Hyunjae Kim
A Federated and Parameter-Efficient Framework for Large Language Model Training in Medicine
Jan 29, 2026Abstract:Large language models (LLMs) have demonstrated strong performance on medical benchmarks, including question answering and diagnosis. To enable their use in clinical settings, LLMs are typically further adapted through continued pretraining or post-training using clinical data. However, most medical LLMs are trained on data from a single institution, which faces limitations in generalizability and safety in heterogeneous systems. Federated learning (FL) is a promising solution for enabling collaborative model development across healthcare institutions. Yet applying FL to LLMs in medicine remains fundamentally limited. First, conventional FL requires transmitting the full model during each communication round, which becomes impractical for multi-billion-parameter LLMs given the limited computational resources. Second, many FL algorithms implicitly assume data homogeneity, whereas real-world clinical data are highly heterogeneous across patients, diseases, and institutional practices. We introduce the model-agnostic and parameter-efficient federated learning framework for adapting LLMs to medical applications. Fed-MedLoRA transmits only low-rank adapter parameters, reducing communication and computation overhead, while Fed-MedLoRA+ further incorporates adaptive, data-aware aggregation to improve convergence under cross-site heterogeneity. We apply the framework to clinical information extraction (IE), which transforms patient narratives into structured medical entities and relations. Accuracy was assessed across five patient cohorts through comparisons with BERT models, and LLaMA-3 and DeepSeek-R1, GPT-4o models. Evaluation settings included (1) in-domain training and testing, (2) external validation on independent cohorts, and (3) a low-resource new-site adaptation scenario using real-world clinical notes from the Yale New Haven Health System.
Benchmarking Direct Preference Optimization for Medical Large Vision-Language Models
Jan 25, 2026Abstract:Large Vision-Language Models (LVLMs) hold significant promise for medical applications, yet their deployment is often constrained by insufficient alignment and reliability. While Direct Preference Optimization (DPO) has emerged as a potent framework for refining model responses, its efficacy in high-stakes medical contexts remains underexplored, lacking the rigorous empirical groundwork necessary to guide future methodological advances. To bridge this gap, we present the first comprehensive examination of diverse DPO variants within the medical domain, evaluating nine distinct formulations across two medical LVLMs: LLaVA-Med and HuatuoGPT-Vision. Our results reveal several critical limitations: current DPO approaches often yield inconsistent gains over supervised fine-tuning, with their efficacy varying significantly across different tasks and backbones. Furthermore, they frequently fail to resolve fundamental visual misinterpretation errors. Building on these insights, we present a targeted preference construction strategy as a proof-of-concept that explicitly addresses visual misinterpretation errors frequently observed in existing DPO models. This design yields a 3.6% improvement over the strongest existing DPO baseline on visual question-answering tasks. To support future research, we release our complete framework, including all training data, model checkpoints, and our codebase at https://github.com/dmis-lab/med-vlm-dpo.
Med-CoReasoner: Reducing Language Disparities in Medical Reasoning via Language-Informed Co-Reasoning
Jan 13, 2026Abstract:While reasoning-enhanced large language models perform strongly on English medical tasks, a persistent multilingual gap remains, with substantially weaker reasoning in local languages, limiting equitable global medical deployment. To bridge this gap, we introduce Med-CoReasoner, a language-informed co-reasoning framework that elicits parallel English and local-language reasoning, abstracts them into structured concepts, and integrates local clinical knowledge into an English logical scaffold via concept-level alignment and retrieval. This design combines the structural robustness of English reasoning with the practice-grounded expertise encoded in local languages. To evaluate multilingual medical reasoning beyond multiple-choice settings, we construct MultiMed-X, a benchmark covering seven languages with expert-annotated long-form question answering and natural language inference tasks, comprising 350 instances per language. Experiments across three benchmarks show that Med-CoReasoner improves multilingual reasoning performance by an average of 5%, with particularly substantial gains in low-resource languages. Moreover, model distillation and expert evaluation analysis further confirm that Med-CoReasoner produces clinically sound and culturally grounded reasoning traces.
Rethinking Retrieval-Augmented Generation for Medicine: A Large-Scale, Systematic Expert Evaluation and Practical Insights
Nov 10, 2025Abstract:Large language models (LLMs) are transforming the landscape of medicine, yet two fundamental challenges persist: keeping up with rapidly evolving medical knowledge and providing verifiable, evidence-grounded reasoning. Retrieval-augmented generation (RAG) has been widely adopted to address these limitations by supplementing model outputs with retrieved evidence. However, whether RAG reliably achieves these goals remains unclear. Here, we present the most comprehensive expert evaluation of RAG in medicine to date. Eighteen medical experts contributed a total of 80,502 annotations, assessing 800 model outputs generated by GPT-4o and Llama-3.1-8B across 200 real-world patient and USMLE-style queries. We systematically decomposed the RAG pipeline into three components: (i) evidence retrieval (relevance of retrieved passages), (ii) evidence selection (accuracy of evidence usage), and (iii) response generation (factuality and completeness of outputs). Contrary to expectation, standard RAG often degraded performance: only 22% of top-16 passages were relevant, evidence selection remained weak (precision 41-43%, recall 27-49%), and factuality and completeness dropped by up to 6% and 5%, respectively, compared with non-RAG variants. Retrieval and evidence selection remain key failure points for the model, contributing to the overall performance drop. We further show that simple yet effective strategies, including evidence filtering and query reformulation, substantially mitigate these issues, improving performance on MedMCQA and MedXpertQA by up to 12% and 8.2%, respectively. These findings call for re-examining RAG's role in medicine and highlight the importance of stage-aware evaluation and deliberate system design for reliable medical LLM applications.
Med-PRM: Medical Reasoning Models with Stepwise, Guideline-verified Process Rewards
Jun 13, 2025Abstract:Large language models have shown promise in clinical decision making, but current approaches struggle to localize and correct errors at specific steps of the reasoning process. This limitation is critical in medicine, where identifying and addressing reasoning errors is essential for accurate diagnosis and effective patient care. We introduce Med-PRM, a process reward modeling framework that leverages retrieval-augmented generation to verify each reasoning step against established medical knowledge bases. By verifying intermediate reasoning steps with evidence retrieved from clinical guidelines and literature, our model can precisely assess the reasoning quality in a fine-grained manner. Evaluations on five medical QA benchmarks and two open-ended diagnostic tasks demonstrate that Med-PRM achieves state-of-the-art performance, with improving the performance of base models by up to 13.50% using Med-PRM. Moreover, we demonstrate the generality of Med-PRM by integrating it in a plug-and-play fashion with strong policy models such as Meerkat, achieving over 80\% accuracy on MedQA for the first time using small-scale models of 8 billion parameters. Our code and data are available at: https://med-prm.github.io/
Benchmarking Next-Generation Reasoning-Focused Large Language Models in Ophthalmology: A Head-to-Head Evaluation on 5,888 Items
Apr 15, 2025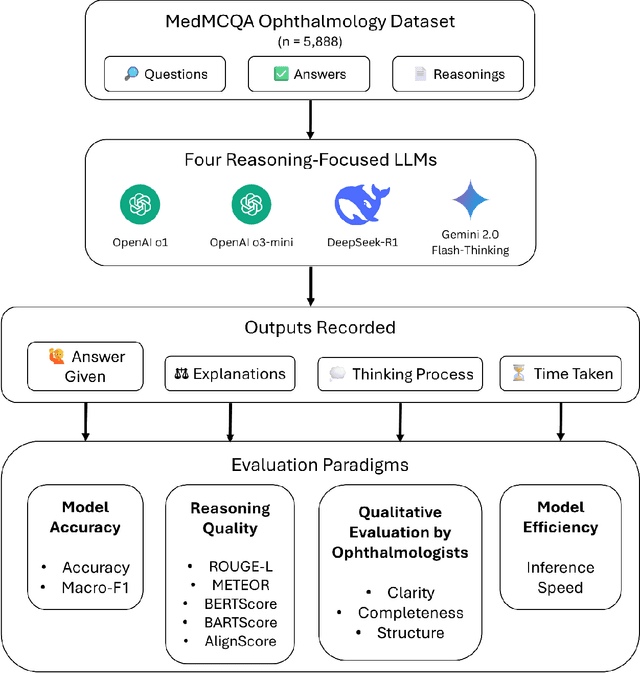
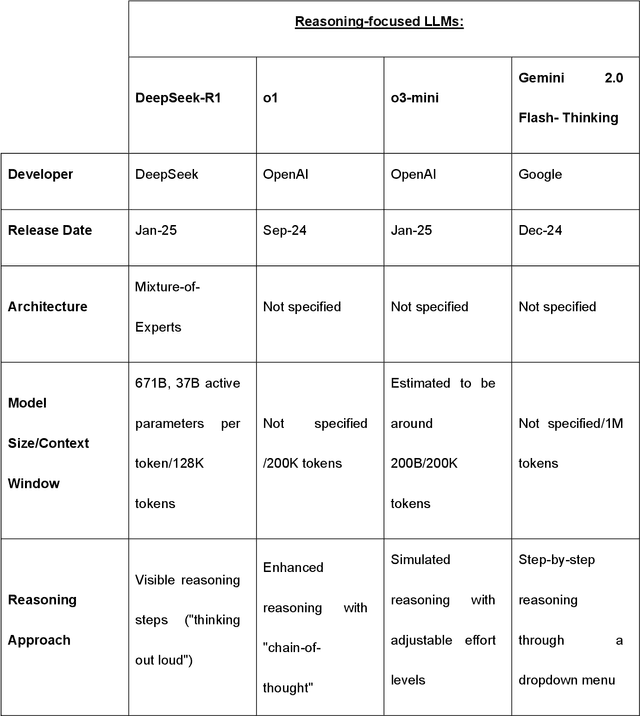
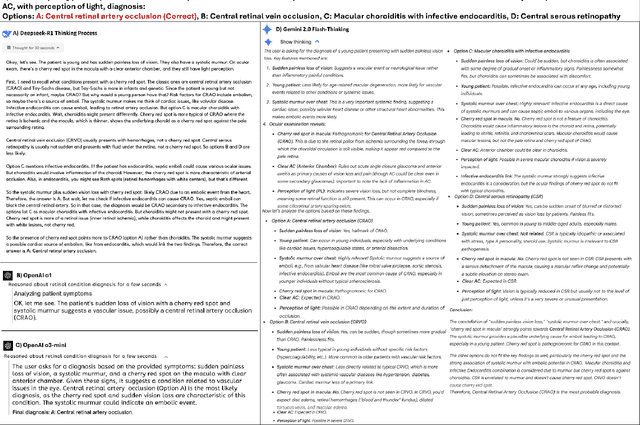
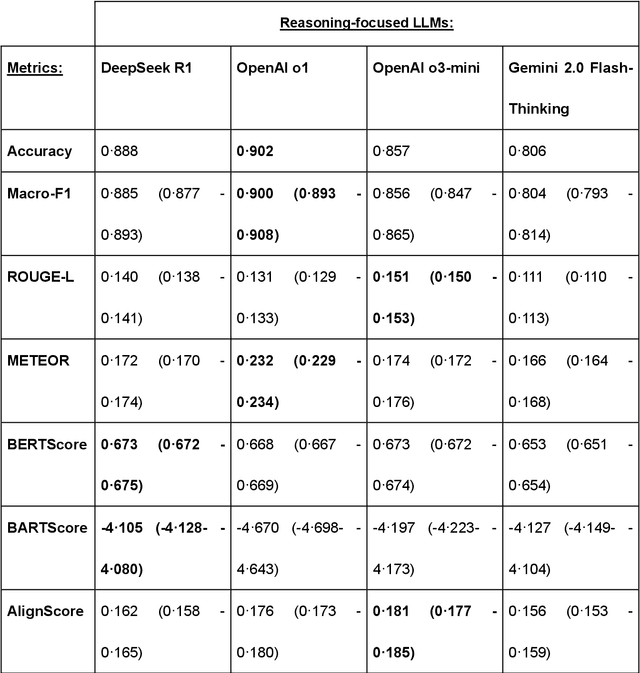
Abstract:Recent advances in reasoning-focused large language models (LLMs) mark a shift from general LLMs toward models designed for complex decision-making, a crucial aspect in medicine. However, their performance in specialized domains like ophthalmology remains underexplored. This study comprehensively evaluated and compared the accuracy and reasoning capabilities of four newly developed reasoning-focused LLMs, namely DeepSeek-R1, OpenAI o1, o3-mini, and Gemini 2.0 Flash-Thinking. Each model was assessed using 5,888 multiple-choice ophthalmology exam questions from the MedMCQA dataset in zero-shot setting. Quantitative evaluation included accuracy, Macro-F1, and five text-generation metrics (ROUGE-L, METEOR, BERTScore, BARTScore, and AlignScore), computed against ground-truth reasonings. Average inference time was recorded for a subset of 100 randomly selected questions. Additionally, two board-certified ophthalmologists qualitatively assessed clarity, completeness, and reasoning structure of responses to differential diagnosis questions.O1 (0.902) and DeepSeek-R1 (0.888) achieved the highest accuracy, with o1 also leading in Macro-F1 (0.900). The performance of models across the text-generation metrics varied: O3-mini excelled in ROUGE-L (0.151), o1 in METEOR (0.232), DeepSeek-R1 and o3-mini tied for BERTScore (0.673), DeepSeek-R1 (-4.105) and Gemini 2.0 Flash-Thinking (-4.127) performed best in BARTScore, while o3-mini (0.181) and o1 (0.176) led AlignScore. Inference time across the models varied, with DeepSeek-R1 being slowest (40.4 seconds) and Gemini 2.0 Flash-Thinking fastest (6.7 seconds). Qualitative evaluation revealed that DeepSeek-R1 and Gemini 2.0 Flash-Thinking tended to provide detailed and comprehensive intermediate reasoning, whereas o1 and o3-mini displayed concise and summarized justifications.
Rationale-Guided Retrieval Augmented Generation for Medical Question Answering
Nov 01, 2024



Abstract:Large language models (LLM) hold significant potential for applications in biomedicine, but they struggle with hallucinations and outdated knowledge. While retrieval-augmented generation (RAG) is generally employed to address these issues, it also has its own set of challenges: (1) LLMs are vulnerable to irrelevant or incorrect context, (2) medical queries are often not well-targeted for helpful information, and (3) retrievers are prone to bias toward the specific source corpus they were trained on. In this study, we present RAG$^2$ (RAtionale-Guided RAG), a new framework for enhancing the reliability of RAG in biomedical contexts. RAG$^2$ incorporates three key innovations: a small filtering model trained on perplexity-based labels of rationales, which selectively augments informative snippets of documents while filtering out distractors; LLM-generated rationales as queries to improve the utility of retrieved snippets; a structure designed to retrieve snippets evenly from a comprehensive set of four biomedical corpora, effectively mitigating retriever bias. Our experiments demonstrate that RAG$^2$ improves the state-of-the-art LLMs of varying sizes, with improvements of up to 6.1\%, and it outperforms the previous best medical RAG model by up to 5.6\% across three medical question-answering benchmarks. Our code is available at https://github.com/dmis-lab/RAG2.
ETHIC: Evaluating Large Language Models on Long-Context Tasks with High Information Coverage
Oct 22, 2024



Abstract:Recent advancements in large language models (LLM) capable of processing extremely long texts highlight the need for a dedicated evaluation benchmark to assess their long-context capabilities. However, existing methods, like the needle-in-a-haystack test, do not effectively assess whether these models fully utilize contextual information, raising concerns about the reliability of current evaluation techniques. To thoroughly examine the effectiveness of existing benchmarks, we introduce a new metric called information coverage (IC), which quantifies the proportion of the input context necessary for answering queries. Our findings indicate that current benchmarks exhibit low IC; although the input context may be extensive, the actual usable context is often limited. To address this, we present ETHIC, a novel benchmark designed to assess LLMs' ability to leverage the entire context. Our benchmark comprises 2,648 test instances spanning four long-context tasks with high IC scores in the domains of books, debates, medicine, and law. Our evaluations reveal significant performance drops in contemporary LLMs, highlighting a critical challenge in managing long contexts. Our benchmark is available at https://github.com/dmis-lab/ETHIC.
Learning from Negative Samples in Generative Biomedical Entity Linking
Aug 29, 2024



Abstract:Generative models have become widely used in biomedical entity linking (BioEL) due to their excellent performance and efficient memory usage. However, these models are usually trained only with positive samples--entities that match the input mention's identifier--and do not explicitly learn from hard negative samples, which are entities that look similar but have different meanings. To address this limitation, we introduce ANGEL (Learning from Negative Samples in Generative Biomedical Entity Linking), the first framework that trains generative BioEL models using negative samples. Specifically, a generative model is initially trained to generate positive samples from the knowledge base for given input entities. Subsequently, both correct and incorrect outputs are gathered from the model's top-k predictions. The model is then updated to prioritize the correct predictions through direct preference optimization. Our models fine-tuned with ANGEL outperform the previous best baseline models by up to an average top-1 accuracy of 1.4% on five benchmarks. When incorporating our framework into pre-training, the performance improvement further increases to 1.7%, demonstrating its effectiveness in both the pre-training and fine-tuning stages. Our code is available at https://github.com/dmis-lab/ANGEL.
Augmenting Biomedical Named Entity Recognition with General-domain Resources
Jun 15, 2024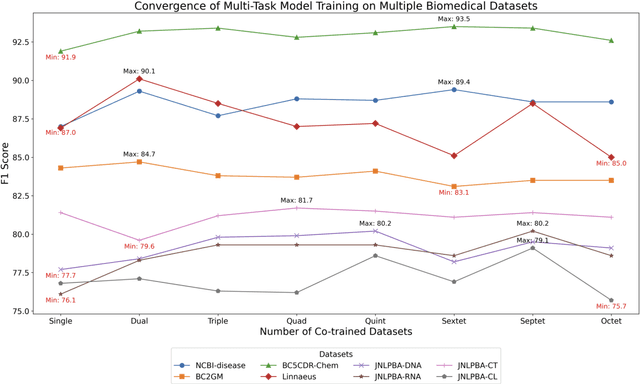
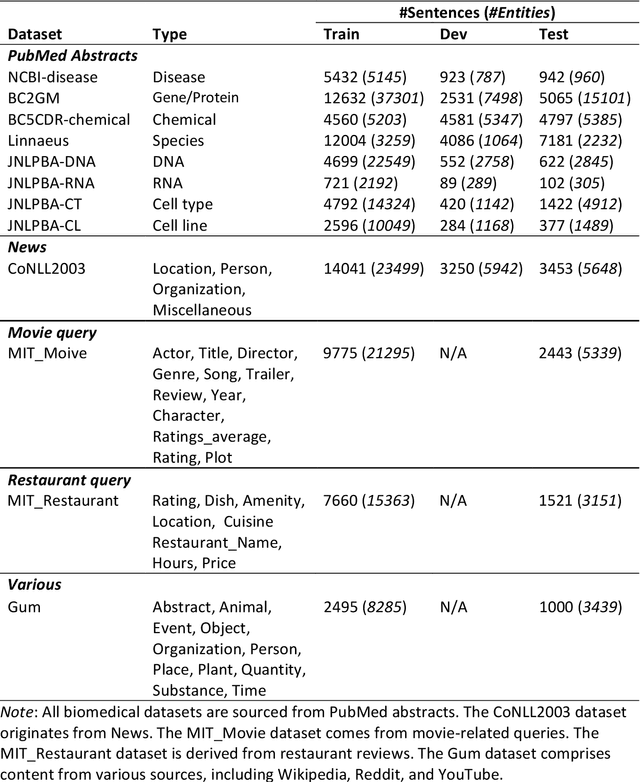

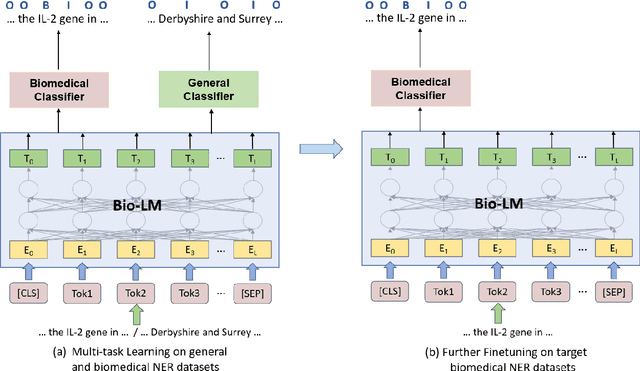
Abstract:Training a neural network-based biomedical named entity recognition (BioNER) model usually requires extensive and costly human annotations. While several studies have employed multi-task learning with multiple BioNER datasets to reduce human effort, this approach does not consistently yield performance improvements and may introduce label ambiguity in different biomedical corpora. We aim to tackle those challenges through transfer learning from easily accessible resources with fewer concept overlaps with biomedical datasets. In this paper, we proposed GERBERA, a simple-yet-effective method that utilized a general-domain NER dataset for training. Specifically, we performed multi-task learning to train a pre-trained biomedical language model with both the target BioNER dataset and the general-domain dataset. Subsequently, we fine-tuned the models specifically for the BioNER dataset. We systematically evaluated GERBERA on five datasets of eight entity types, collectively consisting of 81,410 instances. Despite using fewer biomedical resources, our models demonstrated superior performance compared to baseline models trained with multiple additional BioNER datasets. Specifically, our models consistently outperformed the baselines in six out of eight entity types, achieving an average improvement of 0.9% over the best baseline performance across eight biomedical entity types sourced from five different corpora. Our method was especially effective in amplifying performance on BioNER datasets characterized by limited data, with a 4.7% improvement in F1 scores on the JNLPBA-RNA dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge