Gabriel Girard
Signal Processing Laboratory 5, Department of Computer Science, Université de Sherbrooke, Sherbrooke, Canada
Clinical-ComBAT: a diffusion-weighted MRI harmonization method for clinical applications
Nov 06, 2025Abstract:Diffusion-weighted magnetic resonance imaging (DW-MRI) derived scalar maps are effective for assessing neurodegenerative diseases and microstructural properties of white matter in large number of brain conditions. However, DW-MRI inherently limits the combination of data from multiple acquisition sites without harmonization to mitigate scanner-specific biases. While the widely used ComBAT method reduces site effects in research, its reliance on linear covariate relationships, homogeneous populations, fixed site numbers, and well populated sites constrains its clinical use. To overcome these limitations, we propose Clinical-ComBAT, a method designed for real-world clinical scenarios. Clinical-ComBAT harmonizes each site independently, enabling flexibility as new data and clinics are introduced. It incorporates a non-linear polynomial data model, site-specific harmonization referenced to a normative site, and variance priors adaptable to small cohorts. It further includes hyperparameter tuning and a goodness-of-fit metric for harmonization assessment. We demonstrate its effectiveness on simulated and real data, showing improved alignment of diffusion metrics and enhanced applicability for normative modeling.
ComBAT Harmonization for diffusion MRI: Challenges and Best Practices
May 19, 2025Abstract:Over the years, ComBAT has become the standard method for harmonizing MRI-derived measurements, with its ability to compensate for site-related additive and multiplicative biases while preserving biological variability. However, ComBAT relies on a set of assumptions that, when violated, can result in flawed harmonization. In this paper, we thoroughly review ComBAT's mathematical foundation, outlining these assumptions, and exploring their implications for the demographic composition necessary for optimal results. Through a series of experiments involving a slightly modified version of ComBAT called Pairwise-ComBAT tailored for normative modeling applications, we assess the impact of various population characteristics, including population size, age distribution, the absence of certain covariates, and the magnitude of additive and multiplicative factors. Based on these experiments, we present five essential recommendations that should be carefully considered to enhance consistency and supporting reproducibility, two essential factors for open science, collaborative research, and real-life clinical deployment.
Spatially Regularized Super-Resolved Constrained Spherical Deconvolution (SR$^2$-CSD) of Diffusion MRI Data
Aug 23, 2024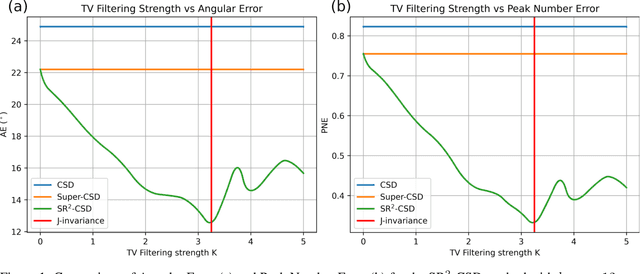
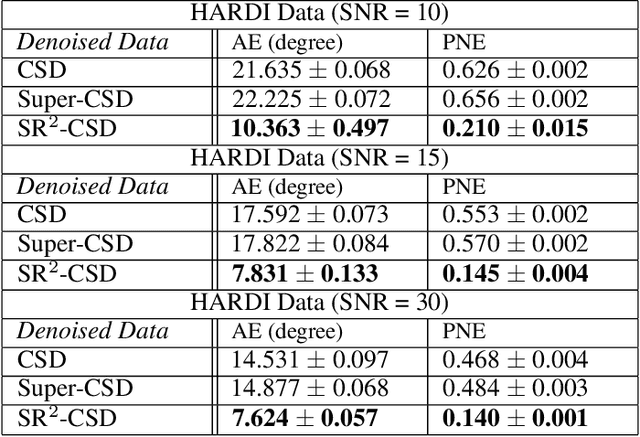
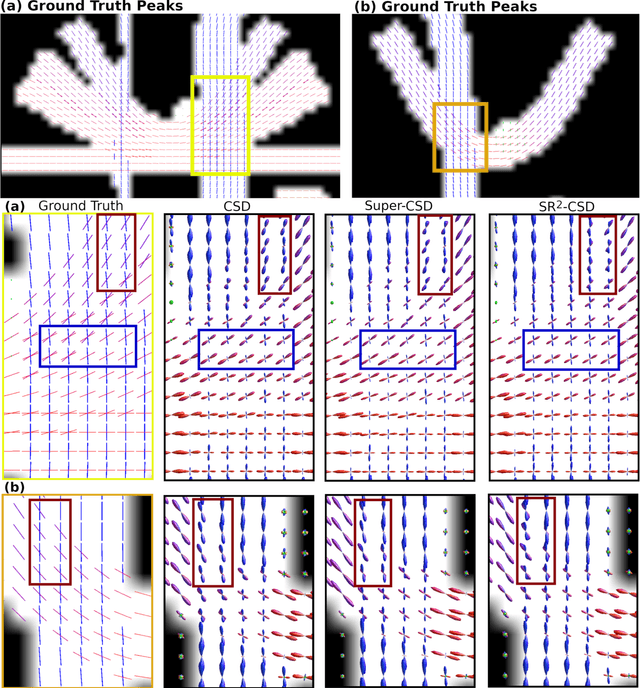
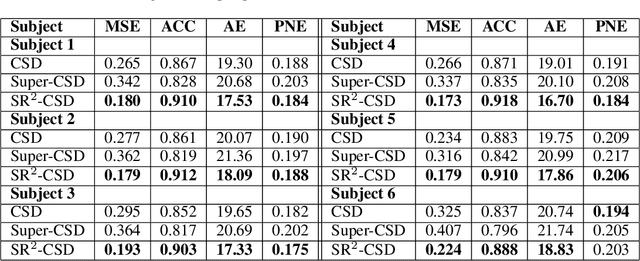
Abstract:Constrained Spherical Deconvolution (CSD) is crucial for estimating white matter fiber orientations using diffusion MRI data. A relevant parameter in CSD is the maximum order $l_{max}$ used in the spherical harmonics series, influencing the angular resolution of the Fiber Orientation Distributions (FODs). Lower $l_{max}$ values produce smoother and more stable estimates, but result in reduced angular resolution. Conversely, higher $l_{max}$ values, as employed in the Super-Resolved CSD variant, are essential for resolving narrow inter-fiber angles but lead to spurious lobes due to increased noise sensitivity. To address this issue, we propose a novel Spatially Regularized Super-Resolved CSD (SR$^2$-CSD) approach, incorporating spatial priors into the CSD framework. This method leverages spatial information among adjacent voxels, enhancing the stability and noise robustness of FOD estimations. SR$^2$-CSD facilitates the practical use of Super-Resolved CSD by including a J-invariant auto-calibrated total variation FOD denoiser. We evaluated the performance of SR$^2$-CSD against standard CSD and Super-Resolved CSD using phantom numerical data and various real brain datasets, including a test-retest sample of six subjects scanned twice. In phantom data, SR$^2$-CSD outperformed both CSD and Super-Resolved CSD, reducing the angular error (AE) by approximately half and the peak number error (PNE) by a factor of three across all noise levels considered. In real data, SR$^2$-CSD produced more continuous FOD estimates with higher spatial-angular coherency. In the test-retest sample, SR$^2$-CSD consistently yielded more reproducible estimates, with reduced AE, PNE, mean squared error, and increased angular correlation coefficient between the FODs estimated from the two scans for each subject.
Cellular EXchange Imaging (CEXI): Evaluation of a diffusion model including water exchange in cells using numerical phantoms of permeable spheres
Apr 12, 2023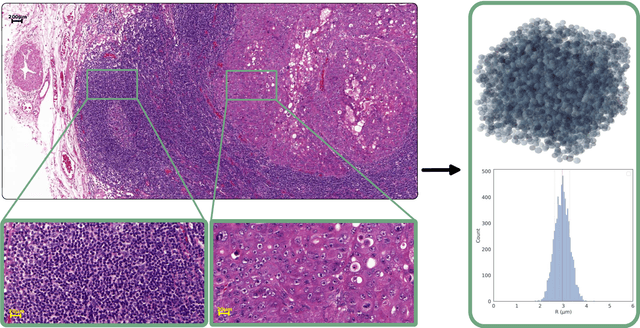

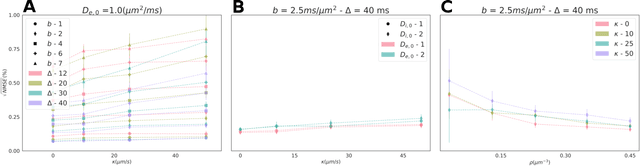

Abstract:Purpose: Biophysical models of diffusion MRI have been developed to characterize microstructure in various tissues, but existing models are not suitable for tissue composed of permeable spherical cells. In this study we introduce Cellular Exchange Imaging (CEXI), a model tailored for permeable spherical cells, and compares its performance to a related Ball \& Sphere (BS) model that neglects permeability. Methods: We generated DW-MRI signals using Monte-Carlo simulations with a PGSE sequence in numerical substrates made of spherical cells and their extracellular space for a range of membrane permeability. From these signals, the properties of the substrates were inferred using both BS and CEXI models. Results: CEXI outperformed the impermeable model by providing more stable estimates cell size and intracellular volume fraction that were diffusion time-independent. Notably, CEXI accurately estimated the exchange time for low to moderate permeability levels previously reported in other studies ($\kappa<25\mu m/s$). However, in highly permeable substrates ($\kappa=50\mu m/s$), the estimated parameters were less stable, particularly the diffusion coefficients. Conclusion: This study highlights the importance of modeling the exchange time to accurately quantify microstructure properties in permeable cellular substrates. Future studies should evaluate CEXI in clinical applications such as lymph nodes, investigate exchange time as a potential biomarker of tumor severity, and develop more appropriate tissue models that account for anisotropic diffusion and highly permeable membranes.
Why is the winner the best?
Mar 30, 2023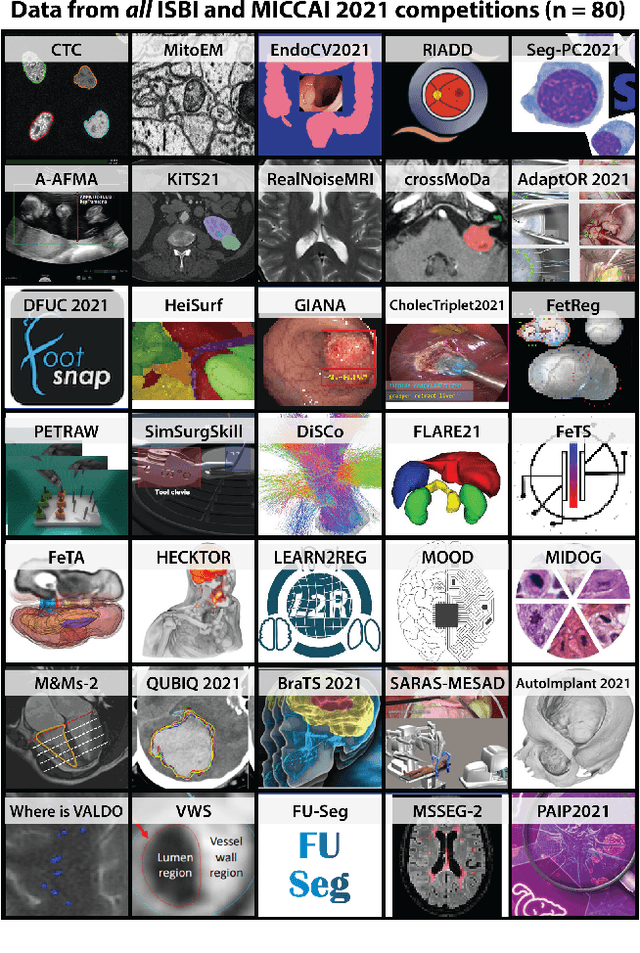
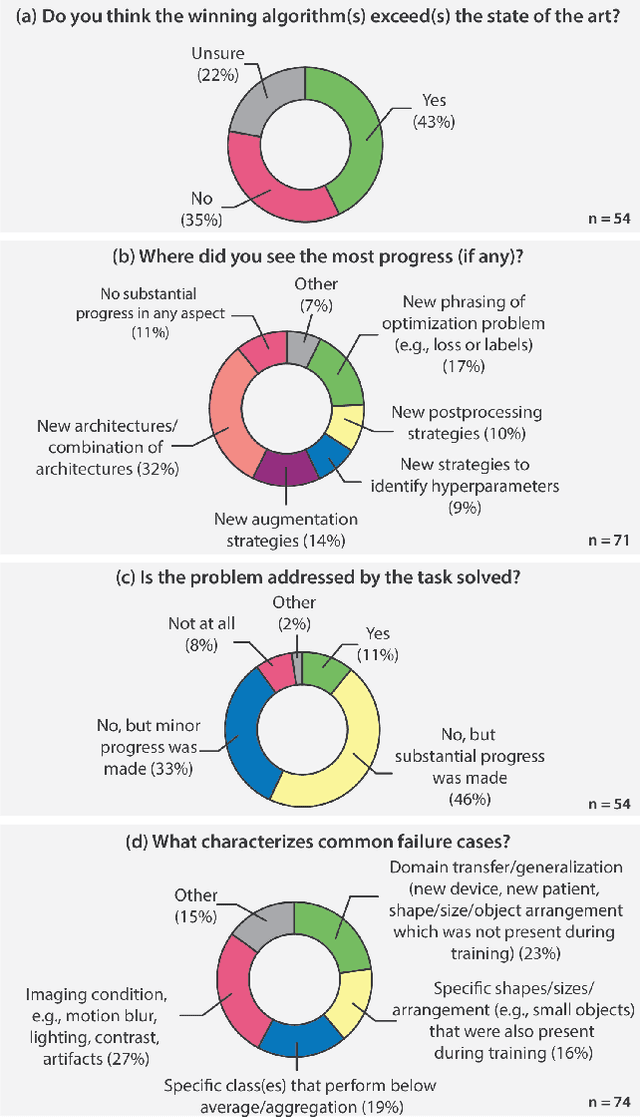
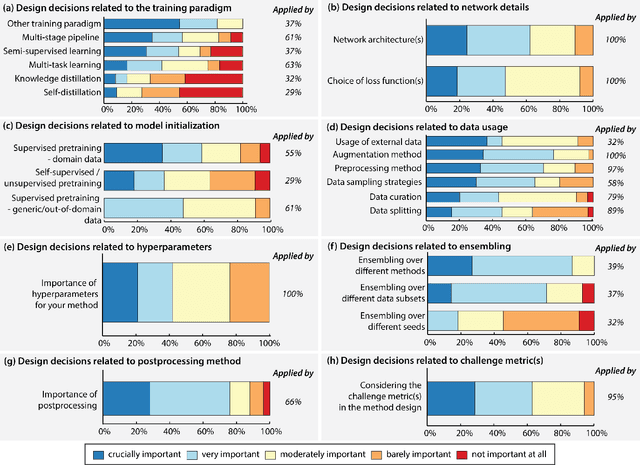
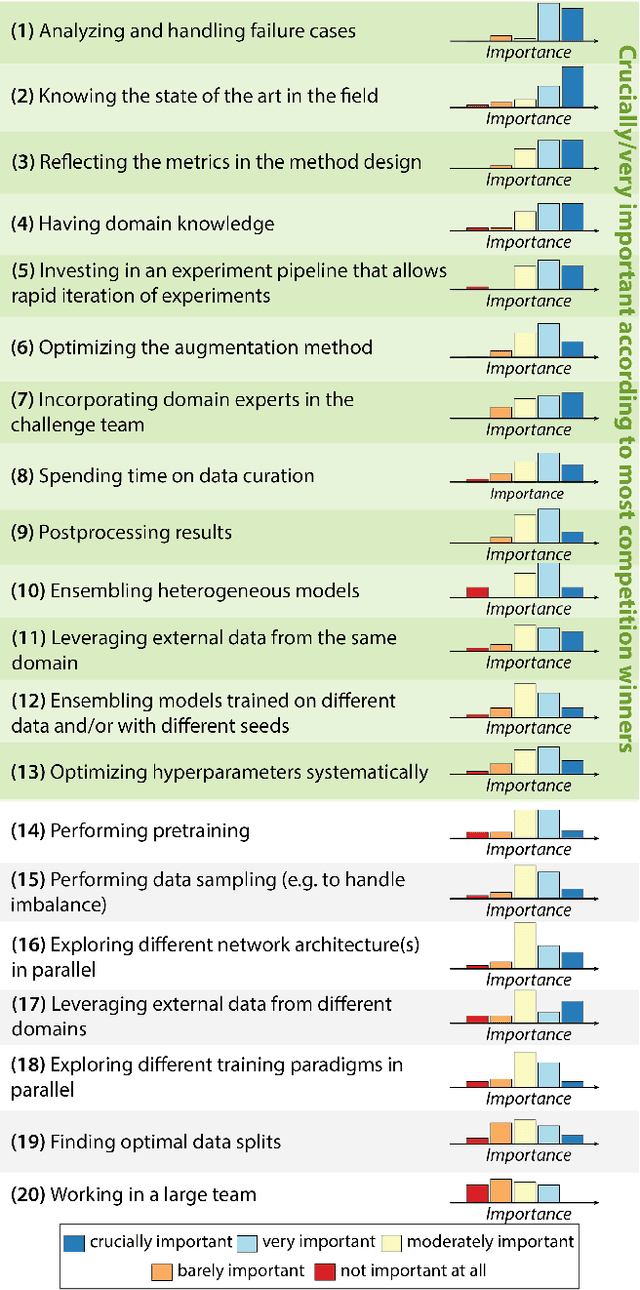
Abstract:International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Microstructure estimation from diffusion-MRI: Compartmentalized models in permeable cellular tissue
Sep 06, 2022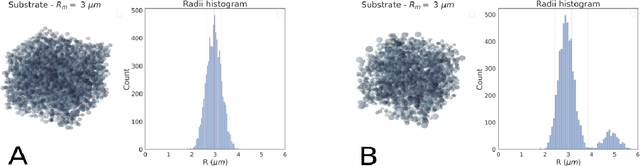

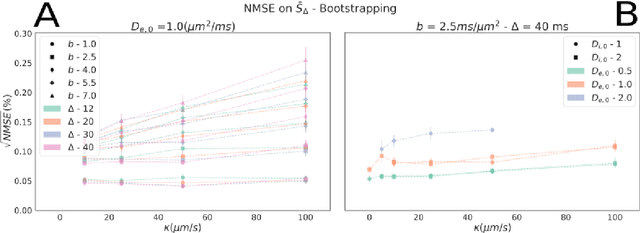

Abstract:Diffusion-weighted magnetic resonance imaging (DW-MRI) is used to characterize brain tissue microstructure employing tissue-specific biophysical models. A current limitation, however, is that most of the proposed models are based on the assumption of negligible water exchange between the intra- and extracellular compartments, which might not be valid in various brain tissues, including unmyelinated axons, gray matter, and tumors. The purpose of this work is to quantify the effect of membrane permeability on the estimates of two popular models neglecting exchange, and compare their performance with a model including exchange. To this aim, DW-MRI experiments were performed in controlled environments with Monte-Carlo simulations. The DW-MRI signals were generated in numerical substrates mimicking biological tissue made of spherical cells with permeable membranes like cancerous tissue or the brain gray matter. From these signals, the substrates properties were estimated using SANDI and VERDICT, the two compartment-based models neglecting exchange, and CEXI, a new model which includes exchange. Our results show that, in cellular permeable tissue, the model with exchange outperformed models without exchange in the estimation of the tissue properties by providing more stable estimates of cell size, intracellular volume fraction and extracellular diffusion coefficient. Moreover, the model with exchange estimated accurately the exchange time in the range of permeability reported for cellular tissue. Finally, the simulations performed in this work showed that the exchange between the intracellular and the extracellular space cannot be neglected in permeable tissue with a conventional PGSE sequence, to obtain accurate estimates. Consequently, existing compartmentalized models of impermeable tissue cannot be used for microstructure estimation of cellular permeable tissue.
Slice estimation in diffusion MRI of neonatal and fetal brains in image and spherical harmonics domains using autoencoders
Aug 29, 2022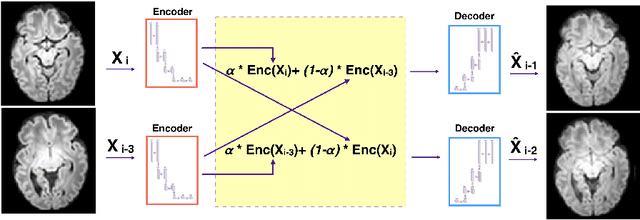
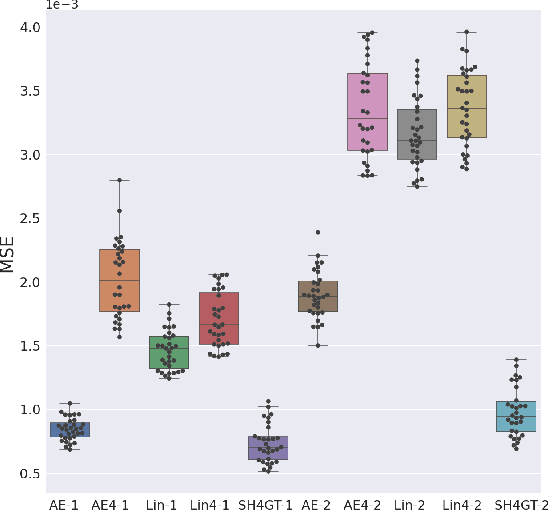
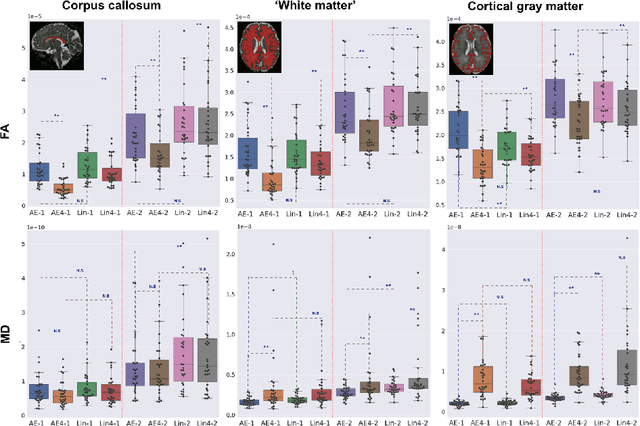
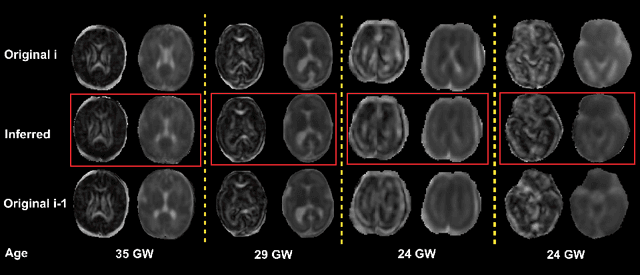
Abstract:Diffusion MRI (dMRI) of the developing brain can provide valuable insights into the white matter development. However, slice thickness in fetal dMRI is typically high (i.e., 3-5 mm) to freeze the in-plane motion, which reduces the sensitivity of the dMRI signal to the underlying anatomy. In this study, we aim at overcoming this problem by using autoencoders to learn unsupervised efficient representations of brain slices in a latent space, using raw dMRI signals and their spherical harmonics (SH) representation. We first learn and quantitatively validate the autoencoders on the developing Human Connectome Project pre-term newborn data, and further test the method on fetal data. Our results show that the autoencoder in the signal domain better synthesized the raw signal. Interestingly, the fractional anisotropy and, to a lesser extent, the mean diffusivity, are best recovered in missing slices by using the autoencoder trained with SH coefficients. A comparison was performed with the same maps reconstructed using an autoencoder trained with raw signals, as well as conventional interpolation methods of raw signals and SH coefficients. From these results, we conclude that the recovery of missing/corrupted slices should be performed in the signal domain if the raw signal is aimed to be recovered, and in the SH domain if diffusion tensor properties (i.e., fractional anisotropy) are targeted. Notably, the trained autoencoders were able to generalize to fetal dMRI data acquired using a much smaller number of diffusion gradients and a lower b-value, where we qualitatively show the consistency of the estimated diffusion tensor maps.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge