Sebastien Tourbier
Spatio-temporal motion correction and iterative reconstruction of in-utero fetal fMRI
Sep 17, 2022Abstract:Resting-state functional Magnetic Resonance Imaging (fMRI) is a powerful imaging technique for studying functional development of the brain in utero. However, unpredictable and excessive movement of fetuses have limited its clinical applicability. Previous studies have focused primarily on the accurate estimation of the motion parameters employing a single step 3D interpolation at each individual time frame to recover a motion-free 4D fMRI image. Using only information from a 3D spatial neighborhood neglects the temporal structure of fMRI and useful information from neighboring timepoints. Here, we propose a novel technique based on four dimensional iterative reconstruction of the motion scattered fMRI slices. Quantitative evaluation of the proposed method on a cohort of real clinical fetal fMRI data indicates improvement of reconstruction quality compared to the conventional 3D interpolation approaches.
Weak labels and anatomical knowledge: making deep learning practical for intracranial aneurysm detection in TOF-MRA
Mar 10, 2021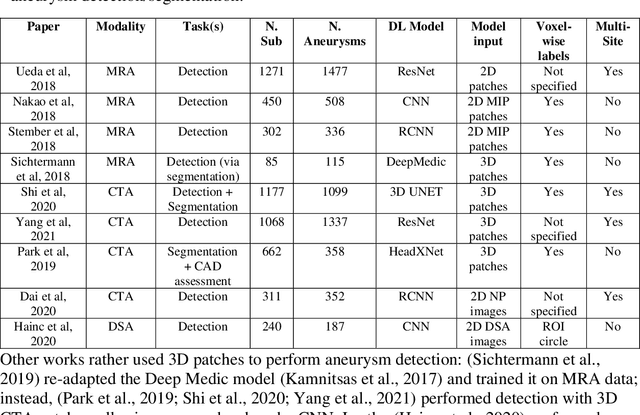
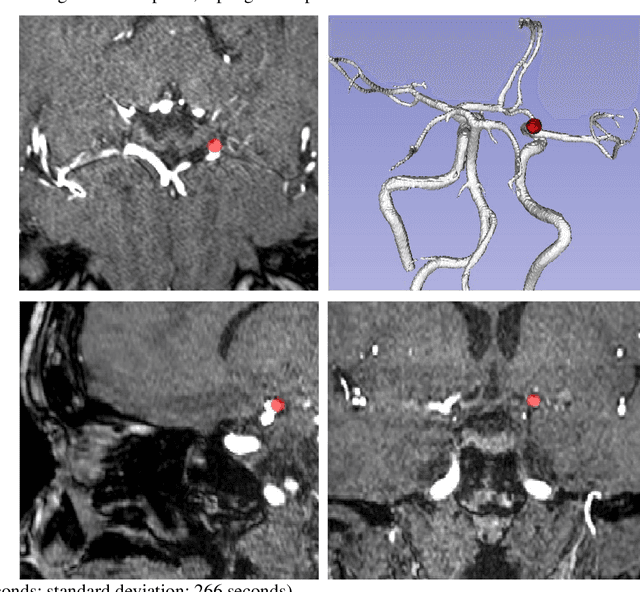
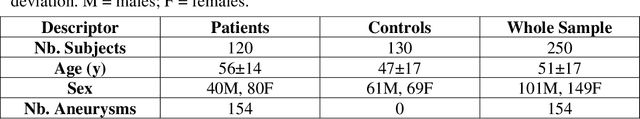
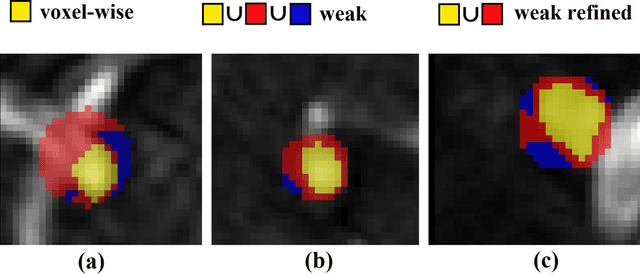
Abstract:Supervised segmentation algorithms yield state-of-the-art results for automated anomaly detection. However, these models require voxel-wise labels which are time-consuming to draw for medical experts. An interesting alternative to voxel-wise annotations is the use of weak labels: these can be coarse or oversized annotations that are less precise, but considerably faster to create. In this work, we address the task of brain aneurysm detection by developing a fully automated, deep neural network that is trained utilizing oversized weak labels. Furthermore, since aneurysms mainly occur in specific anatomical locations, we build our model leveraging the underlying anatomy of the brain vasculature both during training and inference. We apply our model to 250 subjects (120 patients, 130 controls) who underwent Time-Of-Flight Magnetic Resonance Angiography (TOF-MRA) and presented a total of 154 aneurysms. To assess the robustness of the algorithm, we participated in a MICCAI challenge for TOF-MRA data (93 patients, 20 controls, 125 aneurysms) which allowed us to obtain results also for subjects coming from a different institution. Our network achieves an average sensitivity of 77% on our in-house data, with a mean False Positive (FP) rate of 0.72 per patient. Instead, on the challenge data, we attain a sensitivity of 59% with a mean FP rate of 1.18, ranking in 7th/14 position for detection and in 4th/11 for segmentation on the open leaderboard. When computing detection performances with respect to aneurysms' risk of rupture, we found no statistical difference between two risk groups (p = 0.12), although the sensitivity for dangerous aneurysms was higher (78%). Our approach suggests that clinically useful sensitivity can be achieved using weak labels and exploiting prior anatomical knowledge; this expands the feasibility of deep learning studies to hospitals that have limited time and data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge