Patrice Grehten
Pathological MRI Segmentation by Synthetic Pathological Data Generation in Fetuses and Neonates
Jan 31, 2025



Abstract:Developing new methods for the automated analysis of clinical fetal and neonatal MRI data is limited by the scarcity of annotated pathological datasets and privacy concerns that often restrict data sharing, hindering the effectiveness of deep learning models. We address this in two ways. First, we introduce Fetal&Neonatal-DDPM, a novel diffusion model framework designed to generate high-quality synthetic pathological fetal and neonatal MRIs from semantic label images. Second, we enhance training data by modifying healthy label images through morphological alterations to simulate conditions such as ventriculomegaly, cerebellar and pontocerebellar hypoplasia, and microcephaly. By leveraging Fetal&Neonatal-DDPM, we synthesize realistic pathological MRIs from these modified pathological label images. Radiologists rated the synthetic MRIs as significantly (p < 0.05) superior in quality and diagnostic value compared to real MRIs, demonstrating features such as blood vessels and choroid plexus, and improved alignment with label annotations. Synthetic pathological data enhanced state-of-the-art nnUNet segmentation performance, particularly for severe ventriculomegaly cases, with the greatest improvements achieved in ventricle segmentation (Dice scores: 0.9253 vs. 0.7317). This study underscores the potential of generative AI as transformative tool for data augmentation, offering improved segmentation performance in pathological cases. This development represents a significant step towards improving analysis and segmentation accuracy in prenatal imaging, and also offers new ways for data anonymization through the generation of pathologic image data.
A comparison of automatic multi-tissue segmentation methods of the human fetal brain using the FeTA Dataset
Oct 29, 2020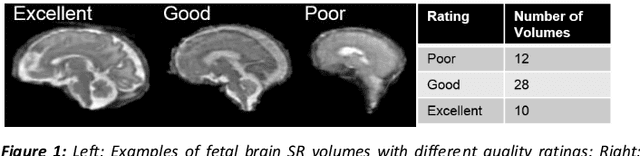
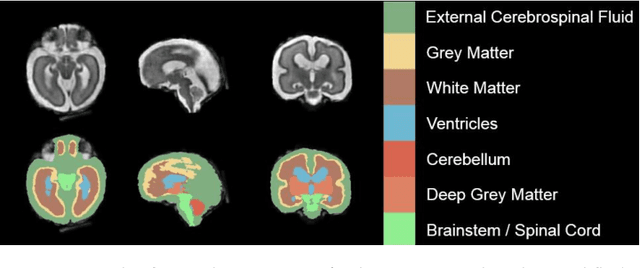
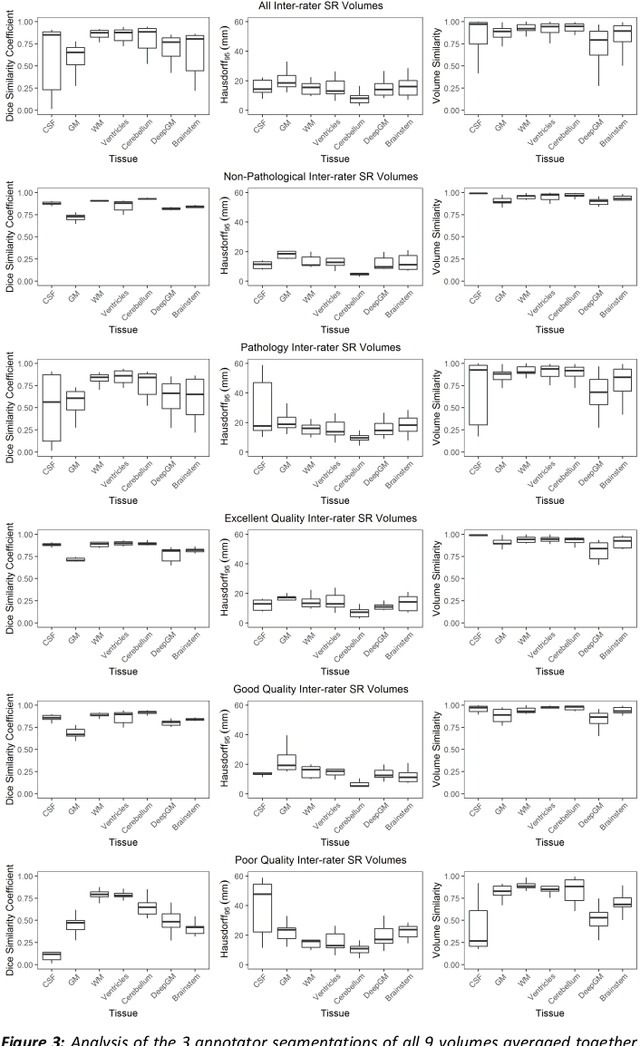
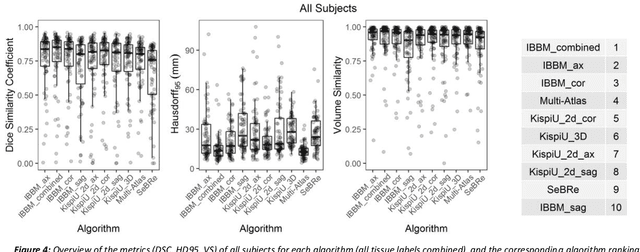
Abstract:It is critical to quantitatively analyse the developing human fetal brain in order to fully understand neurodevelopment in both normal fetuses and those with congenital disorders. To facilitate this analysis, automatic multi-tissue fetal brain segmentation algorithms are needed, which in turn requires open databases of segmented fetal brains. Here we introduce a publicly available database of 50 manually segmented pathological and non-pathological fetal magnetic resonance brain volume reconstructions across a range of gestational ages (20 to 33 weeks) into 7 different tissue categories (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, deep grey matter, brainstem/spinal cord). In addition, we quantitatively evaluate the accuracy of several automatic multi-tissue segmentation algorithms of the developing human fetal brain. Four research groups participated, submitting a total of 10 algorithms, demonstrating the benefits the database for the development of automatic algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge