Hamza Kebiri
Department of Radiology, Lausanne University Hospital, CIBM Center for Biomedical Imaging, Switzerland
Maximizing domain generalization in fetal brain tissue segmentation: the role of synthetic data generation, intensity clustering and real image fine-tuning
Nov 11, 2024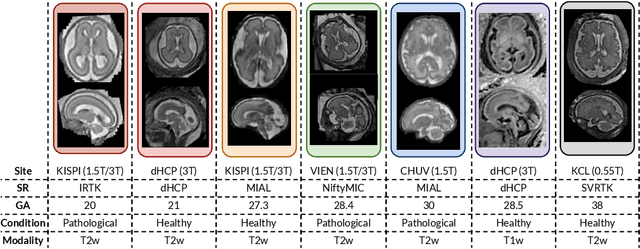
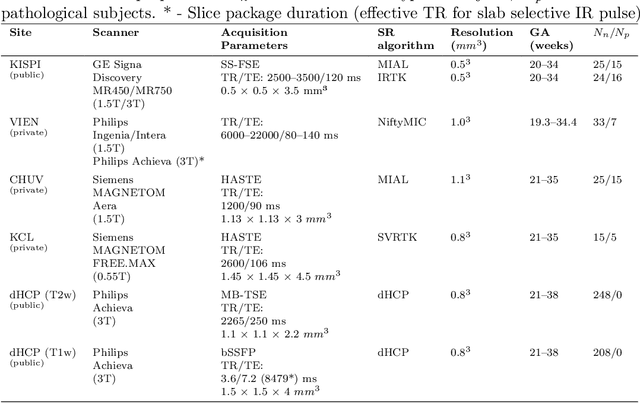
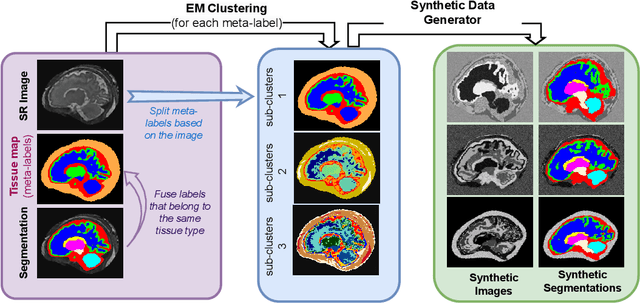
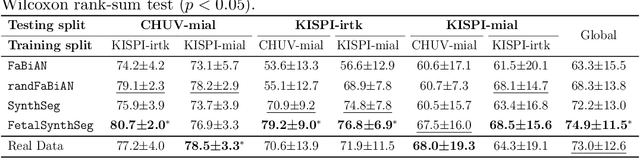
Abstract:Fetal brain tissue segmentation in magnetic resonance imaging (MRI) is a crucial tool that supports the understanding of neurodevelopment, yet it faces challenges due to the heterogeneity of data coming from different scanners and settings, and due to data scarcity. Recent approaches based on domain randomization, like SynthSeg, have shown a great potential for single source domain generalization, by simulating images with randomized contrast and image resolution from the label maps. In this work, we investigate how to maximize the out-of-domain (OOD) generalization potential of SynthSeg-based methods in fetal brain MRI. Specifically, when studying data generation, we demonstrate that the simple Gaussian mixture models used in SynthSeg enable more robust OOD generalization than physics-informed generation methods. We also investigate how intensity clustering can help create more faithful synthetic images, and observe that it is key to achieving a non-trivial OOD generalization capability when few label classes are available. Finally, by combining for the first time SynthSeg with modern fine-tuning approaches based on weight averaging, we show that fine-tuning a model pre-trained on synthetic data on a few real image-segmentation pairs in a new domain can lead to improvements in the target domain, but also in other domains. We summarize our findings as five key recommendations that we believe can guide practitioners who would like to develop SynthSeg-based approaches in other organs or modalities.
Ground-truth effects in learning-based fiber orientation distribution estimation in neonatal brains
Sep 02, 2024



Abstract:Diffusion Magnetic Resonance Imaging (dMRI) is a non-invasive method for depicting brain microstructure in vivo. Fiber orientation distributions (FODs) are mathematical representations extensively used to map white matter fiber configurations. Recently, FOD estimation with deep neural networks has seen growing success, in particular, those of neonates estimated with fewer diffusion measurements. These methods are mostly trained on target FODs reconstructed with multi-shell multi-tissue constrained spherical deconvolution (MSMT-CSD), which might not be the ideal ground truth for developing brains. Here, we investigate this hypothesis by training a state-of-the-art model based on the U-Net architecture on both MSMT-CSD and single-shell three-tissue constrained spherical deconvolution (SS3T-CSD). Our results suggest that SS3T-CSD might be more suited for neonatal brains, given that the ratio between single and multiple fiber-estimated voxels with SS3T-CSD is more realistic compared to MSMT-CSD. Additionally, increasing the number of input gradient directions significantly improves performance with SS3T-CSD over MSMT-CSD. Finally, in an age domain-shift setting, SS3T-CSD maintains robust performance across age groups, indicating its potential for more accurate neonatal brain imaging.
Improving cross-domain brain tissue segmentation in fetal MRI with synthetic data
Mar 22, 2024Abstract:Segmentation of fetal brain tissue from magnetic resonance imaging (MRI) plays a crucial role in the study of in utero neurodevelopment. However, automated tools face substantial domain shift challenges as they must be robust to highly heterogeneous clinical data, often limited in numbers and lacking annotations. Indeed, high variability of the fetal brain morphology, MRI acquisition parameters, and superresolution reconstruction (SR) algorithms adversely affect the model's performance when evaluated out-of-domain. In this work, we introduce FetalSynthSeg, a domain randomization method to segment fetal brain MRI, inspired by SynthSeg. Our results show that models trained solely on synthetic data outperform models trained on real data in out-ofdomain settings, validated on a 120-subject cross-domain dataset. Furthermore, we extend our evaluation to 40 subjects acquired using lowfield (0.55T) MRI and reconstructed with novel SR models, showcasing robustness across different magnetic field strengths and SR algorithms. Leveraging a generative synthetic approach, we tackle the domain shift problem in fetal brain MRI and offer compelling prospects for applications in fields with limited and highly heterogeneous data.
Cross-Age and Cross-Site Domain Shift Impacts on Deep Learning-Based White Matter Fiber Estimation in Newborn and Baby Brains
Dec 22, 2023Abstract:Deep learning models have shown great promise in estimating tissue microstructure from limited diffusion magnetic resonance imaging data. However, these models face domain shift challenges when test and train data are from different scanners and protocols, or when the models are applied to data with inherent variations such as the developing brains of infants and children scanned at various ages. Several techniques have been proposed to address some of these challenges, such as data harmonization or domain adaptation in the adult brain. However, those techniques remain unexplored for the estimation of fiber orientation distribution functions in the rapidly developing brains of infants. In this work, we extensively investigate the age effect and domain shift within and across two different cohorts of 201 newborns and 165 babies using the Method of Moments and fine-tuning strategies. Our results show that reduced variations in the microstructural development of babies in comparison to newborns directly impact the deep learning models' cross-age performance. We also demonstrate that a small number of target domain samples can significantly mitigate domain shift problems.
TBSS++: A novel computational method for Tract-Based Spatial Statistics
Jul 07, 2023Abstract:Diffusion-weighted magnetic resonance imaging (dMRI) is widely used to assess the brain white matter. One of the most common computations in dMRI involves cross-subject tract-specific analysis, whereby dMRI-derived biomarkers are compared between cohorts of subjects. The accuracy and reliability of these studies hinges on the ability to compare precisely the same white matter tracts across subjects. This is an intricate and error-prone computation. Existing computational methods such as Tract-Based Spatial Statistics (TBSS) suffer from a host of shortcomings and limitations that can seriously undermine the validity of the results. We present a new computational framework that overcomes the limitations of existing methods via (i) accurate segmentation of the tracts, and (ii) precise registration of data from different subjects/scans. The registration is based on fiber orientation distributions. To further improve the alignment of cross-subject data, we create detailed atlases of white matter tracts. These atlases serve as an unbiased reference space where the data from all subjects is registered for comparison. Extensive evaluations show that, compared with TBSS, our proposed framework offers significantly higher reproducibility and robustness to data perturbations. Our method promises a drastic improvement in accuracy and reproducibility of cross-subject dMRI studies that are routinely used in neuroscience and medical research.
Direct segmentation of brain white matter tracts in diffusion MRI
Jul 05, 2023



Abstract:The brain white matter consists of a set of tracts that connect distinct regions of the brain. Segmentation of these tracts is often needed for clinical and research studies. Diffusion-weighted MRI offers unique contrast to delineate these tracts. However, existing segmentation methods rely on intermediate computations such as tractography or estimation of fiber orientation density. These intermediate computations, in turn, entail complex computations that can result in unnecessary errors. Moreover, these intermediate computations often require dense multi-shell measurements that are unavailable in many clinical and research applications. As a result, current methods suffer from low accuracy and poor generalizability. Here, we propose a new deep learning method that segments these tracts directly from the diffusion MRI data, thereby sidestepping the intermediate computation errors. Our experiments show that this method can achieve segmentation accuracy that is on par with the state of the art methods (mean Dice Similarity Coefficient of 0.826). Compared with the state of the art, our method offers far superior generalizability to undersampled data that are typical of clinical studies and to data obtained with different acquisition protocols. Moreover, we propose a new method for detecting inaccurate segmentations and show that it is more accurate than standard methods that are based on estimation uncertainty quantification. The new methods can serve many critically important clinical and scientific applications that require accurate and reliable non-invasive segmentation of white matter tracts.
Spatio-temporal motion correction and iterative reconstruction of in-utero fetal fMRI
Sep 17, 2022Abstract:Resting-state functional Magnetic Resonance Imaging (fMRI) is a powerful imaging technique for studying functional development of the brain in utero. However, unpredictable and excessive movement of fetuses have limited its clinical applicability. Previous studies have focused primarily on the accurate estimation of the motion parameters employing a single step 3D interpolation at each individual time frame to recover a motion-free 4D fMRI image. Using only information from a 3D spatial neighborhood neglects the temporal structure of fMRI and useful information from neighboring timepoints. Here, we propose a novel technique based on four dimensional iterative reconstruction of the motion scattered fMRI slices. Quantitative evaluation of the proposed method on a cohort of real clinical fetal fMRI data indicates improvement of reconstruction quality compared to the conventional 3D interpolation approaches.
Slice estimation in diffusion MRI of neonatal and fetal brains in image and spherical harmonics domains using autoencoders
Aug 29, 2022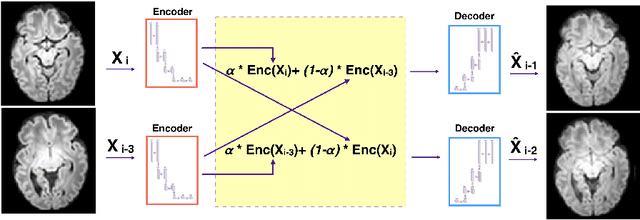
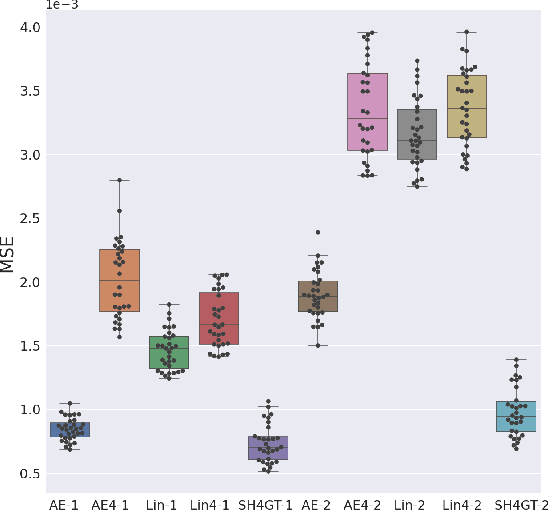
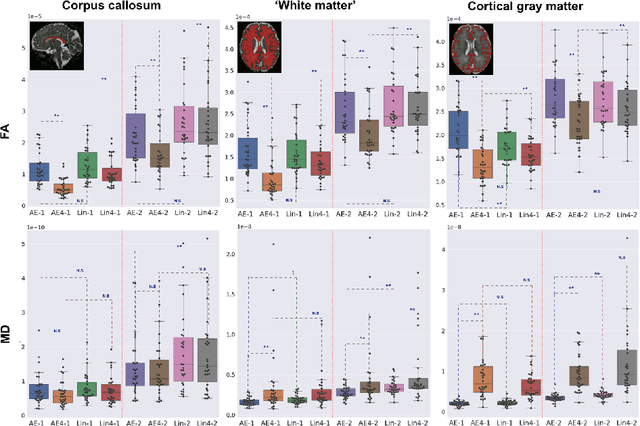
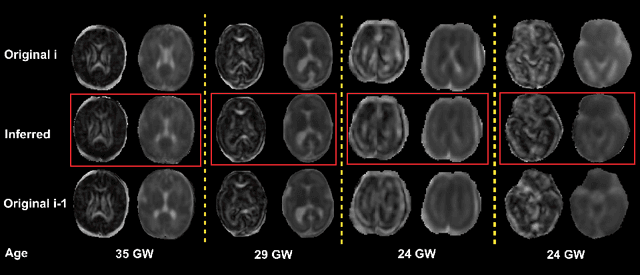
Abstract:Diffusion MRI (dMRI) of the developing brain can provide valuable insights into the white matter development. However, slice thickness in fetal dMRI is typically high (i.e., 3-5 mm) to freeze the in-plane motion, which reduces the sensitivity of the dMRI signal to the underlying anatomy. In this study, we aim at overcoming this problem by using autoencoders to learn unsupervised efficient representations of brain slices in a latent space, using raw dMRI signals and their spherical harmonics (SH) representation. We first learn and quantitatively validate the autoencoders on the developing Human Connectome Project pre-term newborn data, and further test the method on fetal data. Our results show that the autoencoder in the signal domain better synthesized the raw signal. Interestingly, the fractional anisotropy and, to a lesser extent, the mean diffusivity, are best recovered in missing slices by using the autoencoder trained with SH coefficients. A comparison was performed with the same maps reconstructed using an autoencoder trained with raw signals, as well as conventional interpolation methods of raw signals and SH coefficients. From these results, we conclude that the recovery of missing/corrupted slices should be performed in the signal domain if the raw signal is aimed to be recovered, and in the SH domain if diffusion tensor properties (i.e., fractional anisotropy) are targeted. Notably, the trained autoencoders were able to generalize to fetal dMRI data acquired using a much smaller number of diffusion gradients and a lower b-value, where we qualitatively show the consistency of the estimated diffusion tensor maps.
Multi-dimensional topological loss for cortical plate segmentation in fetal brain MRI
Aug 16, 2022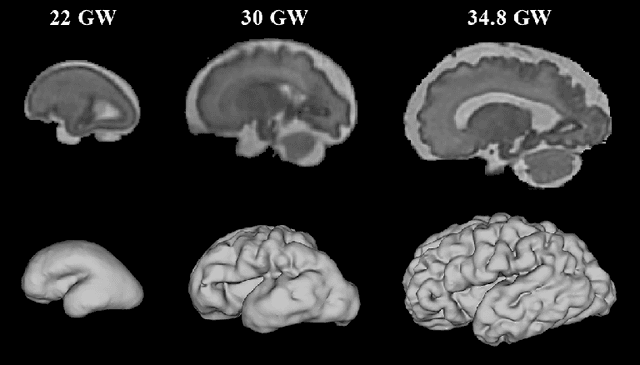
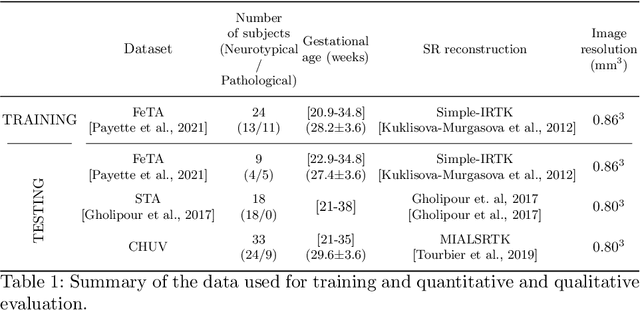
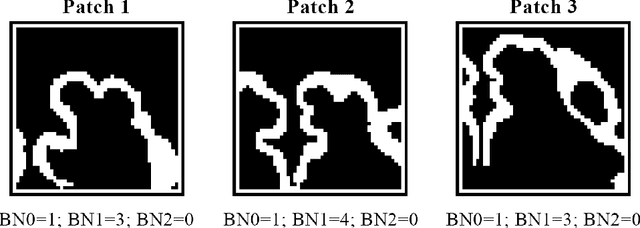
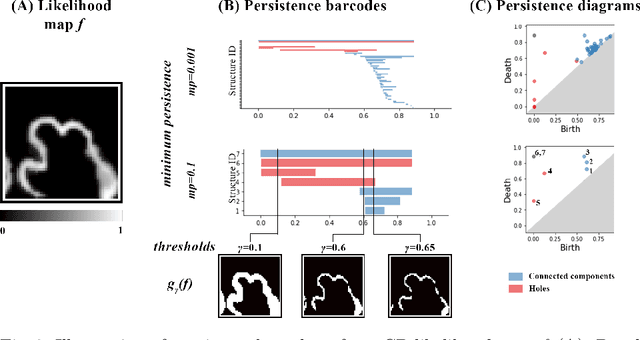
Abstract:The fetal cortical plate (CP) undergoes drastic morphological changes during the in utero development. Therefore, CP growth and folding patterns are key indicator in the assessment of the brain development and maturation. Magnetic resonance imaging (MRI) offers specific insights for the analysis of quantitative imaging biomarkers. Nonetheless, accurate and, more importantly, topologically correct MR image segmentation remains the key baseline to such analysis. In this study, we propose a deep learning segmentation framework for automatic and morphologically consistent segmentation of the CP in fetal brain MRI. Our contribution is two fold. First, we generalized a multi-dimensional topological loss function in order to enhance the topological accuracy. Second, we introduced hole ratio, a new topology-based validation measure that quantifies the size of the topological defects taking into account the size of the structure of interest. Using two publicly available datasets, we quantitatively evaluated our proposed method based on three complementary metrics which are overlap-, distance- and topology-based on 27 fetal brains. Our results evidence that our topology-integrative framework outperforms state-of-the-art training loss functions on super-resolution reconstructed clinical MRI, not only in shape correctness but also in the classical evaluation metrics. Furthermore, results on additional 31 out-of-domain SR reconstructions from clinical acquisitions were qualitatively assessed by three experts. The experts' consensus ranked our TopoCP method as the best segmentation in 100\% of the cases with a high inter-expert agreement. Overall, both quantitative and qualitative results, on a wide range of gestational ages and number of cases, support the generalizability and added value of our topology-guided framework for fetal CP segmentation.
Fetal Brain Tissue Annotation and Segmentation Challenge Results
Apr 20, 2022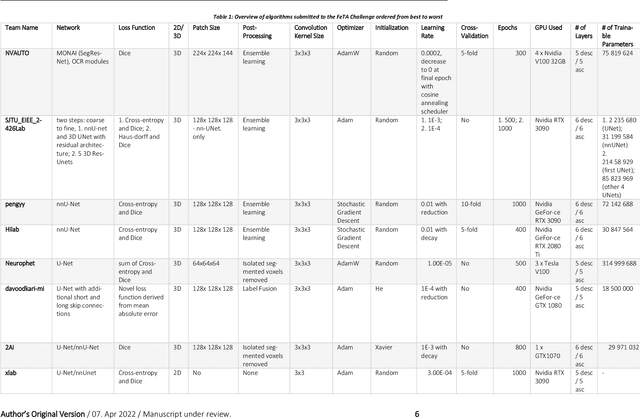
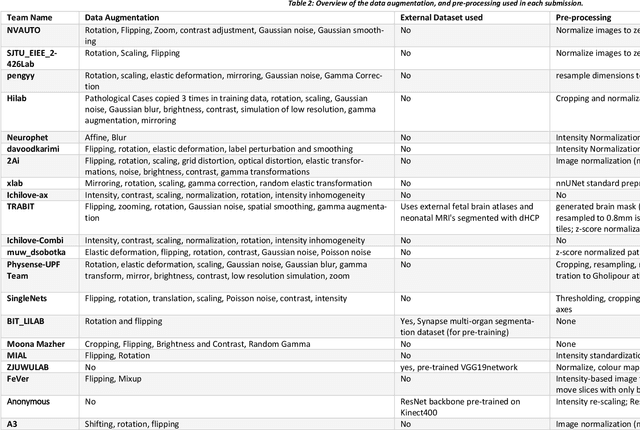
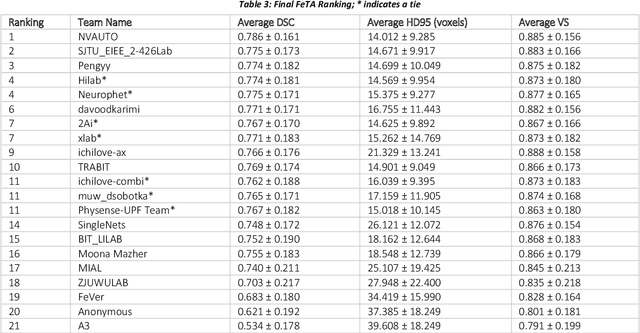
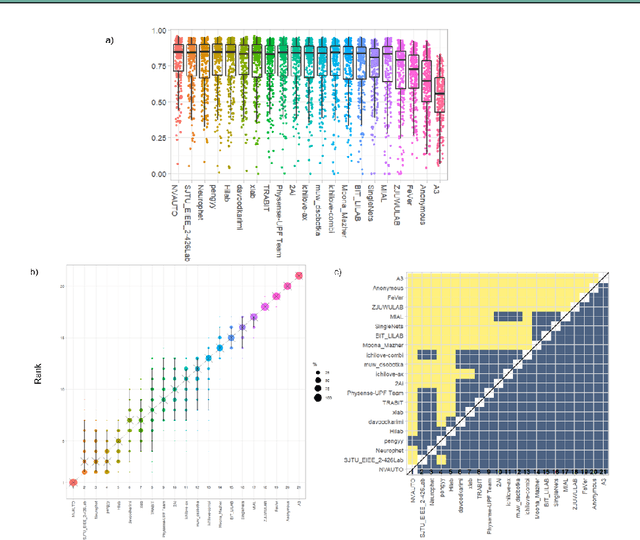
Abstract:In-utero fetal MRI is emerging as an important tool in the diagnosis and analysis of the developing human brain. Automatic segmentation of the developing fetal brain is a vital step in the quantitative analysis of prenatal neurodevelopment both in the research and clinical context. However, manual segmentation of cerebral structures is time-consuming and prone to error and inter-observer variability. Therefore, we organized the Fetal Tissue Annotation (FeTA) Challenge in 2021 in order to encourage the development of automatic segmentation algorithms on an international level. The challenge utilized FeTA Dataset, an open dataset of fetal brain MRI reconstructions segmented into seven different tissues (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, brainstem, deep grey matter). 20 international teams participated in this challenge, submitting a total of 21 algorithms for evaluation. In this paper, we provide a detailed analysis of the results from both a technical and clinical perspective. All participants relied on deep learning methods, mainly U-Nets, with some variability present in the network architecture, optimization, and image pre- and post-processing. The majority of teams used existing medical imaging deep learning frameworks. The main differences between the submissions were the fine tuning done during training, and the specific pre- and post-processing steps performed. The challenge results showed that almost all submissions performed similarly. Four of the top five teams used ensemble learning methods. However, one team's algorithm performed significantly superior to the other submissions, and consisted of an asymmetrical U-Net network architecture. This paper provides a first of its kind benchmark for future automatic multi-tissue segmentation algorithms for the developing human brain in utero.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge