Yasser Alemán-Gómez
Department of Radiology, Lausanne University Hospital
FaBiAN: A Fetal Brain magnetic resonance Acquisition Numerical phantom
Sep 06, 2021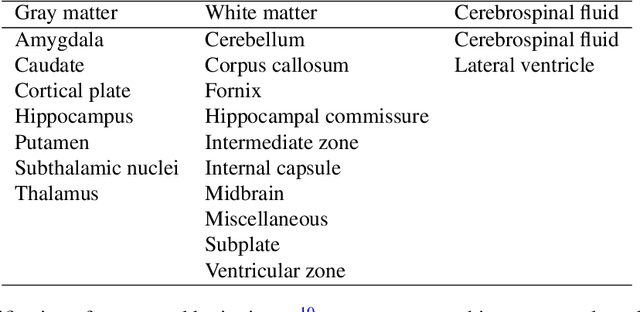
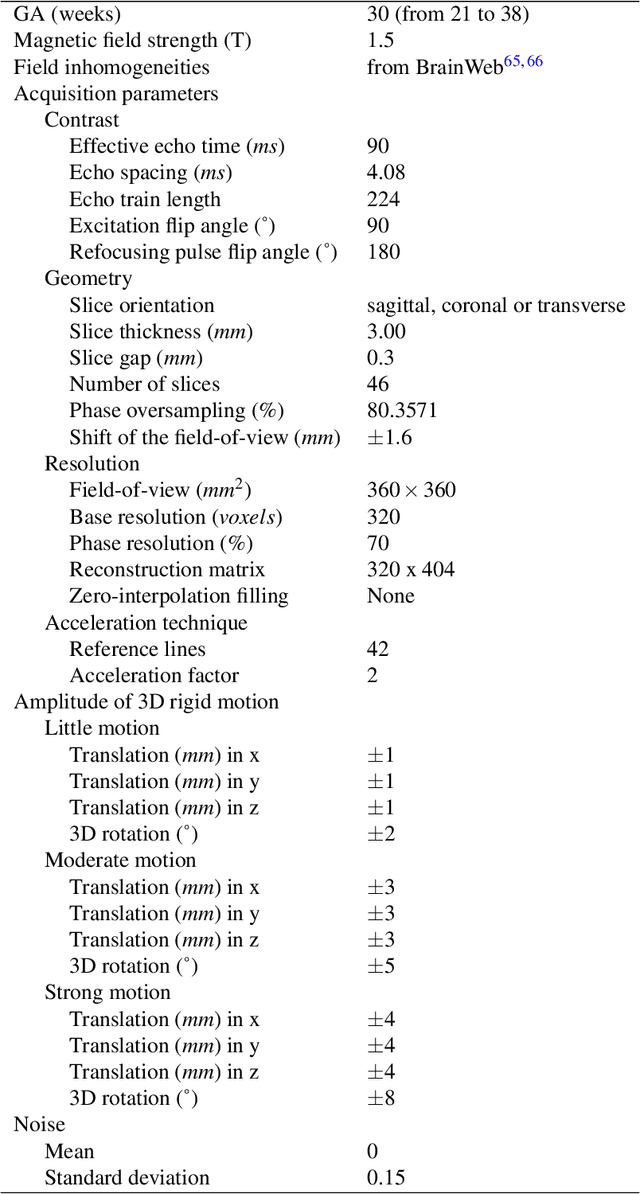
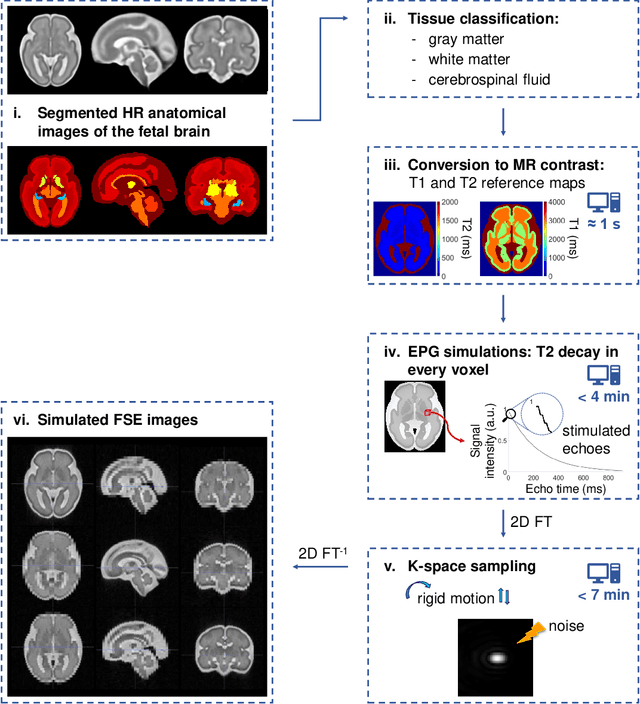

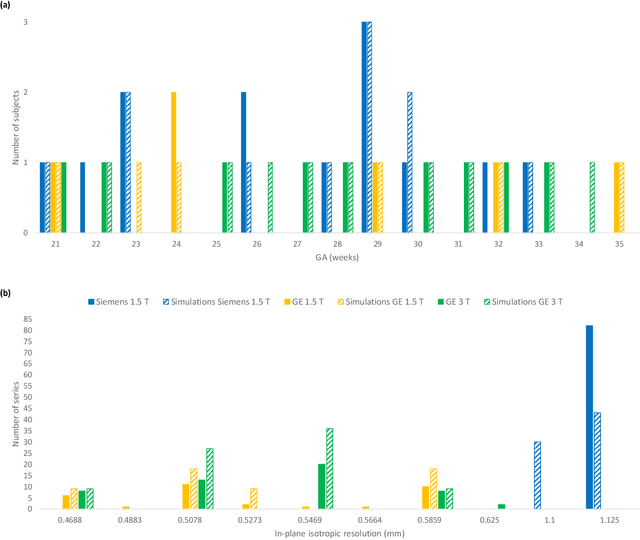
Abstract:Accurate characterization of in utero human brain maturation is critical as it involves complex and interconnected structural and functional processes that may influence health later in life. Magnetic resonance imaging is a powerful tool to investigate equivocal neurological patterns during fetal development. However, the number of acquisitions of satisfactory quality available in this cohort of sensitive subjects remains scarce, thus hindering the validation of advanced image processing techniques. Numerical phantoms can mitigate these limitations by providing a controlled environment with a known ground truth. In this work, we present FaBiAN, an open-source Fetal Brain magnetic resonance Acquisition Numerical phantom that simulates clinical T2-weighted fast spin echo sequences of the fetal brain. This unique tool is based on a general, flexible and realistic setup that includes stochastic fetal movements, thus providing images of the fetal brain throughout maturation comparable to clinical acquisitions. We demonstrate its value to evaluate the robustness and optimize the accuracy of an algorithm for super-resolution fetal brain magnetic resonance imaging from simulated motion-corrupted 2D low-resolution series as compared to a synthetic high-resolution reference volume. We also show that the images generated can complement clinical datasets to support data-intensive deep learning methods for fetal brain tissue segmentation.
Weak labels and anatomical knowledge: making deep learning practical for intracranial aneurysm detection in TOF-MRA
Mar 10, 2021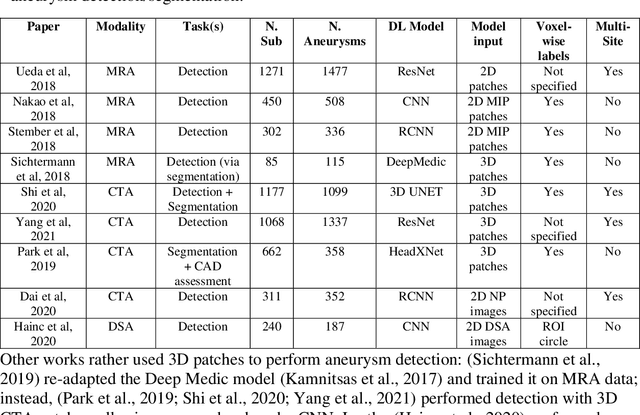
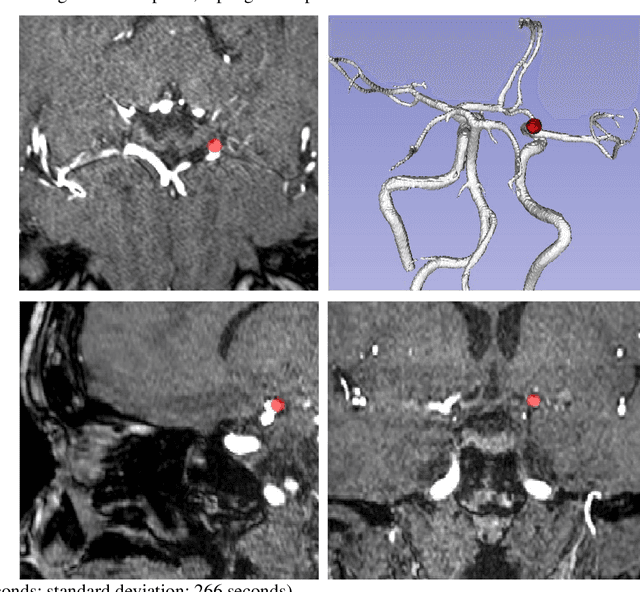
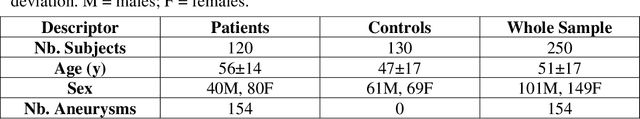
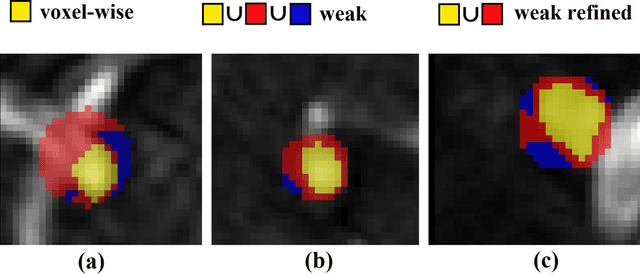
Abstract:Supervised segmentation algorithms yield state-of-the-art results for automated anomaly detection. However, these models require voxel-wise labels which are time-consuming to draw for medical experts. An interesting alternative to voxel-wise annotations is the use of weak labels: these can be coarse or oversized annotations that are less precise, but considerably faster to create. In this work, we address the task of brain aneurysm detection by developing a fully automated, deep neural network that is trained utilizing oversized weak labels. Furthermore, since aneurysms mainly occur in specific anatomical locations, we build our model leveraging the underlying anatomy of the brain vasculature both during training and inference. We apply our model to 250 subjects (120 patients, 130 controls) who underwent Time-Of-Flight Magnetic Resonance Angiography (TOF-MRA) and presented a total of 154 aneurysms. To assess the robustness of the algorithm, we participated in a MICCAI challenge for TOF-MRA data (93 patients, 20 controls, 125 aneurysms) which allowed us to obtain results also for subjects coming from a different institution. Our network achieves an average sensitivity of 77% on our in-house data, with a mean False Positive (FP) rate of 0.72 per patient. Instead, on the challenge data, we attain a sensitivity of 59% with a mean FP rate of 1.18, ranking in 7th/14 position for detection and in 4th/11 for segmentation on the open leaderboard. When computing detection performances with respect to aneurysms' risk of rupture, we found no statistical difference between two risk groups (p = 0.12), although the sensitivity for dangerous aneurysms was higher (78%). Our approach suggests that clinically useful sensitivity can be achieved using weak labels and exploiting prior anatomical knowledge; this expands the feasibility of deep learning studies to hospitals that have limited time and data.
An anatomically-informed 3D CNN for brain aneurysm classification with weak labels
Nov 27, 2020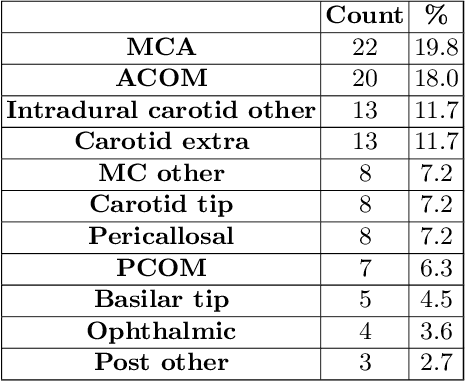

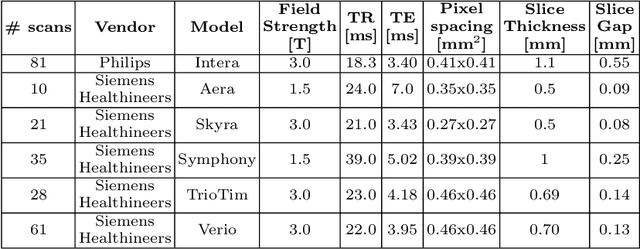
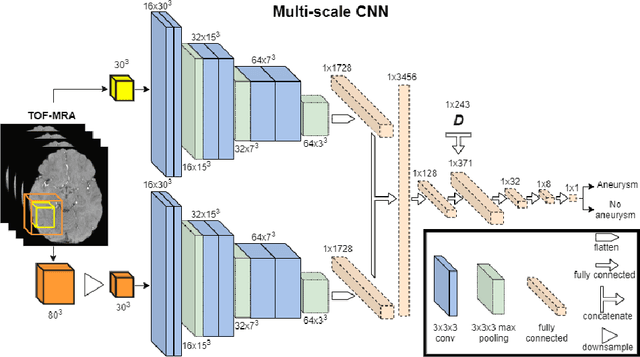
Abstract:A commonly adopted approach to carry out detection tasks in medical imaging is to rely on an initial segmentation. However, this approach strongly depends on voxel-wise annotations which are repetitive and time-consuming to draw for medical experts. An interesting alternative to voxel-wise masks are so-called "weak" labels: these can either be coarse or oversized annotations that are less precise, but noticeably faster to create. In this work, we address the task of brain aneurysm detection as a patch-wise binary classification with weak labels, in contrast to related studies that rather use supervised segmentation methods and voxel-wise delineations. Our approach comes with the non-trivial challenge of the data set creation: as for most focal diseases, anomalous patches (with aneurysm) are outnumbered by those showing no anomaly, and the two classes usually have different spatial distributions. To tackle this frequent scenario of inherently imbalanced, spatially skewed data sets, we propose a novel, anatomically-driven approach by using a multi-scale and multi-input 3D Convolutional Neural Network (CNN). We apply our model to 214 subjects (83 patients, 131 controls) who underwent Time-Of-Flight Magnetic Resonance Angiography (TOF-MRA) and presented a total of 111 unruptured cerebral aneurysms. We compare two strategies for negative patch sampling that have an increasing level of difficulty for the network and we show how this choice can strongly affect the results. To assess whether the added spatial information helps improving performances, we compare our anatomically-informed CNN with a baseline, spatially-agnostic CNN. When considering the more realistic and challenging scenario including vessel-like negative patches, the former model attains the highest classification results (accuracy$\simeq$95\%, AUROC$\simeq$0.95, AUPR$\simeq$0.71), thus outperforming the baseline.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge