Waldo Valenzuela
From Claims to Evidence: A Unified Framework and Critical Analysis of CNN vs. Transformer vs. Mamba in Medical Image Segmentation
Mar 03, 2025



Abstract:While numerous architectures for medical image segmentation have been proposed, achieving competitive performance with state-of-the-art models networks such as nnUNet, still leave room for further innovation. In this work, we introduce nnUZoo, an open source benchmarking framework built upon nnUNet, which incorporates various deep learning architectures, including CNNs, Transformers, and Mamba-based models. Using this framework, we provide a fair comparison to demystify performance claims across different medical image segmentation tasks. Additionally, in an effort to enrich the benchmarking, we explored five new architectures based on Mamba and Transformers, collectively named X2Net, and integrated them into nnUZoo for further evaluation. The proposed models combine the features of conventional U2Net, nnUNet, CNN, Transformer, and Mamba layers and architectures, called X2Net (UNETR2Net (UNETR), SwT2Net (SwinTransformer), SS2D2Net (SwinUMamba), Alt1DM2Net (LightUMamba), and MambaND2Net (MambaND)). We extensively evaluate the performance of different models on six diverse medical image segmentation datasets, including microscopy, ultrasound, CT, MRI, and PET, covering various body parts, organs, and labels. We compare their performance, in terms of dice score and computational efficiency, against their baseline models, U2Net, and nnUNet. CNN models like nnUNet and U2Net demonstrated both speed and accuracy, making them effective choices for medical image segmentation tasks. Transformer-based models, while promising for certain imaging modalities, exhibited high computational costs. Proposed Mamba-based X2Net architecture (SS2D2Net) achieved competitive accuracy with no significantly difference from nnUNet and U2Net, while using fewer parameters. However, they required significantly longer training time, highlighting a trade-off between model efficiency and computational cost.
A Robust Ensemble Algorithm for Ischemic Stroke Lesion Segmentation: Generalizability and Clinical Utility Beyond the ISLES Challenge
Apr 03, 2024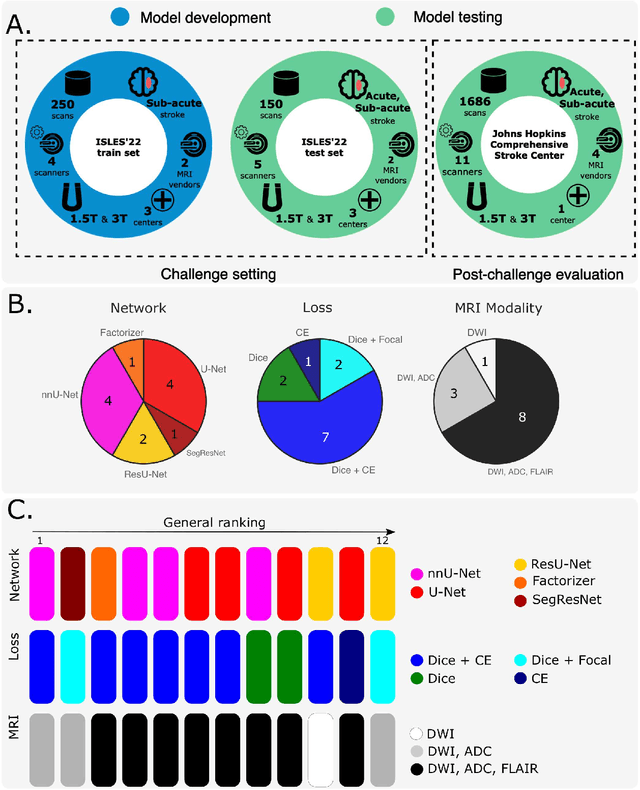
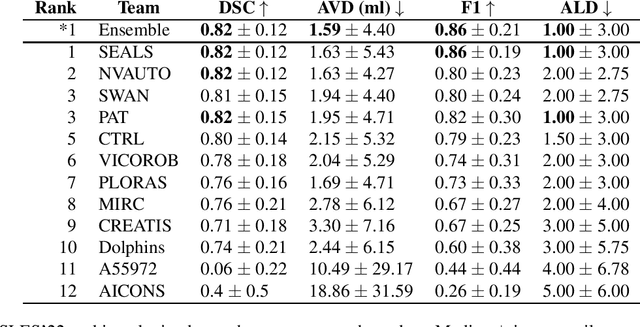
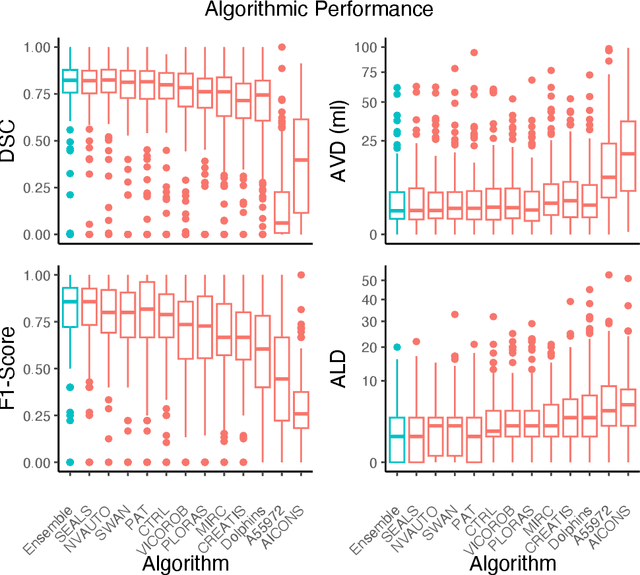
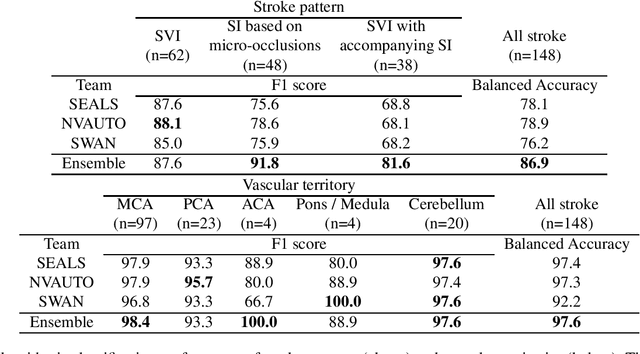
Abstract:Diffusion-weighted MRI (DWI) is essential for stroke diagnosis, treatment decisions, and prognosis. However, image and disease variability hinder the development of generalizable AI algorithms with clinical value. We address this gap by presenting a novel ensemble algorithm derived from the 2022 Ischemic Stroke Lesion Segmentation (ISLES) challenge. ISLES'22 provided 400 patient scans with ischemic stroke from various medical centers, facilitating the development of a wide range of cutting-edge segmentation algorithms by the research community. Through collaboration with leading teams, we combined top-performing algorithms into an ensemble model that overcomes the limitations of individual solutions. Our ensemble model achieved superior ischemic lesion detection and segmentation accuracy on our internal test set compared to individual algorithms. This accuracy generalized well across diverse image and disease variables. Furthermore, the model excelled in extracting clinical biomarkers. Notably, in a Turing-like test, neuroradiologists consistently preferred the algorithm's segmentations over manual expert efforts, highlighting increased comprehensiveness and precision. Validation using a real-world external dataset (N=1686) confirmed the model's generalizability. The algorithm's outputs also demonstrated strong correlations with clinical scores (admission NIHSS and 90-day mRS) on par with or exceeding expert-derived results, underlining its clinical relevance. This study offers two key findings. First, we present an ensemble algorithm (https://github.com/Tabrisrei/ISLES22_Ensemble) that detects and segments ischemic stroke lesions on DWI across diverse scenarios on par with expert (neuro)radiologists. Second, we show the potential for biomedical challenge outputs to extend beyond the challenge's initial objectives, demonstrating their real-world clinical applicability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge