Isaac Shiri
From Claims to Evidence: A Unified Framework and Critical Analysis of CNN vs. Transformer vs. Mamba in Medical Image Segmentation
Mar 03, 2025



Abstract:While numerous architectures for medical image segmentation have been proposed, achieving competitive performance with state-of-the-art models networks such as nnUNet, still leave room for further innovation. In this work, we introduce nnUZoo, an open source benchmarking framework built upon nnUNet, which incorporates various deep learning architectures, including CNNs, Transformers, and Mamba-based models. Using this framework, we provide a fair comparison to demystify performance claims across different medical image segmentation tasks. Additionally, in an effort to enrich the benchmarking, we explored five new architectures based on Mamba and Transformers, collectively named X2Net, and integrated them into nnUZoo for further evaluation. The proposed models combine the features of conventional U2Net, nnUNet, CNN, Transformer, and Mamba layers and architectures, called X2Net (UNETR2Net (UNETR), SwT2Net (SwinTransformer), SS2D2Net (SwinUMamba), Alt1DM2Net (LightUMamba), and MambaND2Net (MambaND)). We extensively evaluate the performance of different models on six diverse medical image segmentation datasets, including microscopy, ultrasound, CT, MRI, and PET, covering various body parts, organs, and labels. We compare their performance, in terms of dice score and computational efficiency, against their baseline models, U2Net, and nnUNet. CNN models like nnUNet and U2Net demonstrated both speed and accuracy, making them effective choices for medical image segmentation tasks. Transformer-based models, while promising for certain imaging modalities, exhibited high computational costs. Proposed Mamba-based X2Net architecture (SS2D2Net) achieved competitive accuracy with no significantly difference from nnUNet and U2Net, while using fewer parameters. However, they required significantly longer training time, highlighting a trade-off between model efficiency and computational cost.
Segmentation-Free Outcome Prediction in Head and Neck Cancer: Deep Learning-based Feature Extraction from Multi-Angle Maximum Intensity Projections (MA-MIPs) of PET Images
May 02, 2024



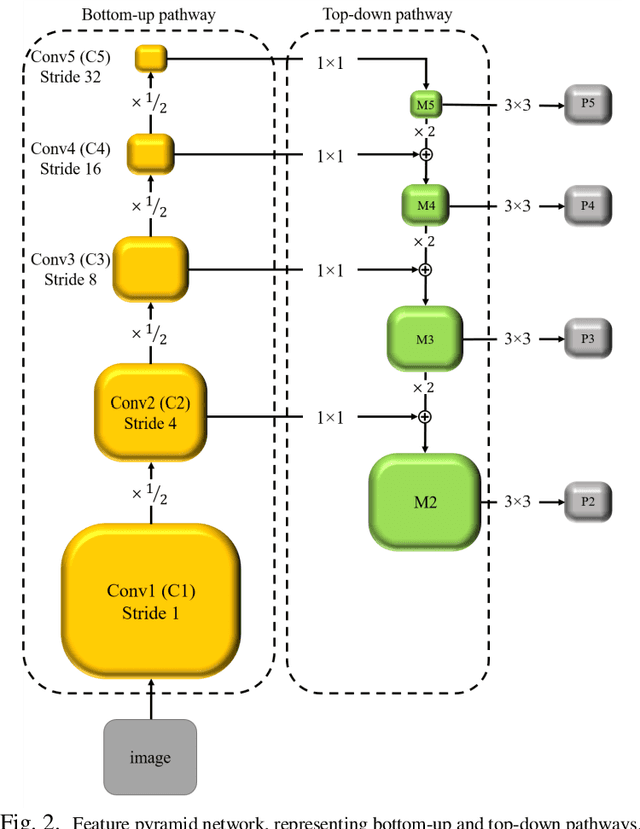
Abstract:We introduce an innovative, simple, effective segmentation-free approach for outcome prediction in head \& neck cancer (HNC) patients. By harnessing deep learning-based feature extraction techniques and multi-angle maximum intensity projections (MA-MIPs) applied to Fluorodeoxyglucose Positron Emission Tomography (FDG-PET) volumes, our proposed method eliminates the need for manual segmentations of regions-of-interest (ROIs) such as primary tumors and involved lymph nodes. Instead, a state-of-the-art object detection model is trained to perform automatic cropping of the head and neck region on the PET volumes. A pre-trained deep convolutional neural network backbone is then utilized to extract deep features from MA-MIPs obtained from 72 multi-angel axial rotations of the cropped PET volumes. These deep features extracted from multiple projection views of the PET volumes are then aggregated and fused, and employed to perform recurrence-free survival analysis on a cohort of 489 HNC patients. The proposed approach outperforms the best performing method on the target dataset for the task of recurrence-free survival analysis. By circumventing the manual delineation of the malignancies on the FDG PET-CT images, our approach eliminates the dependency on subjective interpretations and highly enhances the reproducibility of the proposed survival analysis method.
Semi-supervised learning towards automated segmentation of PET images with limited annotations: Application to lymphoma patients
Dec 24, 2022Abstract:The time-consuming task of manual segmentation challenges routine systematic quantification of disease burden. Convolutional neural networks (CNNs) hold significant promise to reliably identify locations and boundaries of tumors from PET scans. We aimed to leverage the need for annotated data via semi-supervised approaches, with application to PET images of diffuse large B-cell lymphoma (DLBCL) and primary mediastinal large B-cell lymphoma (PMBCL). We analyzed 18F-FDG PET images of 292 patients with PMBCL (n=104) and DLBCL (n=188) (n=232 for training and validation, and n=60 for external testing). We employed FCM and MS losses for training a 3D U-Net with different levels of supervision: i) fully supervised methods with labeled FCM (LFCM) as well as Unified focal and Dice loss functions, ii) unsupervised methods with Robust FCM (RFCM) and Mumford-Shah (MS) loss functions, and iii) Semi-supervised methods based on FCM (RFCM+LFCM), as well as MS loss in combination with supervised Dice loss (MS+Dice). Unified loss function yielded higher Dice score (mean +/- standard deviation (SD)) (0.73 +/- 0.03; 95% CI, 0.67-0.8) compared to Dice loss (p-value<0.01). Semi-supervised (RFCM+alpha*LFCM) with alpha=0.3 showed the best performance, with a Dice score of 0.69 +/- 0.03 (95% CI, 0.45-0.77) outperforming (MS+alpha*Dice) for any supervision level (any alpha) (p<0.01). The best performer among (MS+alpha*Dice) semi-supervised approaches with alpha=0.2 showed a Dice score of 0.60 +/- 0.08 (95% CI, 0.44-0.76) compared to another supervision level in this semi-supervised approach (p<0.01). Semi-supervised learning via FCM loss (RFCM+alpha*LFCM) showed improved performance compared to supervised approaches. Considering the time-consuming nature of expert manual delineations and intra-observer variabilities, semi-supervised approaches have significant potential for automated segmentation workflows.
Tensor Radiomics: Paradigm for Systematic Incorporation of Multi-Flavoured Radiomics Features
Mar 12, 2022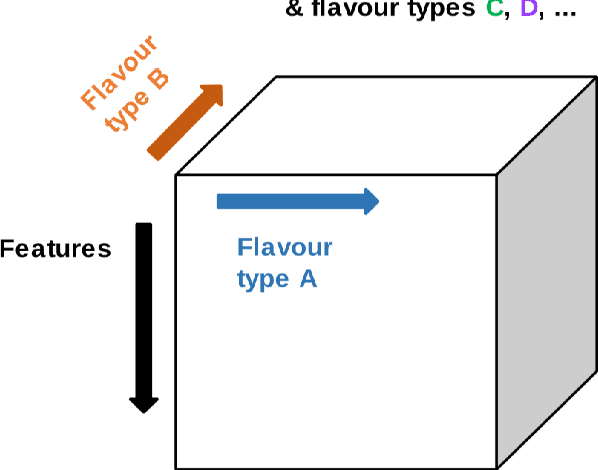
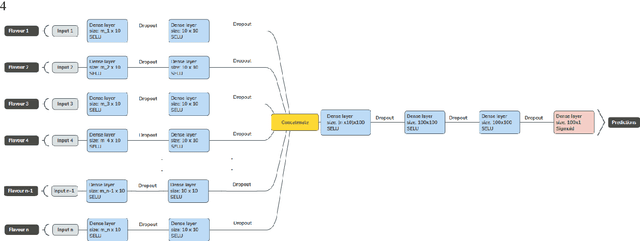
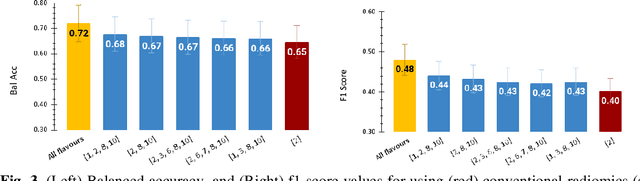
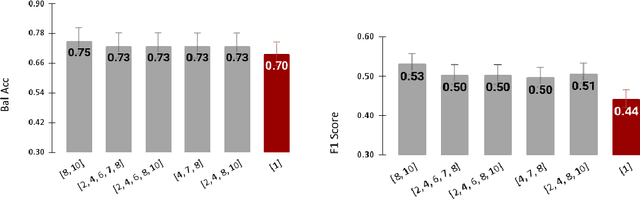
Abstract:Radiomics features extract quantitative information from medical images, towards the derivation of biomarkers for clinical tasks, such as diagnosis, prognosis, or treatment response assessment. Different image discretization parameters (e.g. bin number or size), convolutional filters, segmentation perturbation, or multi-modality fusion levels can be used to generate radiomics features and ultimately signatures. Commonly, only one set of parameters is used; resulting in only one value or flavour for a given RF. We propose tensor radiomics (TR) where tensors of features calculated with multiple combinations of parameters (i.e. flavours) are utilized to optimize the construction of radiomics signatures. We present examples of TR as applied to PET/CT, MRI, and CT imaging invoking machine learning or deep learning solutions, and reproducibility analyses: (1) TR via varying bin sizes on CT images of lung cancer and PET-CT images of head & neck cancer (HNC) for overall survival prediction. A hybrid deep neural network, referred to as TR-Net, along with two ML-based flavour fusion methods showed improved accuracy compared to regular rediomics features. (2) TR built from different segmentation perturbations and different bin sizes for classification of late-stage lung cancer response to first-line immunotherapy using CT images. TR improved predicted patient responses. (3) TR via multi-flavour generated radiomics features in MR imaging showed improved reproducibility when compared to many single-flavour features. (4) TR via multiple PET/CT fusions in HNC. Flavours were built from different fusions using methods, such as Laplacian pyramids and wavelet transforms. TR improved overall survival prediction. Our results suggest that the proposed TR paradigm has the potential to improve performance capabilities in different medical imaging tasks.
Non-Invasive MGMT Status Prediction in GBM Cancer Using Magnetic Resonance Images Radiomics Features: Univariate and Multivariate Machine Learning Radiogenomics Analysis
Jul 08, 2019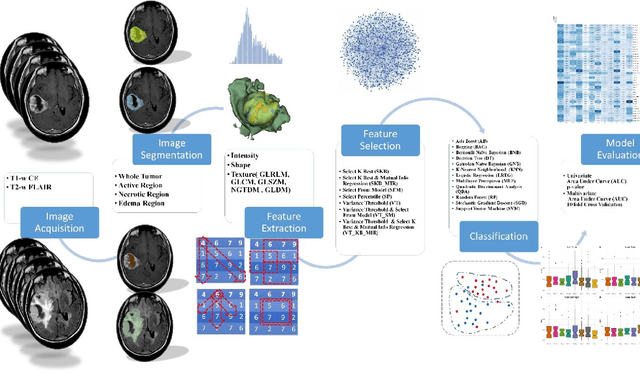
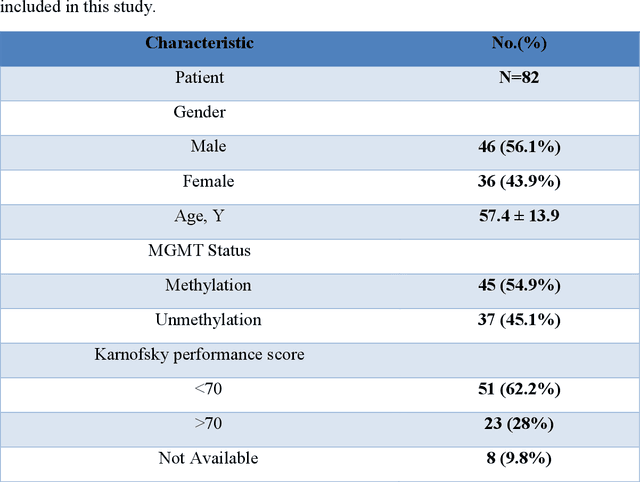
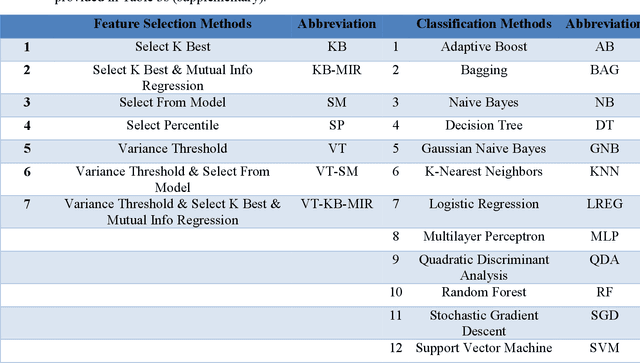
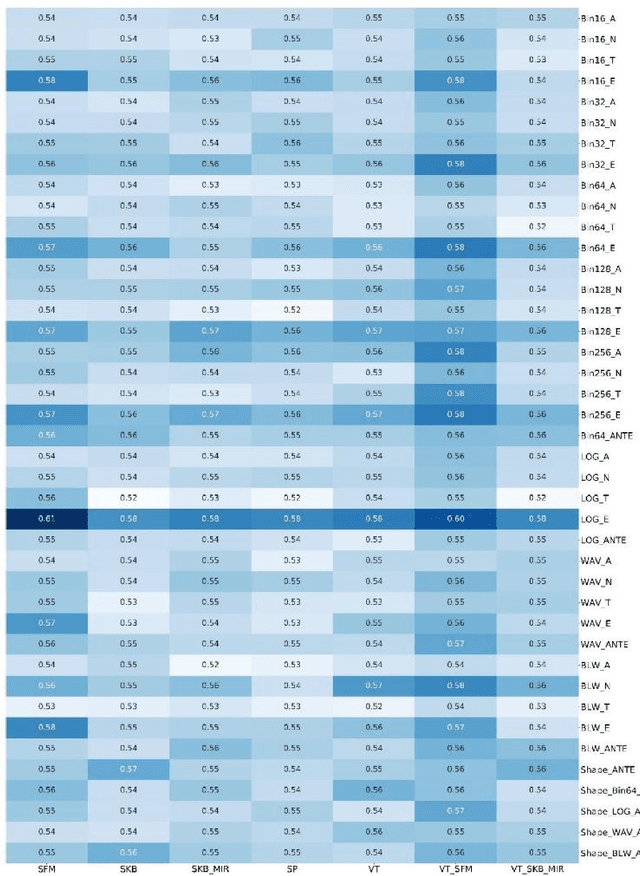
Abstract:Background and aim: This study aimed to predict methylation status of the O-6 methyl guanine-DNA methyl transferase (MGMT) gene promoter status by using MRI radiomics features, as well as univariate and multivariate analysis. Material and Methods: Eighty-two patients who had a MGMT methylation status were include in this study. Tumor were manually segmented in the four regions of MR images, a) whole tumor, b) active/enhanced region, c) necrotic regions and d) edema regions (E). About seven thousand radiomics features were extracted for each patient. Feature selection and classifier were used to predict MGMT status through different machine learning algorithms. The area under the curve (AUC) of receiver operating characteristic (ROC) curve was used for model evaluations. Results: Regarding univariate analysis, the Inverse Variance feature from gray level co-occurrence matrix (GLCM) in Whole Tumor segment with 4.5 mm Sigma of Laplacian of Gaussian filter with AUC: 0.71 (p-value: 0.002) was found to be the best predictor. For multivariate analysis, the decision tree classifier with Select from Model feature selector and LOG filter in Edema region had the highest performance (AUC: 0.78), followed by Ada Boost classifier with Select from Model feature selector and LOG filter in Edema region (AUC: 0.74). Conclusion: This study showed that radiomics using machine learning algorithms is a feasible, noninvasive approach to predict MGMT methylation status in GBM cancer patients Keywords: Radiomics, Radiogenomics, GBM, MRI, MGMT
Next Generation Radiogenomics Sequencing for Prediction of EGFR and KRAS Mutation Status in NSCLC Patients Using Multimodal Imaging and Machine Learning Approaches
Jul 03, 2019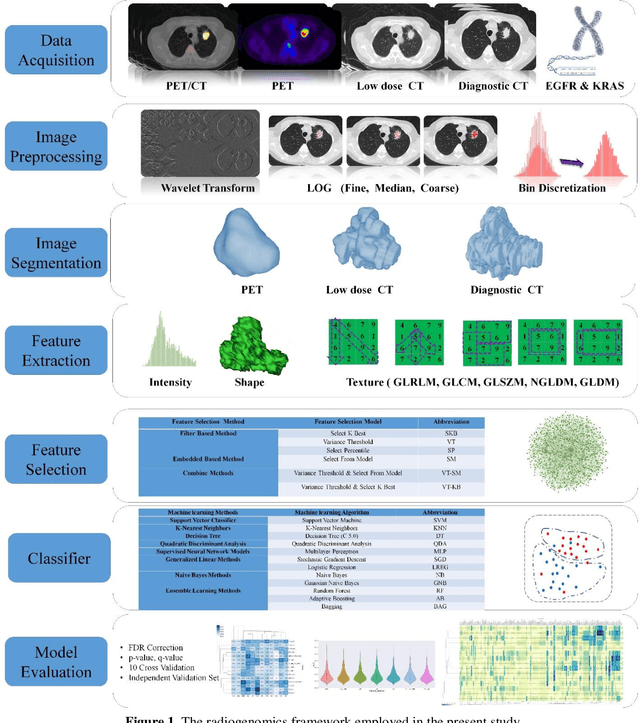
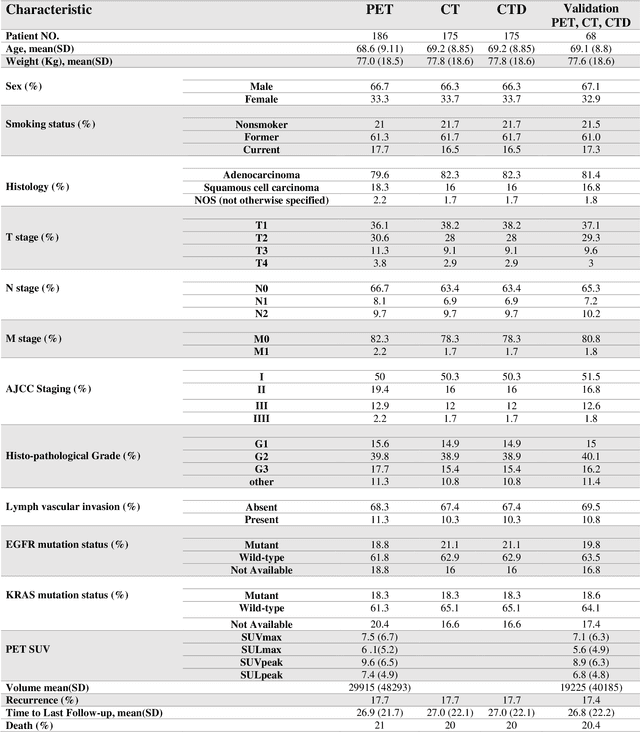

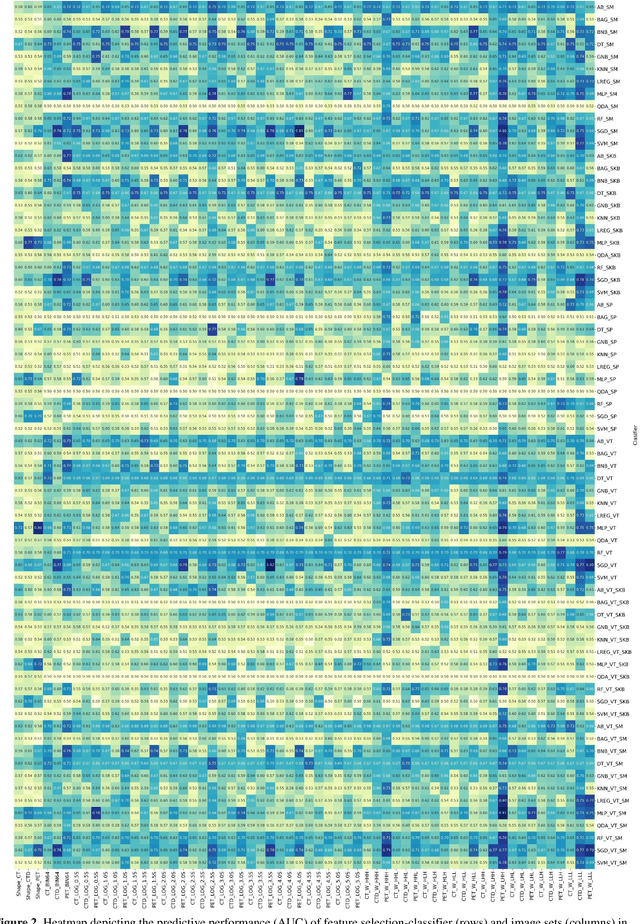
Abstract:Aim: In the present work, we aimed to evaluate a comprehensive radiomics framework that enabled prediction of EGFR and KRAS mutation status in NSCLC cancer patients based on PET and CT multi-modalities radiomic features and machine learning (ML) algorithms. Methods: Our study involved 211 NSCLC cancer patient with PET and CTD images. More than twenty thousand radiomic features from different image-feature sets were extracted Feature value was normalized to obtain Z-scores, followed by student t-test students for comparison, high correlated features were eliminated and the False discovery rate (FDR) correction were performed Six feature selection methods and twelve classifiers were used to predict gene status in patient and model evaluation was reported on independent validation sets (68 patients). Results: The best predictive power of conventional PET parameters was achieved by SUVpeak (AUC: 0.69, P-value = 0.0002) and MTV (AUC: 0.55, P-value = 0.0011) for EGFR and KRAS, respectively. Univariate analysis of radiomics features improved prediction power up to AUC: 75 (q-value: 0.003, Short Run Emphasis feature of GLRLM from LOG preprocessed image of PET with sigma value 1.5) and AUC: 0.71 (q-value 0.00005, The Large Dependence Low Gray Level Emphasis from GLDM in LOG preprocessed image of CTD sigma value 5) for EGFR and KRAS, respectively. Furthermore, the machine learning algorithm improved the perdition power up to AUC: 0.82 for EGFR (LOG preprocessed of PET image set with sigma 3 with VT feature selector and SGD classifier) and AUC: 0.83 for KRAS (CT image set with sigma 3.5 with SM feature selector and SGD classifier). Conclusion: We demonstrated that radiomic features extracted from different image-feature sets could be used for EGFR and KRAS mutation status prediction in NSCLC patients, and showed that they have more predictive power than conventional imaging parameters.
MFP-Unet: A Novel Deep Learning Based Approach for Left Ventricle Segmentation in Echocardiography
Jun 25, 2019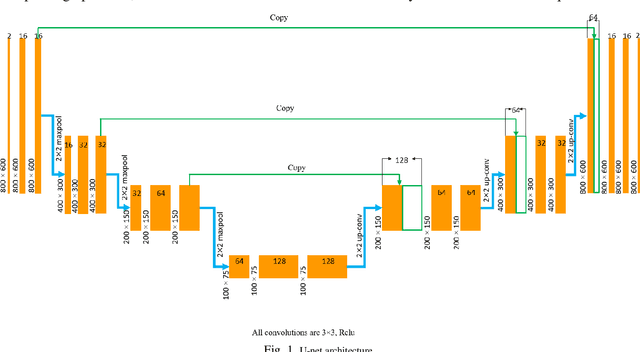
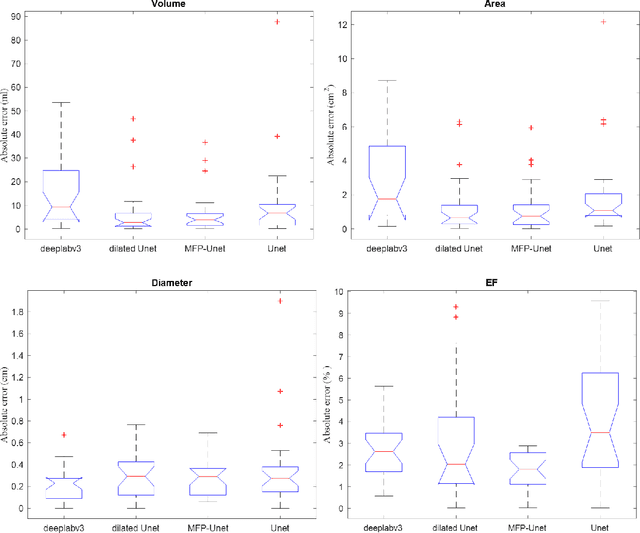
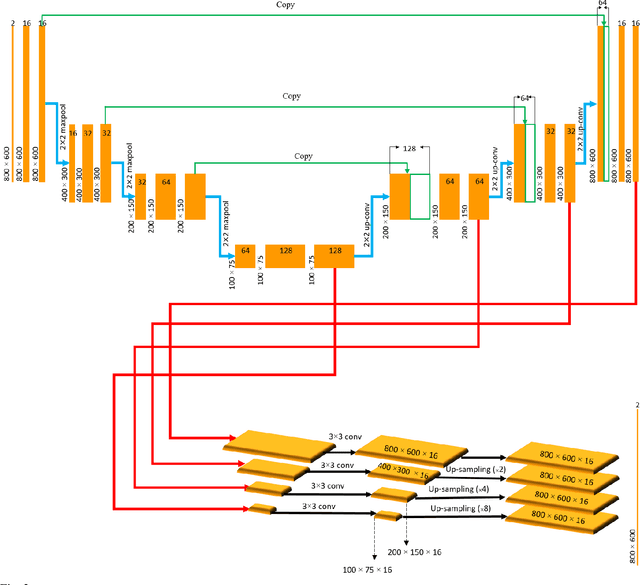
Abstract:Segmentation of the Left ventricle (LV) is a crucial step for quantitative measurements such as area, volume, and ejection fraction. However, the automatic LV segmentation in 2D echocardiographic images is a challenging task due to ill-defined borders, and operator dependence issues (insufficient reproducibility). U-net, which is a well-known architecture in medical image segmentation, addressed this problem through an encoder-decoder path. Despite outstanding overall performance, U-net ignores the contribution of all semantic strengths in the segmentation procedure. In the present study, we have proposed a novel architecture to tackle this drawback. Feature maps in all levels of the decoder path of U-net are concatenated, their depths are equalized, and up-sampled to a fixed dimension. This stack of feature maps would be the input of the semantic segmentation layer. The proposed network yielded state-of-the-art results when comparing with results from U-net, dilated U-net, and deeplabv3, using the same dataset. An average Dice Metric (DM) of 0.945, Hausdorff Distance (HD) of 1.62, Jaccard Coefficient (JC) of 0.97, and Mean Absolute Distance (MAD) of 1.32 are achieved. The correlation graph, bland-altman analysis, and box plot showed a great agreement between automatic and manually calculated volume, area, and length.
PET/CT Radiomic Sequencer for Prediction of EGFR and KRAS Mutation Status in NSCLC Patients
Jun 15, 2019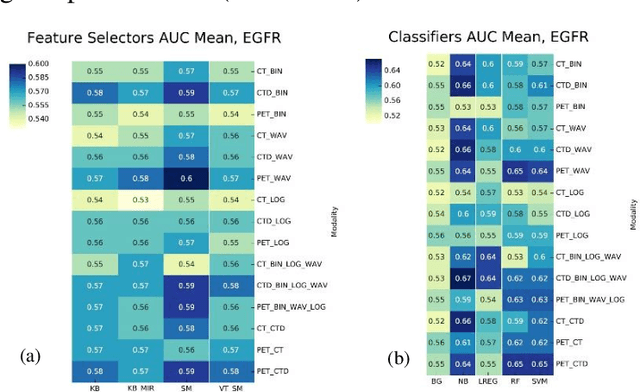
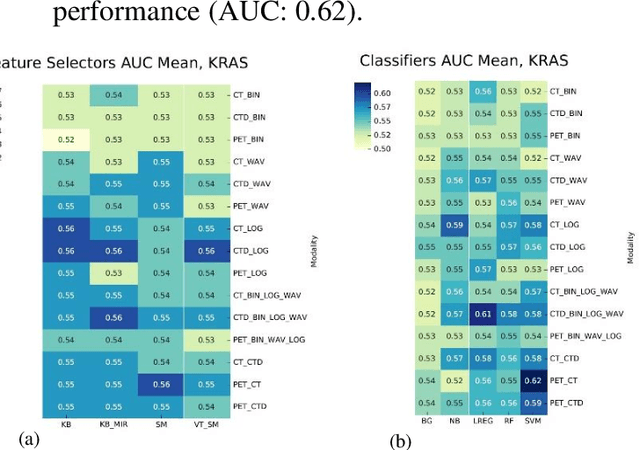
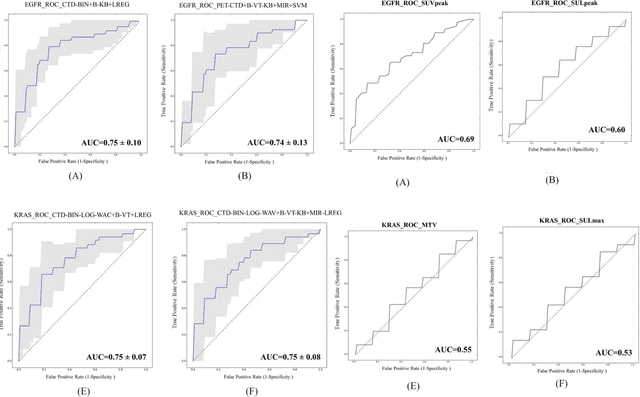
Abstract:The aim of this study was to develop radiomic models using PET/CT radiomic features with different machine learning approaches for finding best predictive epidermal growth factor receptor (EGFR) and Kirsten rat sarcoma viral oncogene (KRAS) mutation status. Patients images including PET and CT [diagnostic (CTD) and low dose CT (CTA)] were pre-processed using wavelet (WAV), Laplacian of Gaussian (LOG) and 64 bin discretization (BIN) (alone or in combinations) and several features from images were extracted. The prediction performance of model was checked using the area under the receiver operator characteristic (ROC) curve (AUC). Results showed a wide range of radiomic model AUC performances up to 0.75 in prediction of EGFR and KRAS mutation status. Combination of K-Best and variance threshold feature selector with logistic regression (LREG) classifier in diagnostic CT scan led to the best performance in EGFR (CTD-BIN+B-KB+LREG, AUC: mean 0.75 sd 0.10) and KRAS (CTD-BIN-LOG-WAV+B-VT+LREG, AUC: mean 0.75 sd 0.07) respectively. Additionally, incorporating PET, kept AUC values at ~0.74. When considering conventional features only, highest predictive performance was achieved by PET SUVpeak (AUC: 0.69) for EGFR and by PET MTV (AUC: 0.55) for KRAS. In comparison with conventional PET parameters such as standard uptake value, radiomic models were found as more predictive. Our findings demonstrated that non-invasive and reliable radiomics analysis can be successfully used to predict EGFR and KRAS mutation status in NSCLC patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge