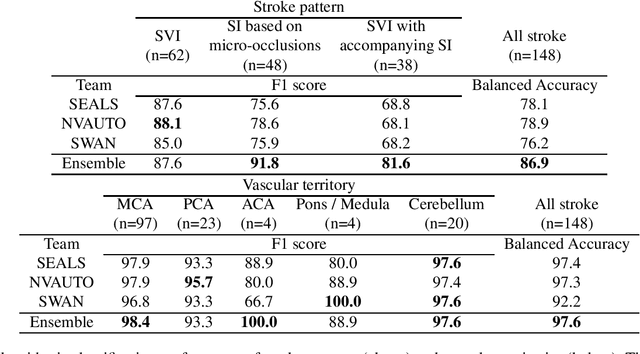
Abstract:Diffusion-weighted MRI (DWI) is essential for stroke diagnosis, treatment decisions, and prognosis. However, image and disease variability hinder the development of generalizable AI algorithms with clinical value. We address this gap by presenting a novel ensemble algorithm derived from the 2022 Ischemic Stroke Lesion Segmentation (ISLES) challenge. ISLES'22 provided 400 patient scans with ischemic stroke from various medical centers, facilitating the development of a wide range of cutting-edge segmentation algorithms by the research community. Through collaboration with leading teams, we combined top-performing algorithms into an ensemble model that overcomes the limitations of individual solutions. Our ensemble model achieved superior ischemic lesion detection and segmentation accuracy on our internal test set compared to individual algorithms. This accuracy generalized well across diverse image and disease variables. Furthermore, the model excelled in extracting clinical biomarkers. Notably, in a Turing-like test, neuroradiologists consistently preferred the algorithm's segmentations over manual expert efforts, highlighting increased comprehensiveness and precision. Validation using a real-world external dataset (N=1686) confirmed the model's generalizability. The algorithm's outputs also demonstrated strong correlations with clinical scores (admission NIHSS and 90-day mRS) on par with or exceeding expert-derived results, underlining its clinical relevance. This study offers two key findings. First, we present an ensemble algorithm (https://github.com/Tabrisrei/ISLES22_Ensemble) that detects and segments ischemic stroke lesions on DWI across diverse scenarios on par with expert (neuro)radiologists. Second, we show the potential for biomedical challenge outputs to extend beyond the challenge's initial objectives, demonstrating their real-world clinical applicability.
Multi-Center Fetal Brain Tissue Annotation (FeTA) Challenge 2022 Results
Feb 08, 2024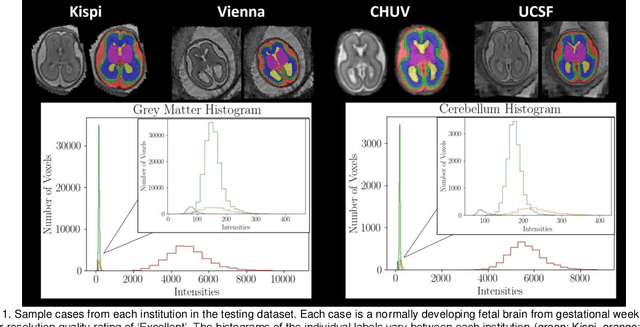
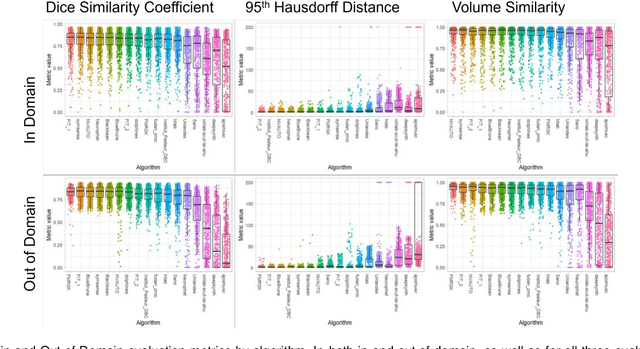
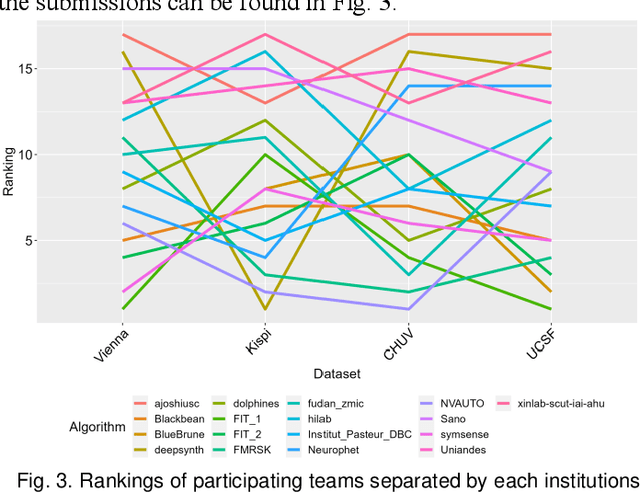
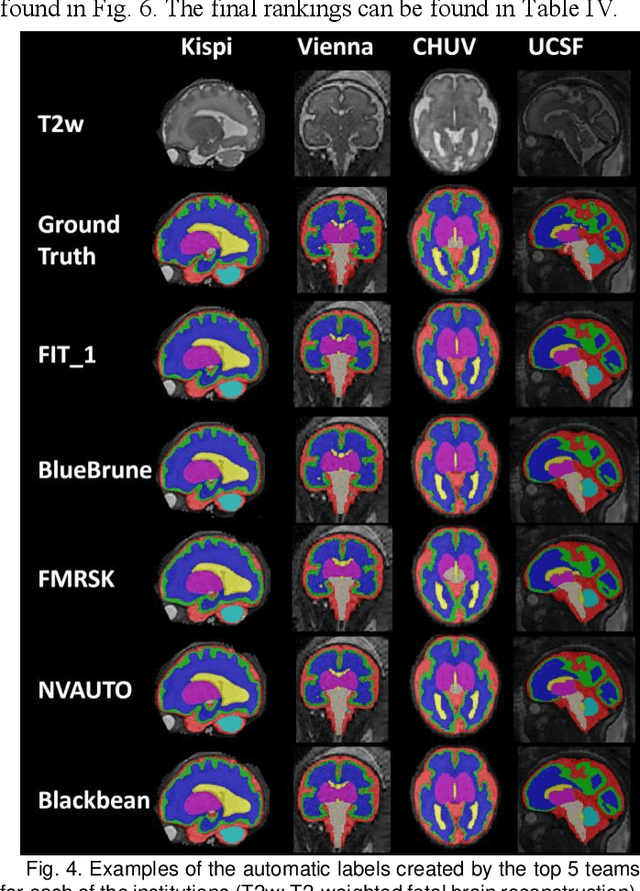
Abstract:Segmentation is a critical step in analyzing the developing human fetal brain. There have been vast improvements in automatic segmentation methods in the past several years, and the Fetal Brain Tissue Annotation (FeTA) Challenge 2021 helped to establish an excellent standard of fetal brain segmentation. However, FeTA 2021 was a single center study, and the generalizability of algorithms across different imaging centers remains unsolved, limiting real-world clinical applicability. The multi-center FeTA Challenge 2022 focuses on advancing the generalizability of fetal brain segmentation algorithms for magnetic resonance imaging (MRI). In FeTA 2022, the training dataset contained images and corresponding manually annotated multi-class labels from two imaging centers, and the testing data contained images from these two imaging centers as well as two additional unseen centers. The data from different centers varied in many aspects, including scanners used, imaging parameters, and fetal brain super-resolution algorithms applied. 16 teams participated in the challenge, and 17 algorithms were evaluated. Here, a detailed overview and analysis of the challenge results are provided, focusing on the generalizability of the submissions. Both in- and out of domain, the white matter and ventricles were segmented with the highest accuracy, while the most challenging structure remains the cerebral cortex due to anatomical complexity. The FeTA Challenge 2022 was able to successfully evaluate and advance generalizability of multi-class fetal brain tissue segmentation algorithms for MRI and it continues to benchmark new algorithms. The resulting new methods contribute to improving the analysis of brain development in utero.
Pseudo Supervised Metrics: Evaluating Unsupervised Image to Image Translation Models In Unsupervised Cross-Domain Classification Frameworks
Mar 18, 2023Abstract:The ability to classify images accurately and efficiently is dependent on having access to large labeled datasets and testing on data from the same domain that the model is trained on. Classification becomes more challenging when dealing with new data from a different domain, where collecting a large labeled dataset and training a new classifier from scratch is time-consuming, expensive, and sometimes infeasible or impossible. Cross-domain classification frameworks were developed to handle this data domain shift problem by utilizing unsupervised image-to-image (UI2I) translation models to translate an input image from the unlabeled domain to the labeled domain. The problem with these unsupervised models lies in their unsupervised nature. For lack of annotations, it is not possible to use the traditional supervised metrics to evaluate these translation models to pick the best-saved checkpoint model. In this paper, we introduce a new method called Pseudo Supervised Metrics that was designed specifically to support cross-domain classification applications contrary to other typically used metrics such as the FID which was designed to evaluate the model in terms of the quality of the generated image from a human-eye perspective. We show that our metric not only outperforms unsupervised metrics such as the FID, but is also highly correlated with the true supervised metrics, robust, and explainable. Furthermore, we demonstrate that it can be used as a standard metric for future research in this field by applying it to a critical real-world problem (the boiling crisis problem).
Brainomaly: Unsupervised Neurologic Disease Detection Utilizing Unannotated T1-weighted Brain MR Images
Feb 18, 2023Abstract:Deep neural networks have revolutionized the field of supervised learning by enabling accurate predictions through learning from large annotated datasets. However, acquiring large annotated medical imaging datasets is a challenging task, especially for rare diseases, due to the high cost, time, and effort required for annotation. In these scenarios, unsupervised disease detection methods, such as anomaly detection, can save significant human effort. A typically used approach for anomaly detection is to learn the images from healthy subjects only, assuming the model will detect the images from diseased subjects as outliers. However, in many real-world scenarios, unannotated datasets with a mix of healthy and diseased individuals are available. Recent studies have shown improvement in unsupervised disease/anomaly detection using such datasets of unannotated images from healthy and diseased individuals compared to datasets that only include images from healthy individuals. A major issue remains unaddressed in these studies, which is selecting the best model for inference from a set of trained models without annotated samples. To address this issue, we propose Brainomaly, a GAN-based image-to-image translation method for neurologic disease detection using unannotated T1-weighted brain MRIs of individuals with neurologic diseases and healthy subjects. Brainomaly is trained to remove the diseased regions from the input brain MRIs and generate MRIs of corresponding healthy brains. Instead of generating the healthy images directly, Brainomaly generates an additive map where each voxel indicates the amount of changes required to make the input image look healthy. In addition, Brainomaly uses a pseudo-AUC metric for inference model selection, which further improves the detection performance. Our Brainomaly outperforms existing state-of-the-art methods by large margins.
The state-of-the-art 3D anisotropic intracranial hemorrhage segmentation on non-contrast head CT: The INSTANCE challenge
Jan 12, 2023Abstract:Automatic intracranial hemorrhage segmentation in 3D non-contrast head CT (NCCT) scans is significant in clinical practice. Existing hemorrhage segmentation methods usually ignores the anisotropic nature of the NCCT, and are evaluated on different in-house datasets with distinct metrics, making it highly challenging to improve segmentation performance and perform objective comparisons among different methods. The INSTANCE 2022 was a grand challenge held in conjunction with the 2022 International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). It is intended to resolve the above-mentioned problems and promote the development of both intracranial hemorrhage segmentation and anisotropic data processing. The INSTANCE released a training set of 100 cases with ground-truth and a validation set with 30 cases without ground-truth labels that were available to the participants. A held-out testing set with 70 cases is utilized for the final evaluation and ranking. The methods from different participants are ranked based on four metrics, including Dice Similarity Coefficient (DSC), Hausdorff Distance (HD), Relative Volume Difference (RVD) and Normalized Surface Dice (NSD). A total of 13 teams submitted distinct solutions to resolve the challenges, making several baseline models, pre-processing strategies and anisotropic data processing techniques available to future researchers. The winner method achieved an average DSC of 0.6925, demonstrating a significant growth over our proposed baseline method. To the best of our knowledge, the proposed INSTANCE challenge releases the first intracranial hemorrhage segmentation benchmark, and is also the first challenge that intended to resolve the anisotropic problem in 3D medical image segmentation, which provides new alternatives in these research fields.
A Generalized Framework for Critical Heat Flux Detection Using Unsupervised Image-to-Image Translation
Dec 25, 2022



Abstract:This work proposes a framework developed to generalize Critical Heat Flux (CHF) detection classification models using an Unsupervised Image-to-Image (UI2I) translation model. The framework enables a typical classification model that was trained and tested on boiling images from domain A to predict boiling images coming from domain B that was never seen by the classification model. This is done by using the UI2I model to transform the domain B images to look like domain A images that the classification model is familiar with. Although CNN was used as the classification model and Fixed-Point GAN (FP-GAN) was used as the UI2I model, the framework is model agnostic. Meaning, that the framework can generalize any image classification model type, making it applicable to a variety of similar applications and not limited to the boiling crisis detection problem. It also means that the more the UI2I models advance, the better the performance of the framework.
Automated head and neck tumor segmentation from 3D PET/CT
Sep 22, 2022



Abstract:Head and neck tumor segmentation challenge (HECKTOR) 2022 offers a platform for researchers to compare their solutions to segmentation of tumors and lymph nodes from 3D CT and PET images. In this work, we describe our solution to HECKTOR 2022 segmentation task. We re-sample all images to a common resolution, crop around head and neck region, and train SegResNet semantic segmentation network from MONAI. We use 5-fold cross validation to select best model checkpoints. The final submission is an ensemble of 15 models from 3 runs. Our solution (team name NVAUTO) achieves the 1st place on the HECKTOR22 challenge leaderboard with an aggregated dice score of 0.78802.
Automated segmentation of intracranial hemorrhages from 3D CT
Sep 21, 2022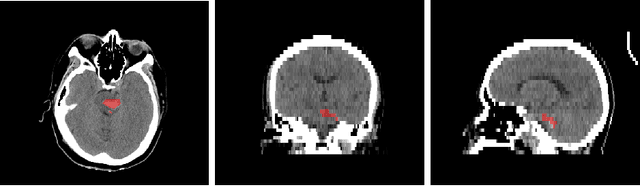
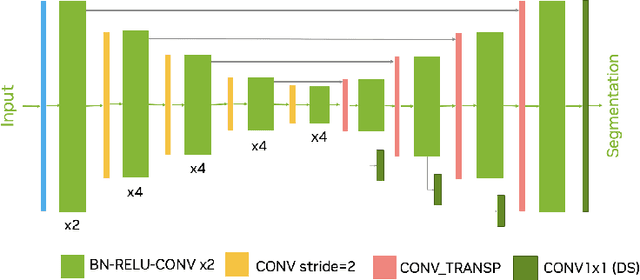
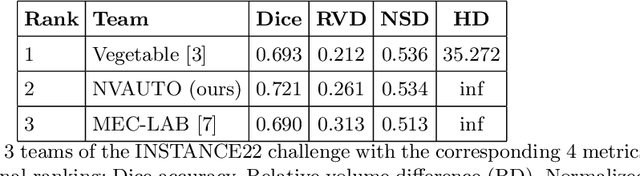
Abstract:Intracranial hemorrhage segmentation challenge (INSTANCE 2022) offers a platform for researchers to compare their solutions to segmentation of hemorrhage stroke regions from 3D CTs. In this work, we describe our solution to INSTANCE 2022. We use a 2D segmentation network, SegResNet from MONAI, operating slice-wise without resampling. The final submission is an ensemble of 18 models. Our solution (team name NVAUTO) achieves the top place in terms of Dice metric (0.721), and overall rank 2. It is implemented with Auto3DSeg.
HealthyGAN: Learning from Unannotated Medical Images to Detect Anomalies Associated with Human Disease
Sep 05, 2022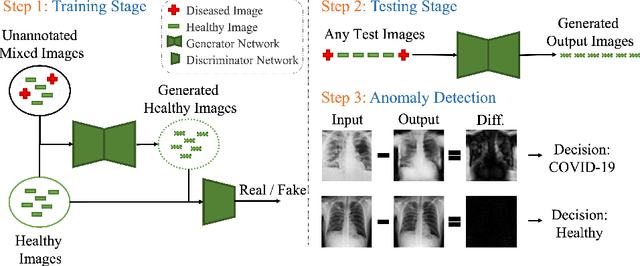
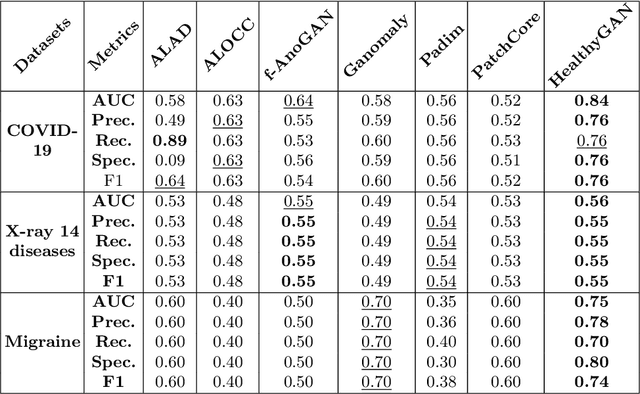
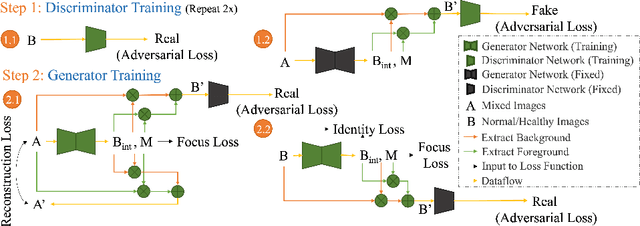
Abstract:Automated anomaly detection from medical images, such as MRIs and X-rays, can significantly reduce human effort in disease diagnosis. Owing to the complexity of modeling anomalies and the high cost of manual annotation by domain experts (e.g., radiologists), a typical technique in the current medical imaging literature has focused on deriving diagnostic models from healthy subjects only, assuming the model will detect the images from patients as outliers. However, in many real-world scenarios, unannotated datasets with a mix of both healthy and diseased individuals are abundant. Therefore, this paper poses the research question of how to improve unsupervised anomaly detection by utilizing (1) an unannotated set of mixed images, in addition to (2) the set of healthy images as being used in the literature. To answer the question, we propose HealthyGAN, a novel one-directional image-to-image translation method, which learns to translate the images from the mixed dataset to only healthy images. Being one-directional, HealthyGAN relaxes the requirement of cycle consistency of existing unpaired image-to-image translation methods, which is unattainable with mixed unannotated data. Once the translation is learned, we generate a difference map for any given image by subtracting its translated output. Regions of significant responses in the difference map correspond to potential anomalies (if any). Our HealthyGAN outperforms the conventional state-of-the-art methods by significant margins on two publicly available datasets: COVID-19 and NIH ChestX-ray14, and one institutional dataset collected from Mayo Clinic. The implementation is publicly available at https://github.com/mahfuzmohammad/HealthyGAN.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge