Gregor Koehler
Beyond Benchmarks of IUGC: Rethinking Requirements of Deep Learning Methods for Intrapartum Ultrasound Biometry from Fetal Ultrasound Videos
Feb 13, 2026Abstract:A substantial proportion (45\%) of maternal deaths, neonatal deaths, and stillbirths occur during the intrapartum phase, with a particularly high burden in low- and middle-income countries. Intrapartum biometry plays a critical role in monitoring labor progression; however, the routine use of ultrasound in resource-limited settings is hindered by a shortage of trained sonographers. To address this challenge, the Intrapartum Ultrasound Grand Challenge (IUGC), co-hosted with MICCAI 2024, was launched. The IUGC introduces a clinically oriented multi-task automatic measurement framework that integrates standard plane classification, fetal head-pubic symphysis segmentation, and biometry, enabling algorithms to exploit complementary task information for more accurate estimation. Furthermore, the challenge releases the largest multi-center intrapartum ultrasound video dataset to date, comprising 774 videos (68,106 frames) collected from three hospitals, providing a robust foundation for model training and evaluation. In this study, we present a comprehensive overview of the challenge design, review the submissions from eight participating teams, and analyze their methods from five perspectives: preprocessing, data augmentation, learning strategy, model architecture, and post-processing. In addition, we perform a systematic analysis of the benchmark results to identify key bottlenecks, explore potential solutions, and highlight open challenges for future research. Although encouraging performance has been achieved, our findings indicate that the field remains at an early stage, and further in-depth investigation is required before large-scale clinical deployment. All benchmark solutions and the complete dataset have been publicly released to facilitate reproducible research and promote continued advances in automatic intrapartum ultrasound biometry.
PSFHS Challenge Report: Pubic Symphysis and Fetal Head Segmentation from Intrapartum Ultrasound Images
Sep 17, 2024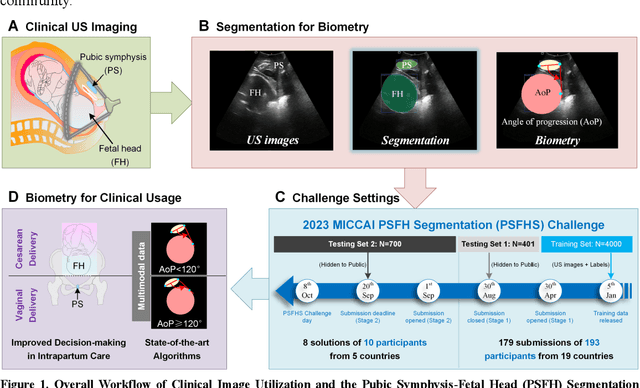
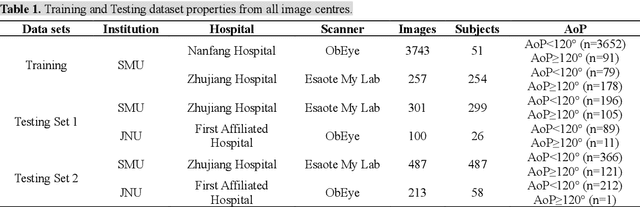
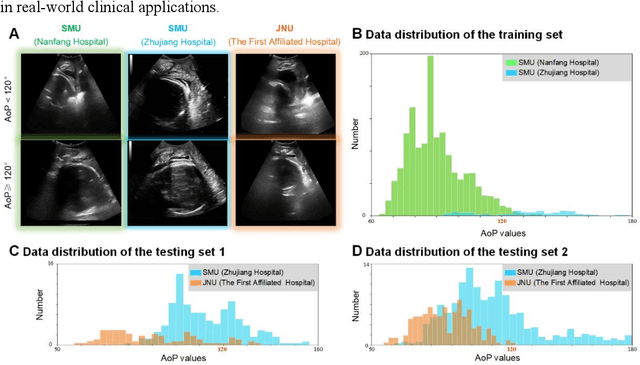

Abstract:Segmentation of the fetal and maternal structures, particularly intrapartum ultrasound imaging as advocated by the International Society of Ultrasound in Obstetrics and Gynecology (ISUOG) for monitoring labor progression, is a crucial first step for quantitative diagnosis and clinical decision-making. This requires specialized analysis by obstetrics professionals, in a task that i) is highly time- and cost-consuming and ii) often yields inconsistent results. The utility of automatic segmentation algorithms for biometry has been proven, though existing results remain suboptimal. To push forward advancements in this area, the Grand Challenge on Pubic Symphysis-Fetal Head Segmentation (PSFHS) was held alongside the 26th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2023). This challenge aimed to enhance the development of automatic segmentation algorithms at an international scale, providing the largest dataset to date with 5,101 intrapartum ultrasound images collected from two ultrasound machines across three hospitals from two institutions. The scientific community's enthusiastic participation led to the selection of the top 8 out of 179 entries from 193 registrants in the initial phase to proceed to the competition's second stage. These algorithms have elevated the state-of-the-art in automatic PSFHS from intrapartum ultrasound images. A thorough analysis of the results pinpointed ongoing challenges in the field and outlined recommendations for future work. The top solutions and the complete dataset remain publicly available, fostering further advancements in automatic segmentation and biometry for intrapartum ultrasound imaging.
New Horizons in Parameter Regularization: A Constraint Approach
Nov 15, 2023



Abstract:This work presents constrained parameter regularization (CPR), an alternative to traditional weight decay. Instead of applying a constant penalty uniformly to all parameters, we enforce an upper bound on a statistical measure (e.g., the L$_2$-norm) of individual parameter groups. This reformulates learning as a constrained optimization problem. To solve this, we utilize an adaptation of the augmented Lagrangian method. Our approach allows for varying regularization strengths across different parameter groups, removing the need for explicit penalty coefficients in the regularization terms. CPR only requires two hyperparameters and introduces no measurable runtime overhead. We offer empirical evidence of CPR's effectiveness through experiments in the "grokking" phenomenon, image classification, and language modeling. Our findings show that CPR can counteract the effects of grokking, and it consistently matches or surpasses the performance of traditional weight decay.
RecycleNet: Latent Feature Recycling Leads to Iterative Decision Refinement
Sep 14, 2023



Abstract:Despite the remarkable success of deep learning systems over the last decade, a key difference still remains between neural network and human decision-making: As humans, we cannot only form a decision on the spot, but also ponder, revisiting an initial guess from different angles, distilling relevant information, arriving at a better decision. Here, we propose RecycleNet, a latent feature recycling method, instilling the pondering capability for neural networks to refine initial decisions over a number of recycling steps, where outputs are fed back into earlier network layers in an iterative fashion. This approach makes minimal assumptions about the neural network architecture and thus can be implemented in a wide variety of contexts. Using medical image segmentation as the evaluation environment, we show that latent feature recycling enables the network to iteratively refine initial predictions even beyond the iterations seen during training, converging towards an improved decision. We evaluate this across a variety of segmentation benchmarks and show consistent improvements even compared with top-performing segmentation methods. This allows trading increased computation time for improved performance, which can be beneficial, especially for safety-critical applications.
Unleashing the Strengths of Unlabeled Data in Pan-cancer Abdominal Organ Quantification: the FLARE22 Challenge
Aug 10, 2023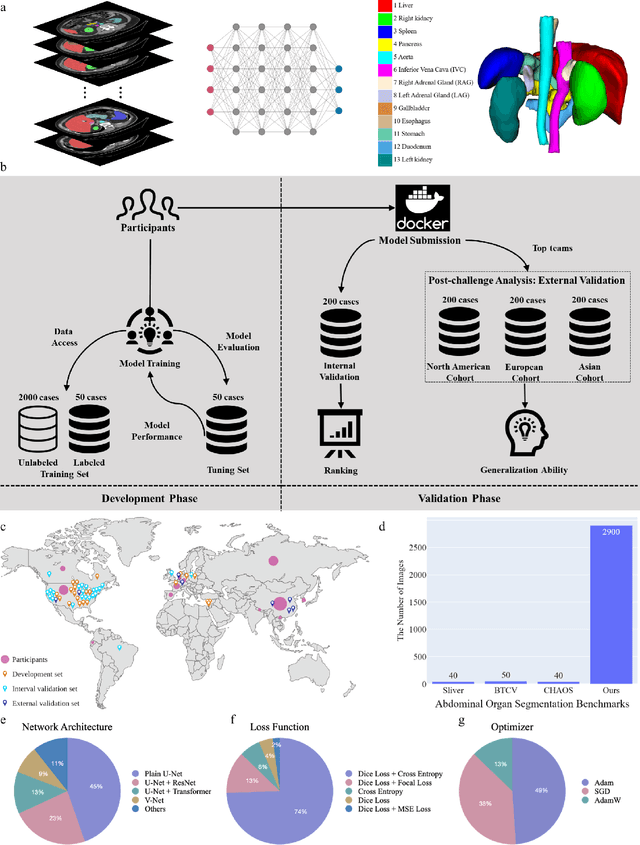
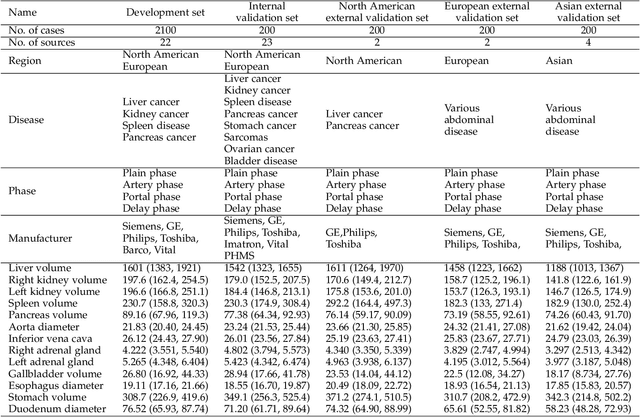
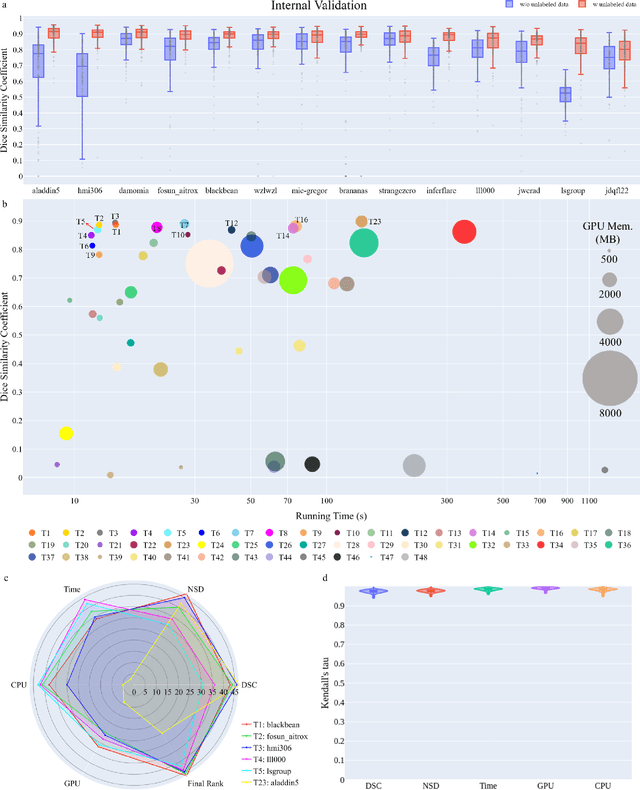
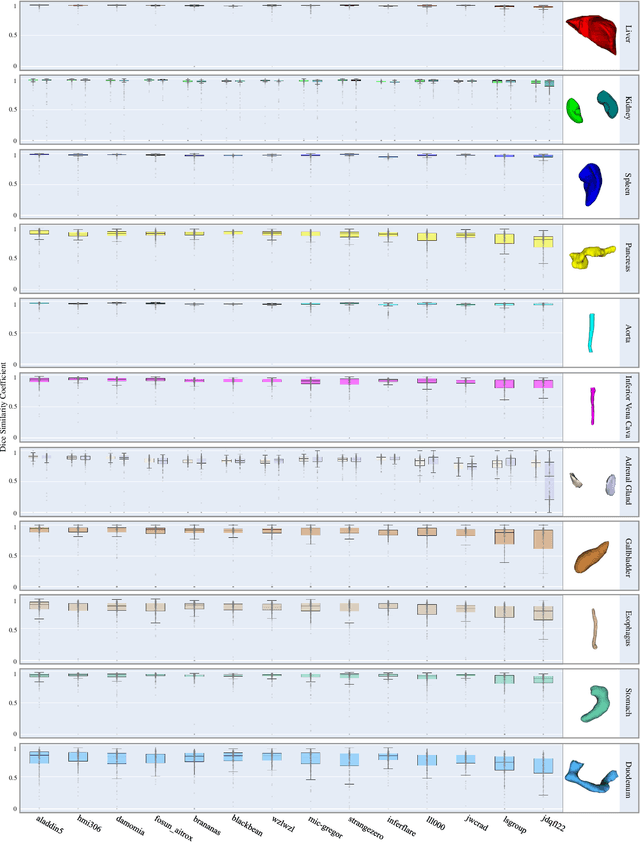
Abstract:Quantitative organ assessment is an essential step in automated abdominal disease diagnosis and treatment planning. Artificial intelligence (AI) has shown great potential to automatize this process. However, most existing AI algorithms rely on many expert annotations and lack a comprehensive evaluation of accuracy and efficiency in real-world multinational settings. To overcome these limitations, we organized the FLARE 2022 Challenge, the largest abdominal organ analysis challenge to date, to benchmark fast, low-resource, accurate, annotation-efficient, and generalized AI algorithms. We constructed an intercontinental and multinational dataset from more than 50 medical groups, including Computed Tomography (CT) scans with different races, diseases, phases, and manufacturers. We independently validated that a set of AI algorithms achieved a median Dice Similarity Coefficient (DSC) of 90.0\% by using 50 labeled scans and 2000 unlabeled scans, which can significantly reduce annotation requirements. The best-performing algorithms successfully generalized to holdout external validation sets, achieving a median DSC of 89.5\%, 90.9\%, and 88.3\% on North American, European, and Asian cohorts, respectively. They also enabled automatic extraction of key organ biology features, which was labor-intensive with traditional manual measurements. This opens the potential to use unlabeled data to boost performance and alleviate annotation shortages for modern AI models.
Exploring new ways: Enforcing representational dissimilarity to learn new features and reduce error consistency
Jul 05, 2023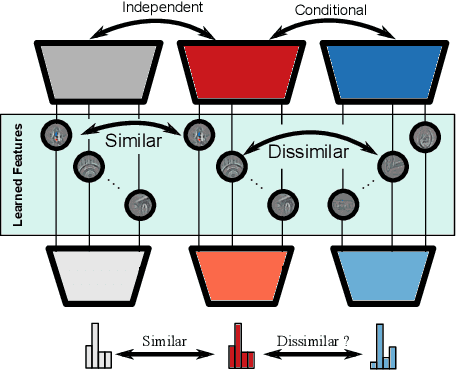
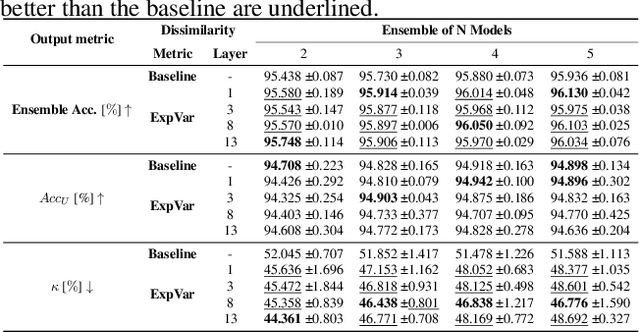

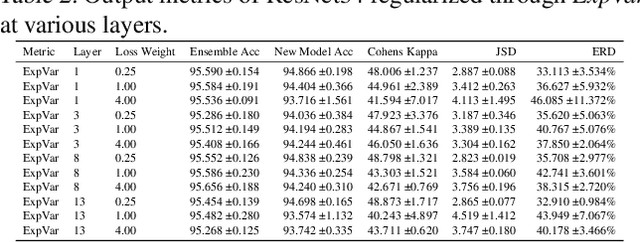
Abstract:Independently trained machine learning models tend to learn similar features. Given an ensemble of independently trained models, this results in correlated predictions and common failure modes. Previous attempts focusing on decorrelation of output predictions or logits yielded mixed results, particularly due to their reduction in model accuracy caused by conflicting optimization objectives. In this paper, we propose the novel idea of utilizing methods of the representational similarity field to promote dissimilarity during training instead of measuring similarity of trained models. To this end, we promote intermediate representations to be dissimilar at different depths between architectures, with the goal of learning robust ensembles with disjoint failure modes. We show that highly dissimilar intermediate representations result in less correlated output predictions and slightly lower error consistency, resulting in higher ensemble accuracy. With this, we shine first light on the connection between intermediate representations and their impact on the output predictions.
SAM.MD: Zero-shot medical image segmentation capabilities of the Segment Anything Model
Apr 10, 2023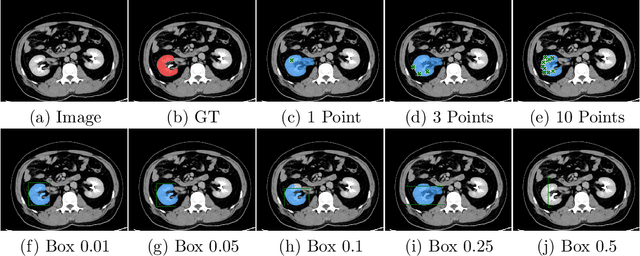
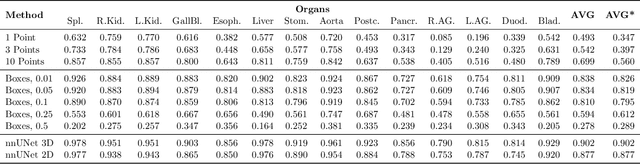
Abstract:Foundation models have taken over natural language processing and image generation domains due to the flexibility of prompting. With the recent introduction of the Segment Anything Model (SAM), this prompt-driven paradigm has entered image segmentation with a hitherto unexplored abundance of capabilities. The purpose of this paper is to conduct an initial evaluation of the out-of-the-box zero-shot capabilities of SAM for medical image segmentation, by evaluating its performance on an abdominal CT organ segmentation task, via point or bounding box based prompting. We show that SAM generalizes well to CT data, making it a potential catalyst for the advancement of semi-automatic segmentation tools for clinicians. We believe that this foundation model, while not reaching state-of-the-art segmentation performance in our investigations, can serve as a highly potent starting point for further adaptations of such models to the intricacies of the medical domain. Keywords: medical image segmentation, SAM, foundation models, zero-shot learning
Transformer Utilization in Medical Image Segmentation Networks
Apr 09, 2023Abstract:Owing to success in the data-rich domain of natural images, Transformers have recently become popular in medical image segmentation. However, the pairing of Transformers with convolutional blocks in varying architectural permutations leaves their relative effectiveness to open interpretation. We introduce Transformer Ablations that replace the Transformer blocks with plain linear operators to quantify this effectiveness. With experiments on 8 models on 2 medical image segmentation tasks, we explore -- 1) the replaceable nature of Transformer-learnt representations, 2) Transformer capacity alone cannot prevent representational replaceability and works in tandem with effective design, 3) The mere existence of explicit feature hierarchies in transformer blocks is more beneficial than accompanying self-attention modules, 4) Major spatial downsampling before Transformer modules should be used with caution.
MedNeXt: Transformer-driven Scaling of ConvNets for Medical Image Segmentation
Mar 22, 2023



Abstract:There has been exploding interest in embracing Transformer-based architectures for medical image segmentation. However, the lack of large-scale annotated medical datasets make achieving performances equivalent to those in natural images challenging. Convolutional networks, in contrast, have higher inductive biases and consequently, are easily trainable to high performance. Recently, the ConvNeXt architecture attempted to modernize the standard ConvNet by mirroring Transformer blocks. In this work, we improve upon this to design a modernized and scalable convolutional architecture customized to challenges of data-scarce medical settings. We introduce MedNeXt, a Transformer-inspired large kernel segmentation network which introduces - 1) A fully ConvNeXt 3D Encoder-Decoder Network for medical image segmentation, 2) Residual ConvNeXt up and downsampling blocks to preserve semantic richness across scales, 3) A novel technique to iteratively increase kernel sizes by upsampling small kernel networks, to prevent performance saturation on limited medical data, 4) Compound scaling at multiple levels (depth, width, kernel size) of MedNeXt. This leads to state-of-the-art performance on 4 tasks on CT and MRI modalities and varying dataset sizes, representing a modernized deep architecture for medical image segmentation.
CRADL: Contrastive Representations for Unsupervised Anomaly Detection and Localization
Jan 05, 2023



Abstract:Unsupervised anomaly detection in medical imaging aims to detect and localize arbitrary anomalies without requiring annotated anomalous data during training. Often, this is achieved by learning a data distribution of normal samples and detecting anomalies as regions in the image which deviate from this distribution. Most current state-of-the-art methods use latent variable generative models operating directly on the images. However, generative models have been shown to mostly capture low-level features, s.a. pixel-intensities, instead of rich semantic features, which also applies to their representations. We circumvent this problem by proposing CRADL whose core idea is to model the distribution of normal samples directly in the low-dimensional representation space of an encoder trained with a contrastive pretext-task. By utilizing the representations of contrastive learning, we aim to fix the over-fixation on low-level features and learn more semantic-rich representations. Our experiments on anomaly detection and localization tasks using three distinct evaluation datasets show that 1) contrastive representations are superior to representations of generative latent variable models and 2) the CRADL framework shows competitive or superior performance to state-of-the-art.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge