Vladyslav Zalevskyi
Advances in Automated Fetal Brain MRI Segmentation and Biometry: Insights from the FeTA 2024 Challenge
May 05, 2025



Abstract:Accurate fetal brain tissue segmentation and biometric analysis are essential for studying brain development in utero. The FeTA Challenge 2024 advanced automated fetal brain MRI analysis by introducing biometry prediction as a new task alongside tissue segmentation. For the first time, our diverse multi-centric test set included data from a new low-field (0.55T) MRI dataset. Evaluation metrics were also expanded to include the topology-specific Euler characteristic difference (ED). Sixteen teams submitted segmentation methods, most of which performed consistently across both high- and low-field scans. However, longitudinal trends indicate that segmentation accuracy may be reaching a plateau, with results now approaching inter-rater variability. The ED metric uncovered topological differences that were missed by conventional metrics, while the low-field dataset achieved the highest segmentation scores, highlighting the potential of affordable imaging systems when paired with high-quality reconstruction. Seven teams participated in the biometry task, but most methods failed to outperform a simple baseline that predicted measurements based solely on gestational age, underscoring the challenge of extracting reliable biometric estimates from image data alone. Domain shift analysis identified image quality as the most significant factor affecting model generalization, with super-resolution pipelines also playing a substantial role. Other factors, such as gestational age, pathology, and acquisition site, had smaller, though still measurable, effects. Overall, FeTA 2024 offers a comprehensive benchmark for multi-class segmentation and biometry estimation in fetal brain MRI, underscoring the need for data-centric approaches, improved topological evaluation, and greater dataset diversity to enable clinically robust and generalizable AI tools.
Maximizing domain generalization in fetal brain tissue segmentation: the role of synthetic data generation, intensity clustering and real image fine-tuning
Nov 11, 2024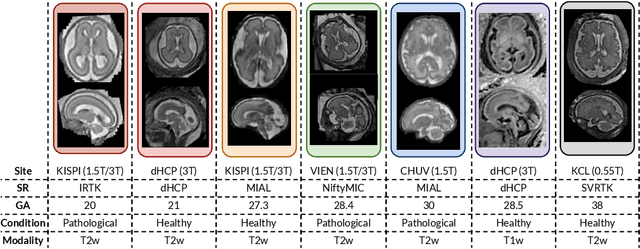
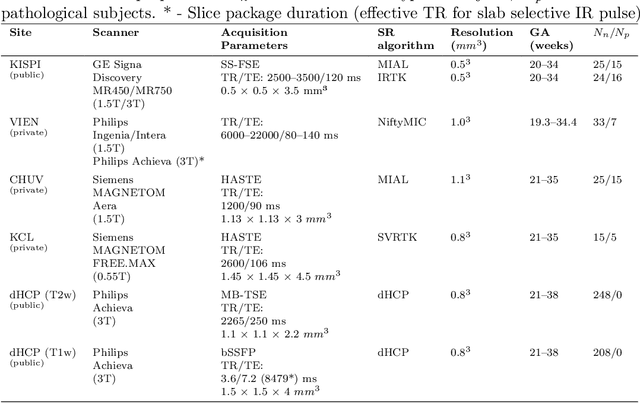
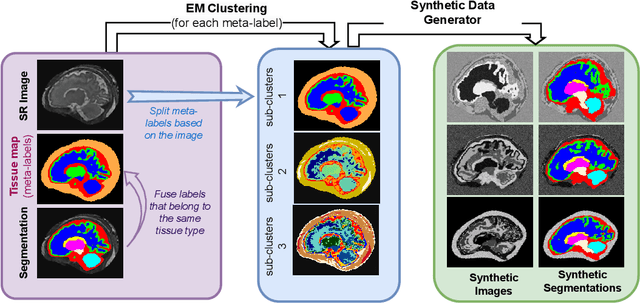
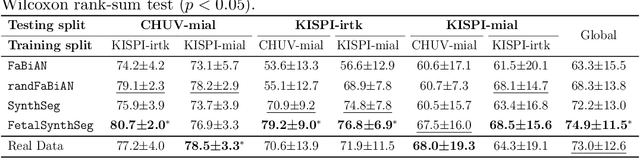
Abstract:Fetal brain tissue segmentation in magnetic resonance imaging (MRI) is a crucial tool that supports the understanding of neurodevelopment, yet it faces challenges due to the heterogeneity of data coming from different scanners and settings, and due to data scarcity. Recent approaches based on domain randomization, like SynthSeg, have shown a great potential for single source domain generalization, by simulating images with randomized contrast and image resolution from the label maps. In this work, we investigate how to maximize the out-of-domain (OOD) generalization potential of SynthSeg-based methods in fetal brain MRI. Specifically, when studying data generation, we demonstrate that the simple Gaussian mixture models used in SynthSeg enable more robust OOD generalization than physics-informed generation methods. We also investigate how intensity clustering can help create more faithful synthetic images, and observe that it is key to achieving a non-trivial OOD generalization capability when few label classes are available. Finally, by combining for the first time SynthSeg with modern fine-tuning approaches based on weight averaging, we show that fine-tuning a model pre-trained on synthetic data on a few real image-segmentation pairs in a new domain can lead to improvements in the target domain, but also in other domains. We summarize our findings as five key recommendations that we believe can guide practitioners who would like to develop SynthSeg-based approaches in other organs or modalities.
Improving cross-domain brain tissue segmentation in fetal MRI with synthetic data
Mar 22, 2024Abstract:Segmentation of fetal brain tissue from magnetic resonance imaging (MRI) plays a crucial role in the study of in utero neurodevelopment. However, automated tools face substantial domain shift challenges as they must be robust to highly heterogeneous clinical data, often limited in numbers and lacking annotations. Indeed, high variability of the fetal brain morphology, MRI acquisition parameters, and superresolution reconstruction (SR) algorithms adversely affect the model's performance when evaluated out-of-domain. In this work, we introduce FetalSynthSeg, a domain randomization method to segment fetal brain MRI, inspired by SynthSeg. Our results show that models trained solely on synthetic data outperform models trained on real data in out-ofdomain settings, validated on a 120-subject cross-domain dataset. Furthermore, we extend our evaluation to 40 subjects acquired using lowfield (0.55T) MRI and reconstructed with novel SR models, showcasing robustness across different magnetic field strengths and SR algorithms. Leveraging a generative synthetic approach, we tackle the domain shift problem in fetal brain MRI and offer compelling prospects for applications in fields with limited and highly heterogeneous data.
SYNCS: Synthetic Data and Contrastive Self-Supervised Training for Central Sulcus Segmentation
Mar 22, 2024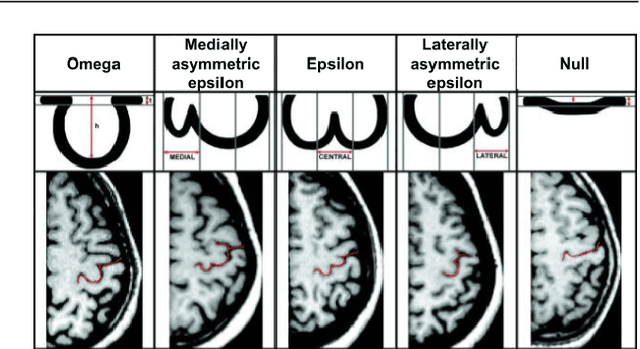
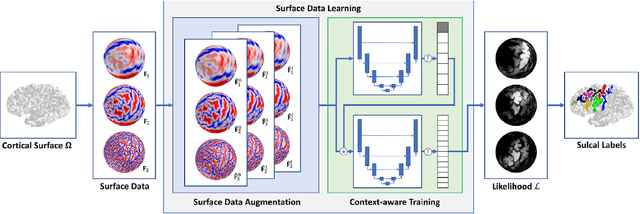
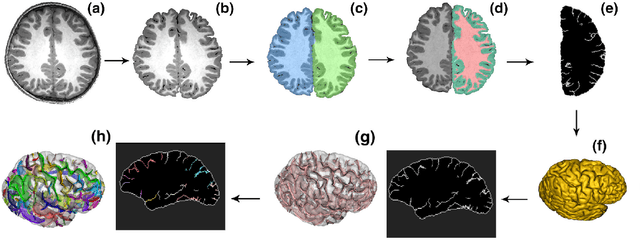
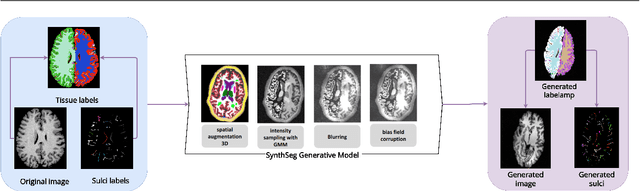
Abstract:Bipolar disorder (BD) and schizophrenia (SZ) are severe mental disorders with profound societal impact. Identifying risk markers early is crucial for understanding disease progression and enabling preventive measures. The Danish High Risk and Resilience Study (VIA) focuses on understanding early disease processes, particularly in children with familial high risk (FHR). Understanding structural brain changes associated with these diseases during early stages is essential for effective interventions. The central sulcus (CS) is a prominent brain landmark related to brain regions involved in motor and sensory processing. Analyzing CS morphology can provide valuable insights into neurodevelopmental abnormalities in the FHR group. However, segmenting the central sulcus (CS) presents challenges due to its variability, especially in adolescents. This study introduces two novel approaches to improve CS segmentation: synthetic data generation to model CS variability and self-supervised pre-training with multi-task learning to adapt models to new cohorts. These methods aim to enhance segmentation performance across diverse populations, eliminating the need for extensive preprocessing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge