Mehrdad Oveisi
AllMetrics: A Unified Python Library for Standardized Metric Evaluation and Robust Data Validation in Machine Learning
May 21, 2025Abstract:Machine learning (ML) models rely heavily on consistent and accurate performance metrics to evaluate and compare their effectiveness. However, existing libraries often suffer from fragmentation, inconsistent implementations, and insufficient data validation protocols, leading to unreliable results. Existing libraries have often been developed independently and without adherence to a unified standard, particularly concerning the specific tasks they aim to support. As a result, each library tends to adopt its conventions for metric computation, input/output formatting, error handling, and data validation protocols. This lack of standardization leads to both implementation differences (ID) and reporting differences (RD), making it difficult to compare results across frameworks or ensure reliable evaluations. To address these issues, we introduce AllMetrics, an open-source unified Python library designed to standardize metric evaluation across diverse ML tasks, including regression, classification, clustering, segmentation, and image-to-image translation. The library implements class-specific reporting for multi-class tasks through configurable parameters to cover all use cases, while incorporating task-specific parameters to resolve metric computation discrepancies across implementations. Various datasets from domains like healthcare, finance, and real estate were applied to our library and compared with Python, Matlab, and R components to identify which yield similar results. AllMetrics combines a modular Application Programming Interface (API) with robust input validation mechanisms to ensure reproducibility and reliability in model evaluation. This paper presents the design principles, architectural components, and empirical analyses demonstrating the ability to mitigate evaluation errors and to enhance the trustworthiness of ML workflows.
Machine Learning Evaluation Metric Discrepancies across Programming Languages and Their Components: Need for Standardization
Nov 18, 2024



Abstract:This study evaluates metrics for tasks such as classification, regression, clustering, correlation analysis, statistical tests, segmentation, and image-to-image (I2I) translation. Metrics were compared across Python libraries, R packages, and Matlab functions to assess their consistency and highlight discrepancies. The findings underscore the need for a unified roadmap to standardize metrics, ensuring reliable and reproducible ML evaluations across platforms. This study examined a wide range of evaluation metrics across various tasks and found only some to be consistent across platforms, such as (i) Accuracy, Balanced Accuracy, Cohens Kappa, F-beta Score, MCC, Geometric Mean, AUC, and Log Loss in binary classification; (ii) Accuracy, Cohens Kappa, and F-beta Score in multi-class classification; (iii) MAE, MSE, RMSE, MAPE, Explained Variance, Median AE, MSLE, and Huber in regression; (iv) Davies-Bouldin Index and Calinski-Harabasz Index in clustering; (v) Pearson, Spearman, Kendall's Tau, Mutual Information, Distance Correlation, Percbend, Shepherd, and Partial Correlation in correlation analysis; (vi) Paired t-test, Chi-Square Test, ANOVA, Kruskal-Wallis Test, Shapiro-Wilk Test, Welchs t-test, and Bartlett's test in statistical tests; (vii) Accuracy, Precision, and Recall in 2D segmentation; (viii) Accuracy in 3D segmentation; (ix) MAE, MSE, RMSE, and R-Squared in 2D-I2I translation; and (x) MAE, MSE, and RMSE in 3D-I2I translation. Given observation of discrepancies in a number of metrics (e.g. precision, recall and F1 score in binary classification, WCSS in clustering, multiple statistical tests, and IoU in segmentation, amongst multiple metrics), this study concludes that ML evaluation metrics require standardization and recommends that future research use consistent metrics for different tasks to effectively compare ML techniques and solutions.
Non-Invasive MGMT Status Prediction in GBM Cancer Using Magnetic Resonance Images Radiomics Features: Univariate and Multivariate Machine Learning Radiogenomics Analysis
Jul 08, 2019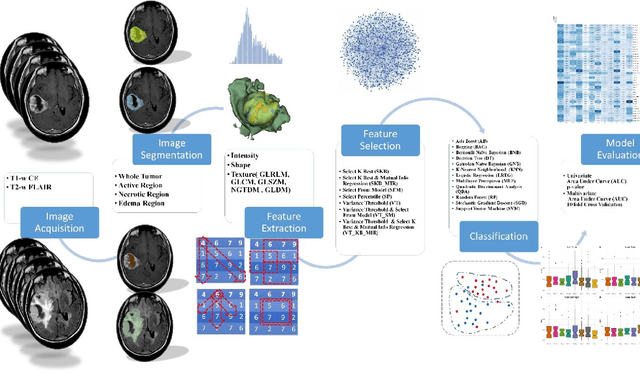
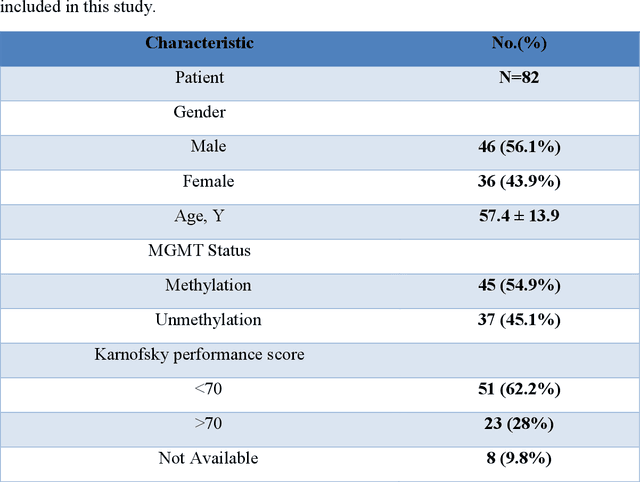
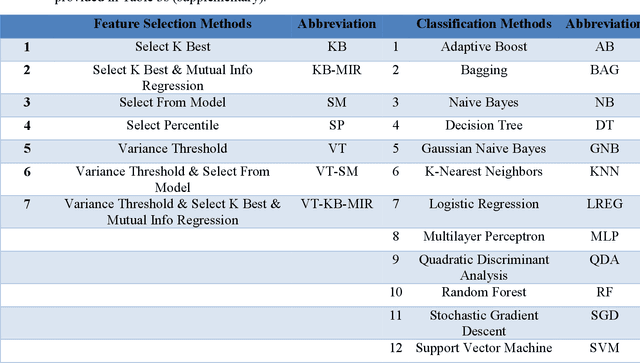
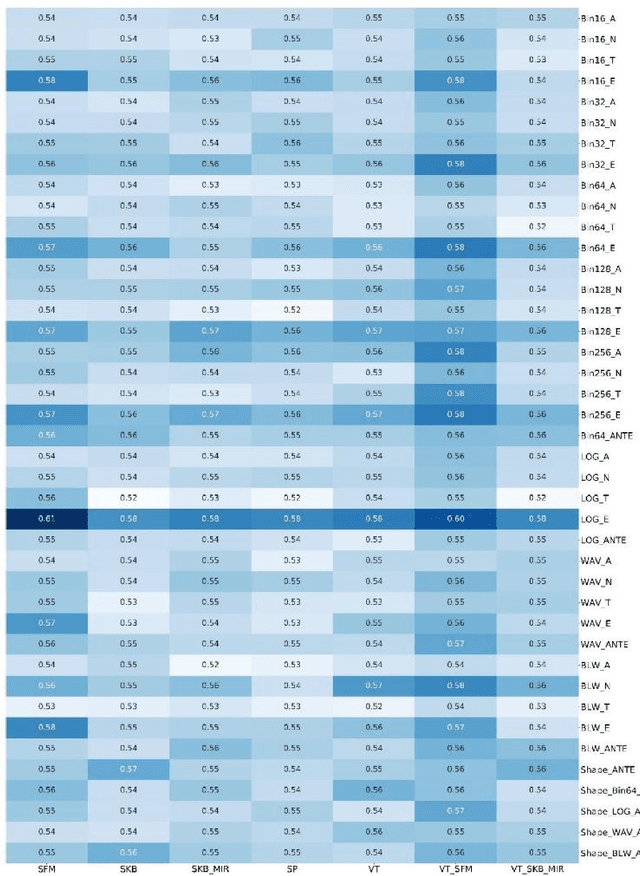
Abstract:Background and aim: This study aimed to predict methylation status of the O-6 methyl guanine-DNA methyl transferase (MGMT) gene promoter status by using MRI radiomics features, as well as univariate and multivariate analysis. Material and Methods: Eighty-two patients who had a MGMT methylation status were include in this study. Tumor were manually segmented in the four regions of MR images, a) whole tumor, b) active/enhanced region, c) necrotic regions and d) edema regions (E). About seven thousand radiomics features were extracted for each patient. Feature selection and classifier were used to predict MGMT status through different machine learning algorithms. The area under the curve (AUC) of receiver operating characteristic (ROC) curve was used for model evaluations. Results: Regarding univariate analysis, the Inverse Variance feature from gray level co-occurrence matrix (GLCM) in Whole Tumor segment with 4.5 mm Sigma of Laplacian of Gaussian filter with AUC: 0.71 (p-value: 0.002) was found to be the best predictor. For multivariate analysis, the decision tree classifier with Select from Model feature selector and LOG filter in Edema region had the highest performance (AUC: 0.78), followed by Ada Boost classifier with Select from Model feature selector and LOG filter in Edema region (AUC: 0.74). Conclusion: This study showed that radiomics using machine learning algorithms is a feasible, noninvasive approach to predict MGMT methylation status in GBM cancer patients Keywords: Radiomics, Radiogenomics, GBM, MRI, MGMT
Next Generation Radiogenomics Sequencing for Prediction of EGFR and KRAS Mutation Status in NSCLC Patients Using Multimodal Imaging and Machine Learning Approaches
Jul 03, 2019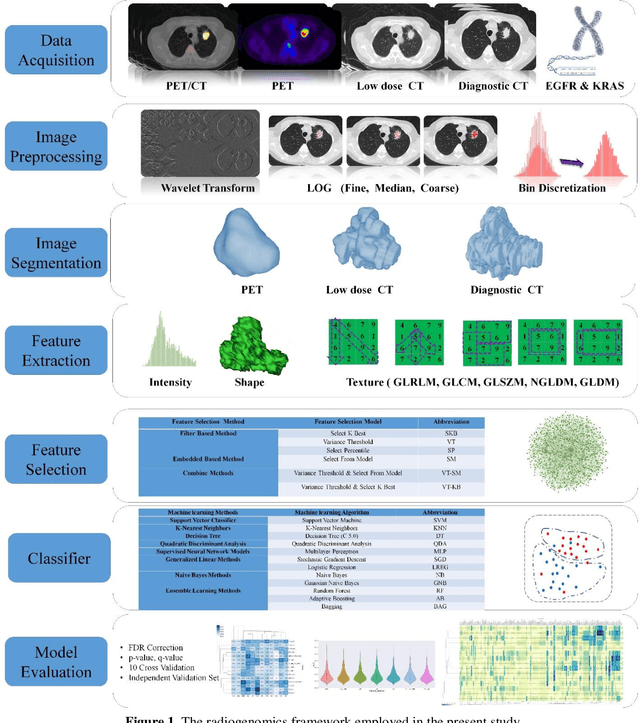
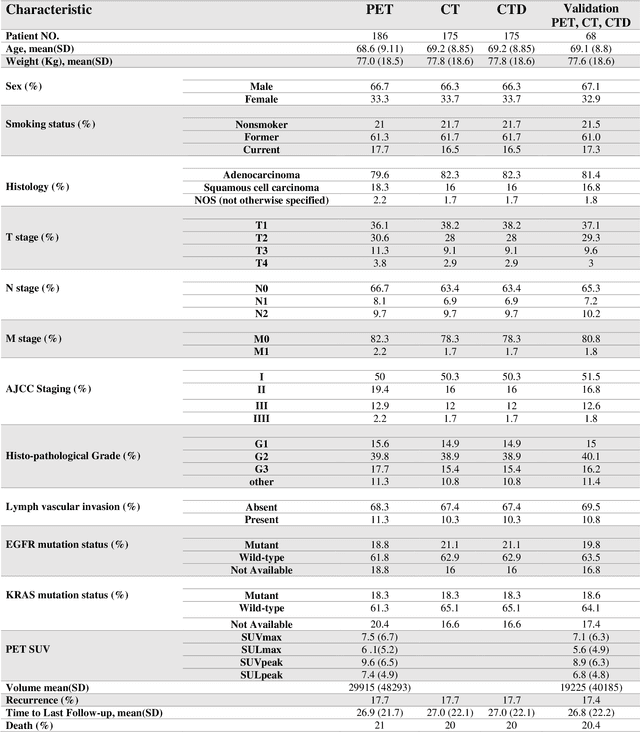

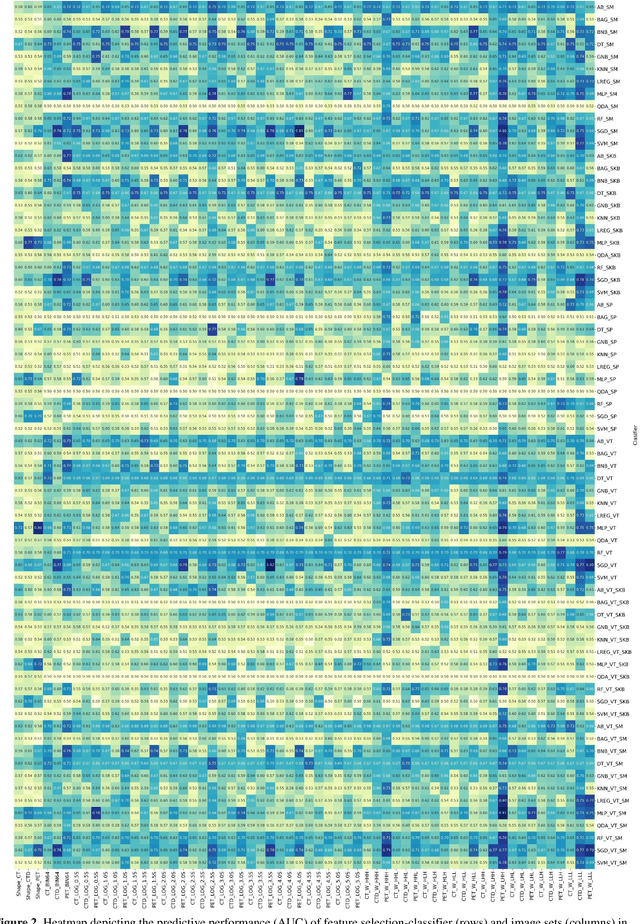
Abstract:Aim: In the present work, we aimed to evaluate a comprehensive radiomics framework that enabled prediction of EGFR and KRAS mutation status in NSCLC cancer patients based on PET and CT multi-modalities radiomic features and machine learning (ML) algorithms. Methods: Our study involved 211 NSCLC cancer patient with PET and CTD images. More than twenty thousand radiomic features from different image-feature sets were extracted Feature value was normalized to obtain Z-scores, followed by student t-test students for comparison, high correlated features were eliminated and the False discovery rate (FDR) correction were performed Six feature selection methods and twelve classifiers were used to predict gene status in patient and model evaluation was reported on independent validation sets (68 patients). Results: The best predictive power of conventional PET parameters was achieved by SUVpeak (AUC: 0.69, P-value = 0.0002) and MTV (AUC: 0.55, P-value = 0.0011) for EGFR and KRAS, respectively. Univariate analysis of radiomics features improved prediction power up to AUC: 75 (q-value: 0.003, Short Run Emphasis feature of GLRLM from LOG preprocessed image of PET with sigma value 1.5) and AUC: 0.71 (q-value 0.00005, The Large Dependence Low Gray Level Emphasis from GLDM in LOG preprocessed image of CTD sigma value 5) for EGFR and KRAS, respectively. Furthermore, the machine learning algorithm improved the perdition power up to AUC: 0.82 for EGFR (LOG preprocessed of PET image set with sigma 3 with VT feature selector and SGD classifier) and AUC: 0.83 for KRAS (CT image set with sigma 3.5 with SM feature selector and SGD classifier). Conclusion: We demonstrated that radiomic features extracted from different image-feature sets could be used for EGFR and KRAS mutation status prediction in NSCLC patients, and showed that they have more predictive power than conventional imaging parameters.
MFP-Unet: A Novel Deep Learning Based Approach for Left Ventricle Segmentation in Echocardiography
Jun 25, 2019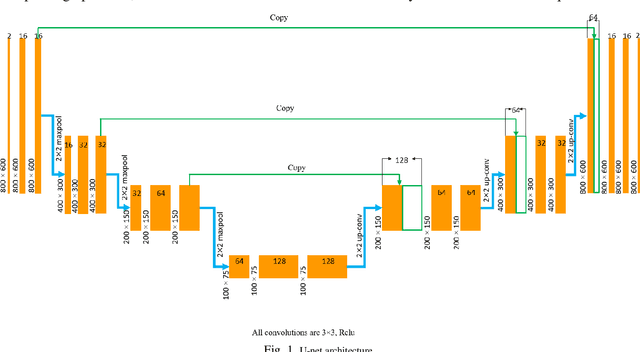
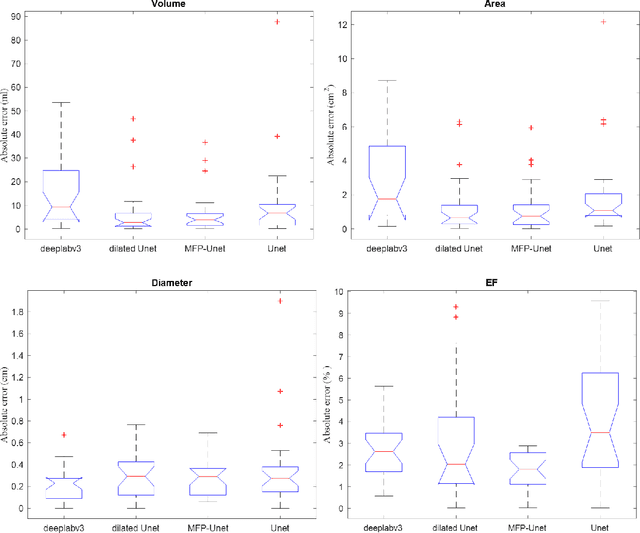
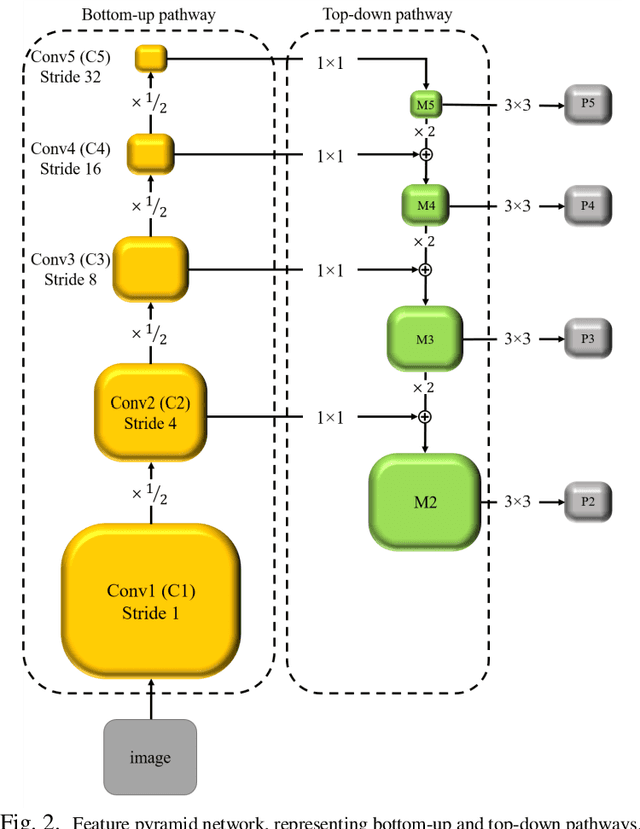
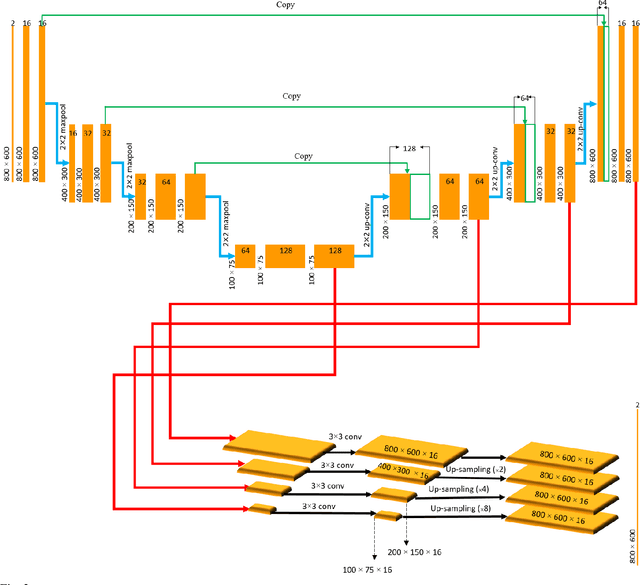
Abstract:Segmentation of the Left ventricle (LV) is a crucial step for quantitative measurements such as area, volume, and ejection fraction. However, the automatic LV segmentation in 2D echocardiographic images is a challenging task due to ill-defined borders, and operator dependence issues (insufficient reproducibility). U-net, which is a well-known architecture in medical image segmentation, addressed this problem through an encoder-decoder path. Despite outstanding overall performance, U-net ignores the contribution of all semantic strengths in the segmentation procedure. In the present study, we have proposed a novel architecture to tackle this drawback. Feature maps in all levels of the decoder path of U-net are concatenated, their depths are equalized, and up-sampled to a fixed dimension. This stack of feature maps would be the input of the semantic segmentation layer. The proposed network yielded state-of-the-art results when comparing with results from U-net, dilated U-net, and deeplabv3, using the same dataset. An average Dice Metric (DM) of 0.945, Hausdorff Distance (HD) of 1.62, Jaccard Coefficient (JC) of 0.97, and Mean Absolute Distance (MAD) of 1.32 are achieved. The correlation graph, bland-altman analysis, and box plot showed a great agreement between automatic and manually calculated volume, area, and length.
PET/CT Radiomic Sequencer for Prediction of EGFR and KRAS Mutation Status in NSCLC Patients
Jun 15, 2019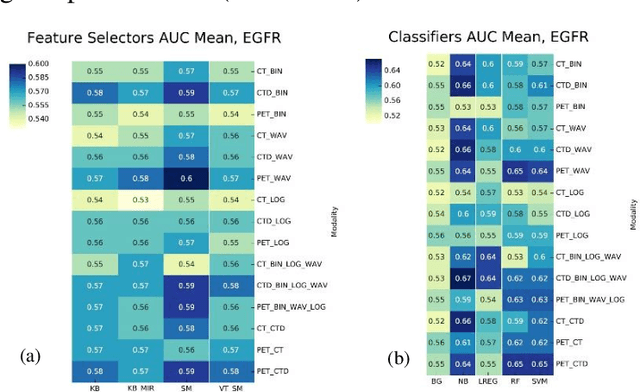
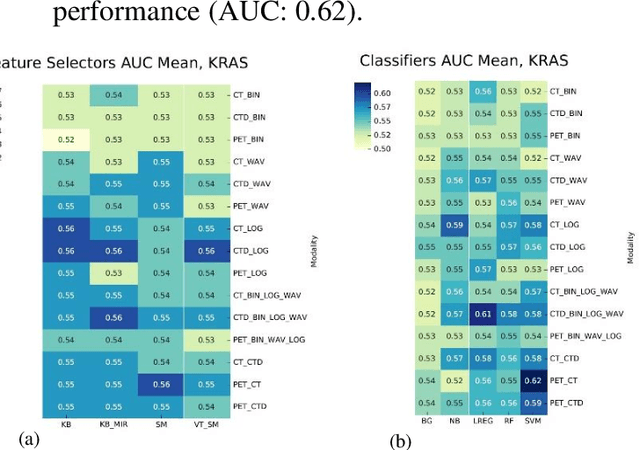
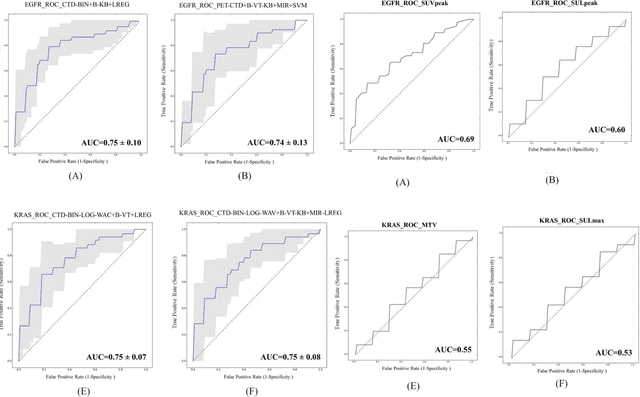
Abstract:The aim of this study was to develop radiomic models using PET/CT radiomic features with different machine learning approaches for finding best predictive epidermal growth factor receptor (EGFR) and Kirsten rat sarcoma viral oncogene (KRAS) mutation status. Patients images including PET and CT [diagnostic (CTD) and low dose CT (CTA)] were pre-processed using wavelet (WAV), Laplacian of Gaussian (LOG) and 64 bin discretization (BIN) (alone or in combinations) and several features from images were extracted. The prediction performance of model was checked using the area under the receiver operator characteristic (ROC) curve (AUC). Results showed a wide range of radiomic model AUC performances up to 0.75 in prediction of EGFR and KRAS mutation status. Combination of K-Best and variance threshold feature selector with logistic regression (LREG) classifier in diagnostic CT scan led to the best performance in EGFR (CTD-BIN+B-KB+LREG, AUC: mean 0.75 sd 0.10) and KRAS (CTD-BIN-LOG-WAV+B-VT+LREG, AUC: mean 0.75 sd 0.07) respectively. Additionally, incorporating PET, kept AUC values at ~0.74. When considering conventional features only, highest predictive performance was achieved by PET SUVpeak (AUC: 0.69) for EGFR and by PET MTV (AUC: 0.55) for KRAS. In comparison with conventional PET parameters such as standard uptake value, radiomic models were found as more predictive. Our findings demonstrated that non-invasive and reliable radiomics analysis can be successfully used to predict EGFR and KRAS mutation status in NSCLC patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge