Jason Holmes
Diffusion Transformer-based Universal Dose Denoising for Pencil Beam Scanning Proton Therapy
Jun 04, 2025Abstract:Purpose: Intensity-modulated proton therapy (IMPT) offers precise tumor coverage while sparing organs at risk (OARs) in head and neck (H&N) cancer. However, its sensitivity to anatomical changes requires frequent adaptation through online adaptive radiation therapy (oART), which depends on fast, accurate dose calculation via Monte Carlo (MC) simulations. Reducing particle count accelerates MC but degrades accuracy. To address this, denoising low-statistics MC dose maps is proposed to enable fast, high-quality dose generation. Methods: We developed a diffusion transformer-based denoising framework. IMPT plans and 3D CT images from 80 H&N patients were used to generate noisy and high-statistics dose maps using MCsquare (1 min and 10 min per plan, respectively). Data were standardized into uniform chunks with zero-padding, normalized, and transformed into quasi-Gaussian distributions. Testing was done on 10 H&N, 10 lung, 10 breast, and 10 prostate cancer cases, preprocessed identically. The model was trained with noisy dose maps and CT images as input and high-statistics dose maps as ground truth, using a combined loss of mean square error (MSE), residual loss, and regional MAE (focusing on top/bottom 10% dose voxels). Performance was assessed via MAE, 3D Gamma passing rate, and DVH indices. Results: The model achieved MAEs of 0.195 (H&N), 0.120 (lung), 0.172 (breast), and 0.376 Gy[RBE] (prostate). 3D Gamma passing rates exceeded 92% (3%/2mm) across all sites. DVH indices for clinical target volumes (CTVs) and OARs closely matched the ground truth. Conclusion: A diffusion transformer-based denoising framework was developed and, though trained only on H&N data, generalizes well across multiple disease sites.
Fine-Tuning Open-Source Large Language Models to Improve Their Performance on Radiation Oncology Tasks: A Feasibility Study to Investigate Their Potential Clinical Applications in Radiation Oncology
Jan 28, 2025



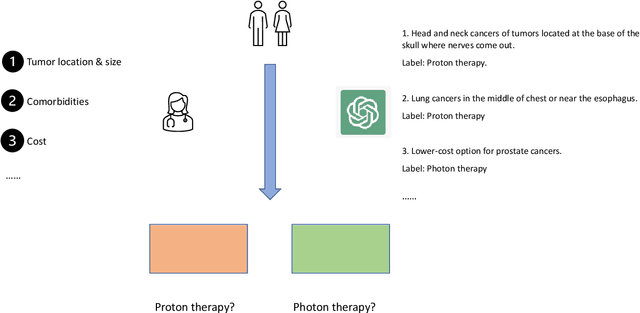
Abstract:Background: The radiation oncology clinical practice involves many steps relying on the dynamic interplay of abundant text data. Large language models have displayed remarkable capabilities in processing complex text information. But their direct applications in specific fields like radiation oncology remain underexplored. Purpose: This study aims to investigate whether fine-tuning LLMs with domain knowledge can improve the performance on Task (1) treatment regimen generation, Task (2) treatment modality selection (photon, proton, electron, or brachytherapy), and Task (3) ICD-10 code prediction in radiation oncology. Methods: Data for 15,724 patient cases were extracted. Cases where patients had a single diagnostic record, and a clearly identifiable primary treatment plan were selected for preprocessing and manual annotation to have 7,903 cases of the patient diagnosis, treatment plan, treatment modality, and ICD-10 code. Each case was used to construct a pair consisting of patient diagnostics details and an answer (treatment regimen, treatment modality, or ICD-10 code respectively) for the supervised fine-tuning of these three tasks. Open source LLaMA2-7B and Mistral-7B models were utilized for the fine-tuning with the Low-Rank Approximations method. Accuracy and ROUGE-1 score were reported for the fine-tuned models and original models. Clinical evaluation was performed on Task (1) by radiation oncologists, while precision, recall, and F-1 score were evaluated for Task (2) and (3). One-sided Wilcoxon signed-rank tests were used to statistically analyze the results. Results: Fine-tuned LLMs outperformed original LLMs across all tasks with p-value <= 0.001. Clinical evaluation demonstrated that over 60% of the fine-tuned LLMs-generated treatment regimens were clinically acceptable. Precision, recall, and F1-score showed improved performance of fine-tuned LLMs.
Evaluating The Performance of Using Large Language Models to Automate Summarization of CT Simulation Orders in Radiation Oncology
Jan 27, 2025



Abstract:Purpose: This study aims to use a large language model (LLM) to automate the generation of summaries from the CT simulation orders and evaluate its performance. Materials and Methods: A total of 607 CT simulation orders for patients were collected from the Aria database at our institution. A locally hosted Llama 3.1 405B model, accessed via the Application Programming Interface (API) service, was used to extract keywords from the CT simulation orders and generate summaries. The downloaded CT simulation orders were categorized into seven groups based on treatment modalities and disease sites. For each group, a customized instruction prompt was developed collaboratively with therapists to guide the Llama 3.1 405B model in generating summaries. The ground truth for the corresponding summaries was manually derived by carefully reviewing each CT simulation order and subsequently verified by therapists. The accuracy of the LLM-generated summaries was evaluated by therapists using the verified ground truth as a reference. Results: About 98% of the LLM-generated summaries aligned with the manually generated ground truth in terms of accuracy. Our evaluations showed an improved consistency in format and enhanced readability of the LLM-generated summaries compared to the corresponding therapists-generated summaries. This automated approach demonstrated a consistent performance across all groups, regardless of modality or disease site. Conclusions: This study demonstrated the high precision and consistency of the Llama 3.1 405B model in extracting keywords and summarizing CT simulation orders, suggesting that LLMs have great potential to help with this task, reduce the workload of therapists and improve workflow efficiency.
A recent evaluation on the performance of LLMs on radiation oncology physics using questions of randomly shuffled options
Dec 14, 2024



Abstract:Purpose: We present an updated study evaluating the performance of large language models (LLMs) in answering radiation oncology physics questions, focusing on the latest released models. Methods: A set of 100 multiple-choice radiation oncology physics questions, previously created by us, was used for this study. The answer options of the questions were randomly shuffled to create "new" exam sets. Five LLMs -- OpenAI o1-preview, GPT-4o, LLaMA 3.1 (405B), Gemini 1.5 Pro, and Claude 3.5 Sonnet -- with the versions released before September 30, 2024, were queried using these new exams. To evaluate their deductive reasoning abilities, the correct answer options in the questions were replaced with "None of the above." Then, the explain-first and step-by-step instruction prompt was used to test if it improved their reasoning abilities. The performance of the LLMs was compared to medical physicists in majority-vote scenarios. Results: All models demonstrated expert-level performance on these questions, with o1-preview even surpassing medical physicists in majority-vote scenarios. When substituting the correct answer options with "None of the above," all models exhibited a considerable decline in performance, suggesting room for improvement. The explain-first and step-by-step instruction prompt helped enhance the reasoning abilities of LLaMA 3.1 (405B), Gemini 1.5 Pro, and Claude 3.5 Sonnet models. Conclusion: These latest LLMs demonstrated expert-level performance in answering radiation oncology physics questions, exhibiting great potential for assisting in radiation oncology physics education.
Evaluation of OpenAI o1: Opportunities and Challenges of AGI
Sep 27, 2024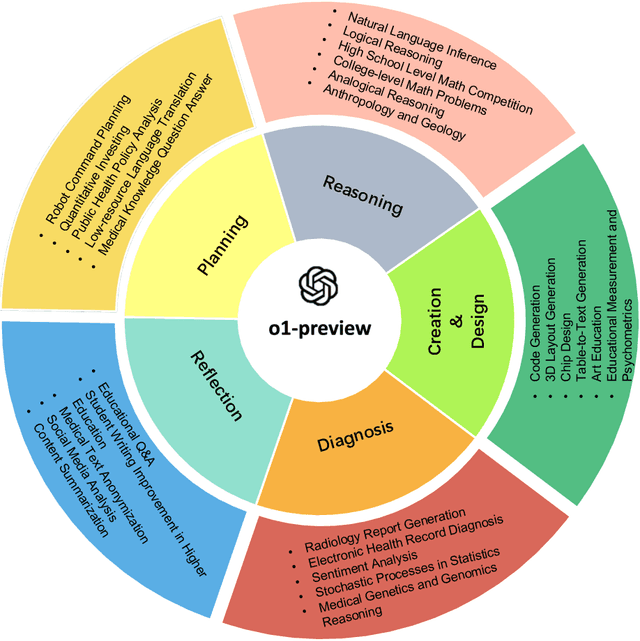

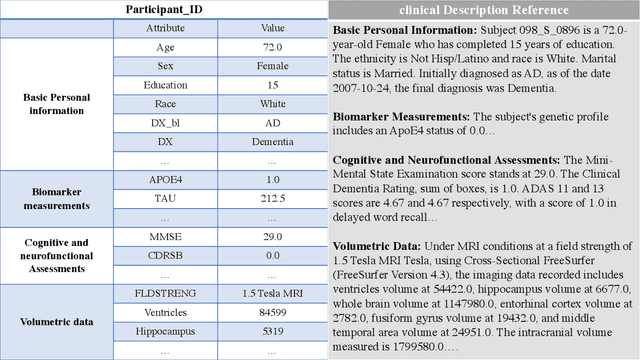
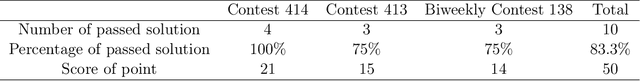
Abstract:This comprehensive study evaluates the performance of OpenAI's o1-preview large language model across a diverse array of complex reasoning tasks, spanning multiple domains, including computer science, mathematics, natural sciences, medicine, linguistics, and social sciences. Through rigorous testing, o1-preview demonstrated remarkable capabilities, often achieving human-level or superior performance in areas ranging from coding challenges to scientific reasoning and from language processing to creative problem-solving. Key findings include: -83.3% success rate in solving complex competitive programming problems, surpassing many human experts. -Superior ability in generating coherent and accurate radiology reports, outperforming other evaluated models. -100% accuracy in high school-level mathematical reasoning tasks, providing detailed step-by-step solutions. -Advanced natural language inference capabilities across general and specialized domains like medicine. -Impressive performance in chip design tasks, outperforming specialized models in areas such as EDA script generation and bug analysis. -Remarkable proficiency in anthropology and geology, demonstrating deep understanding and reasoning in these specialized fields. -Strong capabilities in quantitative investing. O1 has comprehensive financial knowledge and statistical modeling skills. -Effective performance in social media analysis, including sentiment analysis and emotion recognition. The model excelled particularly in tasks requiring intricate reasoning and knowledge integration across various fields. While some limitations were observed, including occasional errors on simpler problems and challenges with certain highly specialized concepts, the overall results indicate significant progress towards artificial general intelligence.
The Radiation Oncology NLP Database
Jan 19, 2024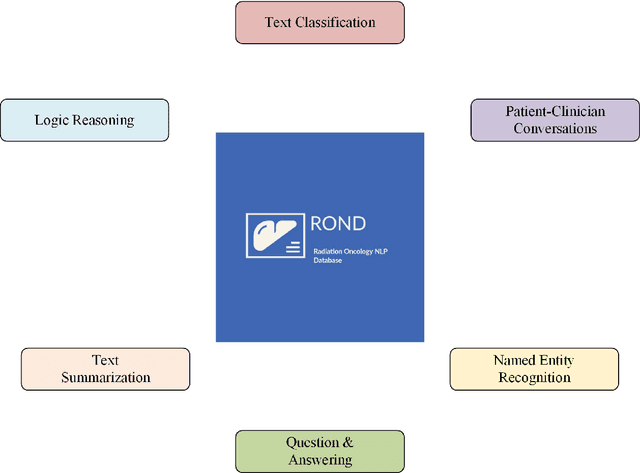


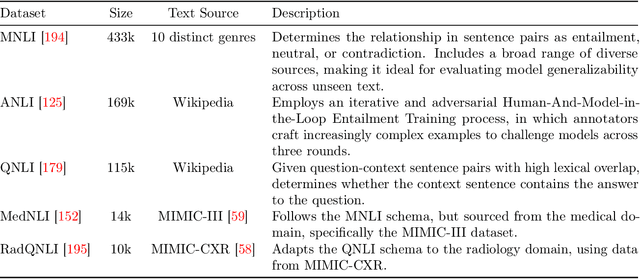
Abstract:We present the Radiation Oncology NLP Database (ROND), the first dedicated Natural Language Processing (NLP) dataset for radiation oncology, an important medical specialty that has received limited attention from the NLP community in the past. With the advent of Artificial General Intelligence (AGI), there is an increasing need for specialized datasets and benchmarks to facilitate research and development. ROND is specifically designed to address this gap in the domain of radiation oncology, a field that offers many opportunities for NLP exploration. It encompasses various NLP tasks including Logic Reasoning, Text Classification, Named Entity Recognition (NER), Question Answering (QA), Text Summarization, and Patient-Clinician Conversations, each with a distinct focus on radiation oncology concepts and application cases. In addition, we have developed an instruction-tuning dataset consisting of over 20k instruction pairs (based on ROND) and trained a large language model, CancerChat. This serves to demonstrate the potential of instruction-tuning large language models within a highly-specialized medical domain. The evaluation results in this study could serve as baseline results for future research. ROND aims to stimulate advancements in radiation oncology and clinical NLP by offering a platform for testing and improving algorithms and models in a domain-specific context. The ROND dataset is a joint effort of multiple U.S. health institutions. The data is available at https://github.com/zl-liu/Radiation-Oncology-NLP-Database.
Evaluating Large Language Models in Ophthalmology
Nov 07, 2023Abstract:Purpose: The performance of three different large language models (LLMS) (GPT-3.5, GPT-4, and PaLM2) in answering ophthalmology professional questions was evaluated and compared with that of three different professional populations (medical undergraduates, medical masters, and attending physicians). Methods: A 100-item ophthalmology single-choice test was administered to three different LLMs (GPT-3.5, GPT-4, and PaLM2) and three different professional levels (medical undergraduates, medical masters, and attending physicians), respectively. The performance of LLM was comprehensively evaluated and compared with the human group in terms of average score, stability, and confidence. Results: Each LLM outperformed undergraduates in general, with GPT-3.5 and PaLM2 being slightly below the master's level, while GPT-4 showed a level comparable to that of attending physicians. In addition, GPT-4 showed significantly higher answer stability and confidence than GPT-3.5 and PaLM2. Conclusion: Our study shows that LLM represented by GPT-4 performs better in the field of ophthalmology. With further improvements, LLM will bring unexpected benefits in medical education and clinical decision making in the near future.
Evaluating multiple large language models in pediatric ophthalmology
Nov 07, 2023



Abstract:IMPORTANCE The response effectiveness of different large language models (LLMs) and various individuals, including medical students, graduate students, and practicing physicians, in pediatric ophthalmology consultations, has not been clearly established yet. OBJECTIVE Design a 100-question exam based on pediatric ophthalmology to evaluate the performance of LLMs in highly specialized scenarios and compare them with the performance of medical students and physicians at different levels. DESIGN, SETTING, AND PARTICIPANTS This survey study assessed three LLMs, namely ChatGPT (GPT-3.5), GPT-4, and PaLM2, were assessed alongside three human cohorts: medical students, postgraduate students, and attending physicians, in their ability to answer questions related to pediatric ophthalmology. It was conducted by administering questionnaires in the form of test papers through the LLM network interface, with the valuable participation of volunteers. MAIN OUTCOMES AND MEASURES Mean scores of LLM and humans on 100 multiple-choice questions, as well as the answer stability, correlation, and response confidence of each LLM. RESULTS GPT-4 performed comparably to attending physicians, while ChatGPT (GPT-3.5) and PaLM2 outperformed medical students but slightly trailed behind postgraduate students. Furthermore, GPT-4 exhibited greater stability and confidence when responding to inquiries compared to ChatGPT (GPT-3.5) and PaLM2. CONCLUSIONS AND RELEVANCE Our results underscore the potential for LLMs to provide medical assistance in pediatric ophthalmology and suggest significant capacity to guide the education of medical students.
Evaluating the Potential of Leading Large Language Models in Reasoning Biology Questions
Nov 05, 2023Abstract:Recent advances in Large Language Models (LLMs) have presented new opportunities for integrating Artificial General Intelligence (AGI) into biological research and education. This study evaluated the capabilities of leading LLMs, including GPT-4, GPT-3.5, PaLM2, Claude2, and SenseNova, in answering conceptual biology questions. The models were tested on a 108-question multiple-choice exam covering biology topics in molecular biology, biological techniques, metabolic engineering, and synthetic biology. Among the models, GPT-4 achieved the highest average score of 90 and demonstrated the greatest consistency across trials with different prompts. The results indicated GPT-4's proficiency in logical reasoning and its potential to aid biology research through capabilities like data analysis, hypothesis generation, and knowledge integration. However, further development and validation are still required before the promise of LLMs in accelerating biological discovery can be realized.
Benchmarking a foundation LLM on its ability to re-label structure names in accordance with the AAPM TG-263 report
Oct 05, 2023



Abstract:Purpose: To introduce the concept of using large language models (LLMs) to re-label structure names in accordance with the American Association of Physicists in Medicine (AAPM) Task Group (TG)-263 standard, and to establish a benchmark for future studies to reference. Methods and Materials: The Generative Pre-trained Transformer (GPT)-4 application programming interface (API) was implemented as a Digital Imaging and Communications in Medicine (DICOM) storage server, which upon receiving a structure set DICOM file, prompts GPT-4 to re-label the structure names of both target volumes and normal tissues according to the AAPM TG-263. Three disease sites, prostate, head and neck, and thorax were selected for evaluation. For each disease site category, 150 patients were randomly selected for manually tuning the instructions prompt (in batches of 50) and 50 patients were randomly selected for evaluation. Structure names that were considered were those that were most likely to be relevant for studies utilizing structure contours for many patients. Results: The overall re-labeling accuracy of both target volumes and normal tissues for prostate, head and neck, and thorax cases was 96.0%, 98.5%, and 96.9% respectively. Re-labeling of target volumes was less accurate on average except for prostate - 100%, 93.1%, and 91.1% respectively. Conclusions: Given the accuracy of GPT-4 in re-labeling structure names of both target volumes and normal tissues as presented in this work, LLMs are poised to be the preferred method for standardizing structure names in radiation oncology, especially considering the rapid advancements in LLM capabilities that are likely to continue.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge