Synthesis of realistic fetal MRI with conditional Generative Adversarial Networks
Paper and Code
Sep 20, 2022
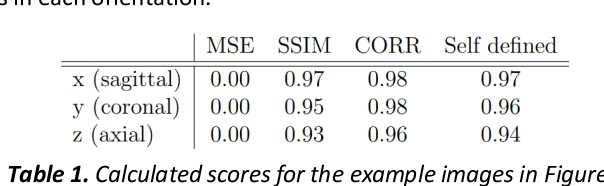
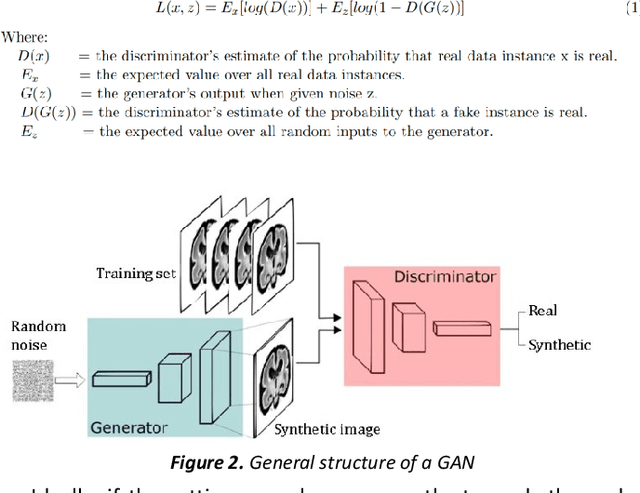
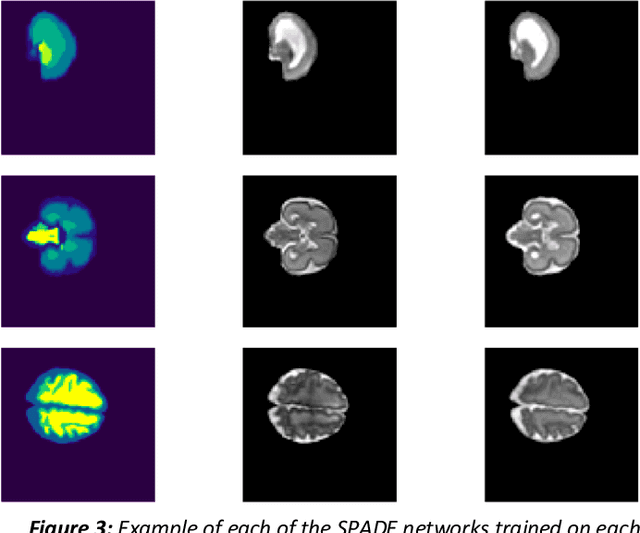
Fetal brain magnetic resonance imaging serves as an emerging modality for prenatal counseling and diagnosis in disorders affecting the brain. Machine learning based segmentation plays an important role in the quantification of brain development. However, a limiting factor is the lack of sufficiently large, labeled training data. Our study explored the application of SPADE, a conditional general adversarial network (cGAN), which learns the mapping from the label to the image space. The input to the network was super-resolution T2-weighted cerebral MRI data of 120 fetuses (gestational age range: 20-35 weeks, normal and pathological), which were annotated for 7 different tissue categories. SPADE networks were trained on 256*256 2D slices of the reconstructed volumes (image and label pairs) in each orthogonal orientation. To combine the generated volumes from each orientation into one image, a simple mean of the outputs of the three networks was taken. Based on the label maps only, we synthesized highly realistic images. However, some finer details, like small vessels were not synthesized. A structural similarity index (SSIM) of 0.972+-0.016 and correlation coefficient of 0.974+-0.008 were achieved. To demonstrate the capacity of the cGAN to create new anatomical variants, we artificially dilated the ventricles in the segmentation map and created synthetic MRI of different degrees of fetal hydrocephalus. cGANs, such as the SPADE algorithm, allow the generation of hypothetically unseen scenarios and anatomical configurations in the label space, which data in turn can be utilized for training various machine learning algorithms. In the future, this algorithm would be used for generating large, synthetic datasets representing fetal brain development. These datasets would potentially improve the performance of currently available segmentation networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge