Constantin Ulrich
MedNeXt-v2: Scaling 3D ConvNeXts for Large-Scale Supervised Representation Learning in Medical Image Segmentation
Dec 19, 2025Abstract:Large-scale supervised pretraining is rapidly reshaping 3D medical image segmentation. However, existing efforts focus primarily on increasing dataset size and overlook the question of whether the backbone network is an effective representation learner at scale. In this work, we address this gap by revisiting ConvNeXt-based architectures for volumetric segmentation and introducing MedNeXt-v2, a compound-scaled 3D ConvNeXt that leverages improved micro-architecture and data scaling to deliver state-of-the-art performance. First, we show that routinely used backbones in large-scale pretraining pipelines are often suboptimal. Subsequently, we use comprehensive backbone benchmarking prior to scaling and demonstrate that stronger from scratch performance reliably predicts stronger downstream performance after pretraining. Guided by these findings, we incorporate a 3D Global Response Normalization module and use depth, width, and context scaling to improve our architecture for effective representation learning. We pretrain MedNeXt-v2 on 18k CT volumes and demonstrate state-of-the-art performance when fine-tuning across six challenging CT and MR benchmarks (144 structures), showing consistent gains over seven publicly released pretrained models. Beyond improvements, our benchmarking of these models also reveals that stronger backbones yield better results on similar data, representation scaling disproportionately benefits pathological segmentation, and that modality-specific pretraining offers negligible benefit once full finetuning is applied. In conclusion, our results establish MedNeXt-v2 as a strong backbone for large-scale supervised representation learning in 3D Medical Image Segmentation. Our code and pretrained models are made available with the official nnUNet repository at: https://www.github.com/MIC-DKFZ/nnUNet
CRONOS: Continuous Time Reconstruction for 4D Medical Longitudinal Series
Dec 18, 2025Abstract:Forecasting how 3D medical scans evolve over time is important for disease progression, treatment planning, and developmental assessment. Yet existing models either rely on a single prior scan, fixed grid times, or target global labels, which limits voxel-level forecasting under irregular sampling. We present CRONOS, a unified framework for many-to-one prediction from multiple past scans that supports both discrete (grid-based) and continuous (real-valued) timestamps in one model, to the best of our knowledge the first to achieve continuous sequence-to-image forecasting for 3D medical data. CRONOS learns a spatio-temporal velocity field that transports context volumes toward a target volume at an arbitrary time, while operating directly in 3D voxel space. Across three public datasets spanning Cine-MRI, perfusion CT, and longitudinal MRI, CRONOS outperforms other baselines, while remaining computationally competitive. We will release code and evaluation protocols to enable reproducible, multi-dataset benchmarking of multi-context, continuous-time forecasting.
VoxTell: Free-Text Promptable Universal 3D Medical Image Segmentation
Nov 14, 2025Abstract:We introduce VoxTell, a vision-language model for text-prompted volumetric medical image segmentation. It maps free-form descriptions, from single words to full clinical sentences, to 3D masks. Trained on 62K+ CT, MRI, and PET volumes spanning over 1K anatomical and pathological classes, VoxTell uses multi-stage vision-language fusion across decoder layers to align textual and visual features at multiple scales. It achieves state-of-the-art zero-shot performance across modalities on unseen datasets, excelling on familiar concepts while generalizing to related unseen classes. Extensive experiments further demonstrate strong cross-modality transfer, robustness to linguistic variations and clinical language, as well as accurate instance-specific segmentation from real-world text. Code is available at: https://www.github.com/MIC-DKFZ/VoxTell
The Missing Piece: A Case for Pre-Training in 3D Medical Object Detection
Sep 19, 2025Abstract:Large-scale pre-training holds the promise to advance 3D medical object detection, a crucial component of accurate computer-aided diagnosis. Yet, it remains underexplored compared to segmentation, where pre-training has already demonstrated significant benefits. Existing pre-training approaches for 3D object detection rely on 2D medical data or natural image pre-training, failing to fully leverage 3D volumetric information. In this work, we present the first systematic study of how existing pre-training methods can be integrated into state-of-the-art detection architectures, covering both CNNs and Transformers. Our results show that pre-training consistently improves detection performance across various tasks and datasets. Notably, reconstruction-based self-supervised pre-training outperforms supervised pre-training, while contrastive pre-training provides no clear benefit for 3D medical object detection. Our code is publicly available at: https://github.com/MIC-DKFZ/nnDetection-finetuning.
* MICCAI 2025
Temporal Flow Matching for Learning Spatio-Temporal Trajectories in 4D Longitudinal Medical Imaging
Aug 29, 2025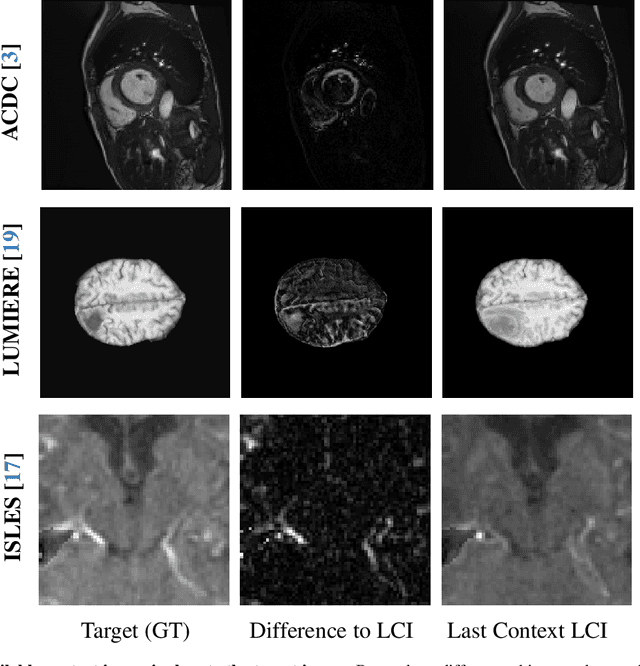
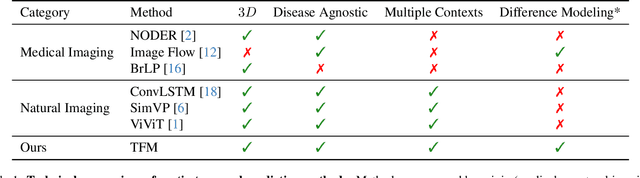

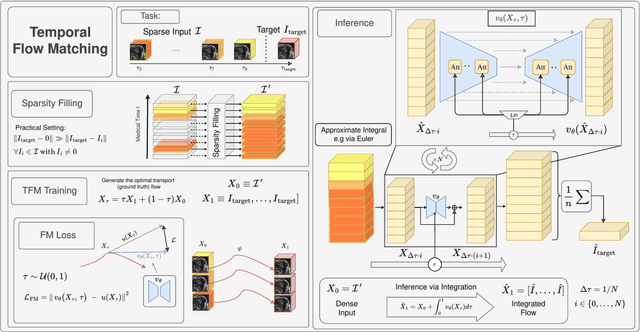
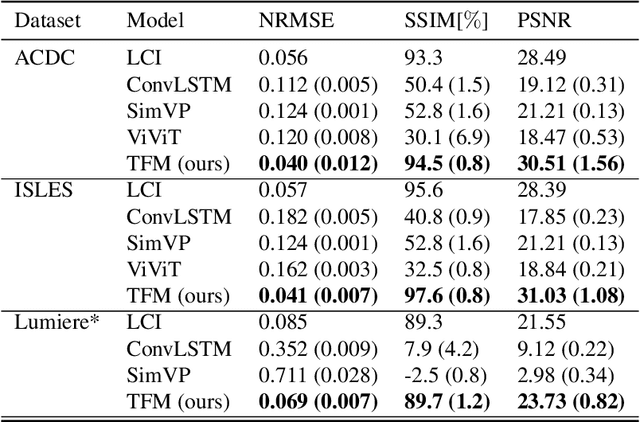
Abstract:Understanding temporal dynamics in medical imaging is crucial for applications such as disease progression modeling, treatment planning and anatomical development tracking. However, most deep learning methods either consider only single temporal contexts, or focus on tasks like classification or regression, limiting their ability for fine-grained spatial predictions. While some approaches have been explored, they are often limited to single timepoints, specific diseases or have other technical restrictions. To address this fundamental gap, we introduce Temporal Flow Matching (TFM), a unified generative trajectory method that (i) aims to learn the underlying temporal distribution, (ii) by design can fall back to a nearest image predictor, i.e. predicting the last context image (LCI), as a special case, and (iii) supports $3D$ volumes, multiple prior scans, and irregular sampling. Extensive benchmarks on three public longitudinal datasets show that TFM consistently surpasses spatio-temporal methods from natural imaging, establishing a new state-of-the-art and robust baseline for $4D$ medical image prediction.
Large Scale Supervised Pretraining For Traumatic Brain Injury Segmentation
Apr 09, 2025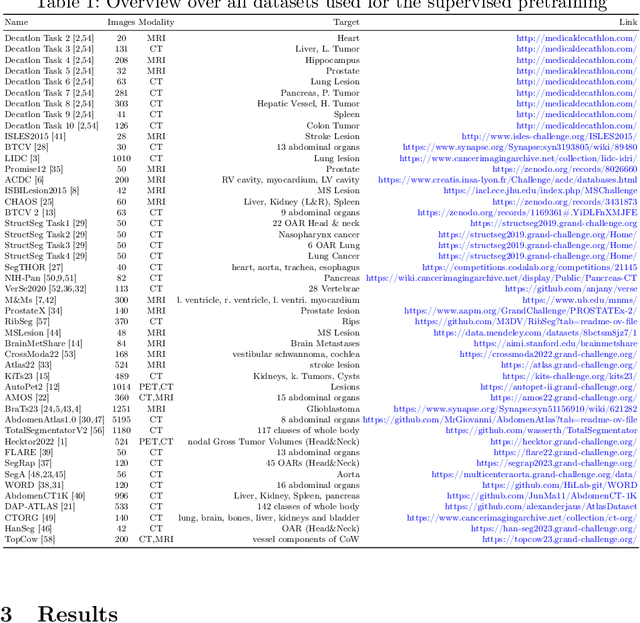
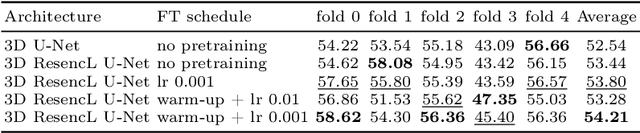
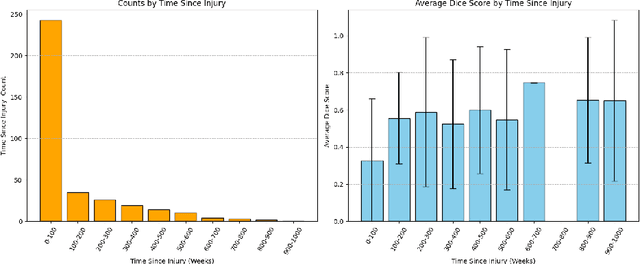
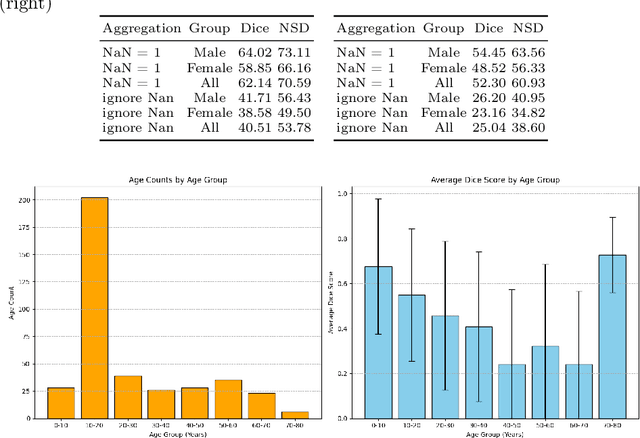
Abstract:The segmentation of lesions in Moderate to Severe Traumatic Brain Injury (msTBI) presents a significant challenge in neuroimaging due to the diverse characteristics of these lesions, which vary in size, shape, and distribution across brain regions and tissue types. This heterogeneity complicates traditional image processing techniques, resulting in critical errors in tasks such as image registration and brain parcellation. To address these challenges, the AIMS-TBI Segmentation Challenge 2024 aims to advance innovative segmentation algorithms specifically designed for T1-weighted MRI data, the most widely utilized imaging modality in clinical practice. Our proposed solution leverages a large-scale multi-dataset supervised pretraining approach inspired by the MultiTalent method. We train a Resenc L network on a comprehensive collection of datasets covering various anatomical and pathological structures, which equips the model with a robust understanding of brain anatomy and pathology. Following this, the model is fine-tuned on msTBI-specific data to optimize its performance for the unique characteristics of T1-weighted MRI scans and outperforms the baseline without pretraining up to 2 Dice points.
Enhanced Diagnostic Fidelity in Pathology Whole Slide Image Compression via Deep Learning
Mar 14, 2025Abstract:Accurate diagnosis of disease often depends on the exhaustive examination of Whole Slide Images (WSI) at microscopic resolution. Efficient handling of these data-intensive images requires lossy compression techniques. This paper investigates the limitations of the widely-used JPEG algorithm, the current clinical standard, and reveals severe image artifacts impacting diagnostic fidelity. To overcome these challenges, we introduce a novel deep-learning (DL)-based compression method tailored for pathology images. By enforcing feature similarity of deep features between the original and compressed images, our approach achieves superior Peak Signal-to-Noise Ratio (PSNR), Multi-Scale Structural Similarity Index (MS-SSIM), and Learned Perceptual Image Patch Similarity (LPIPS) scores compared to JPEG-XL, Webp, and other DL compression methods.
nnInteractive: Redefining 3D Promptable Segmentation
Mar 11, 2025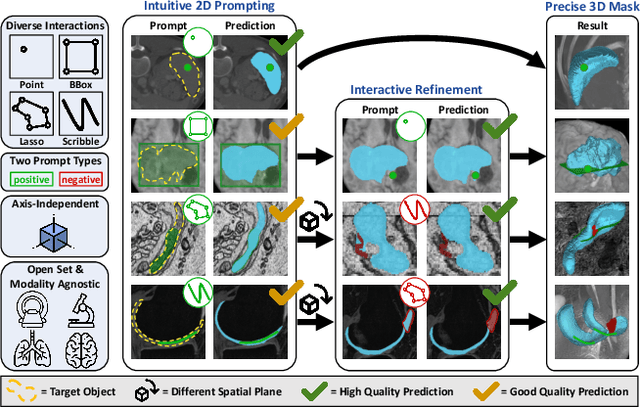
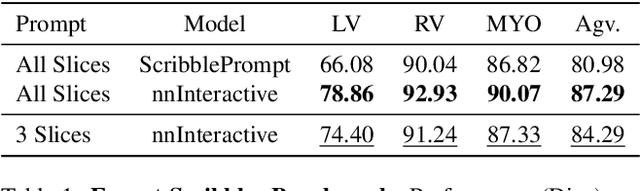
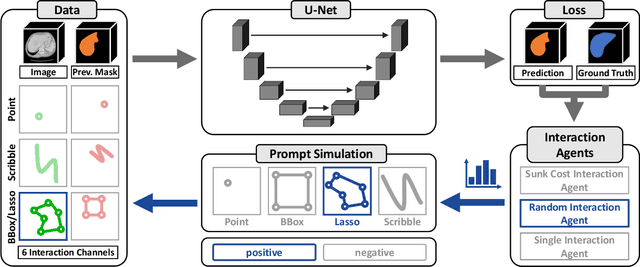
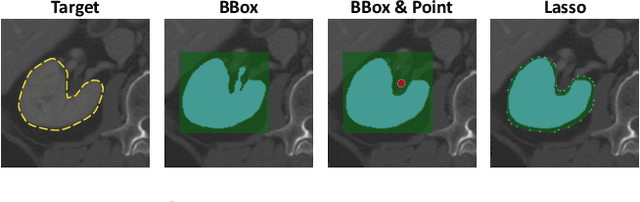
Abstract:Accurate and efficient 3D segmentation is essential for both clinical and research applications. While foundation models like SAM have revolutionized interactive segmentation, their 2D design and domain shift limitations make them ill-suited for 3D medical images. Current adaptations address some of these challenges but remain limited, either lacking volumetric awareness, offering restricted interactivity, or supporting only a small set of structures and modalities. Usability also remains a challenge, as current tools are rarely integrated into established imaging platforms and often rely on cumbersome web-based interfaces with restricted functionality. We introduce nnInteractive, the first comprehensive 3D interactive open-set segmentation method. It supports diverse prompts-including points, scribbles, boxes, and a novel lasso prompt-while leveraging intuitive 2D interactions to generate full 3D segmentations. Trained on 120+ diverse volumetric 3D datasets (CT, MRI, PET, 3D Microscopy, etc.), nnInteractive sets a new state-of-the-art in accuracy, adaptability, and usability. Crucially, it is the first method integrated into widely used image viewers (e.g., Napari, MITK), ensuring broad accessibility for real-world clinical and research applications. Extensive benchmarking demonstrates that nnInteractive far surpasses existing methods, setting a new standard for AI-driven interactive 3D segmentation. nnInteractive is publicly available: https://github.com/MIC-DKFZ/napari-nninteractive (Napari plugin), https://www.mitk.org/MITK-nnInteractive (MITK integration), https://github.com/MIC-DKFZ/nnInteractive (Python backend).
Primus: Enforcing Attention Usage for 3D Medical Image Segmentation
Mar 03, 2025
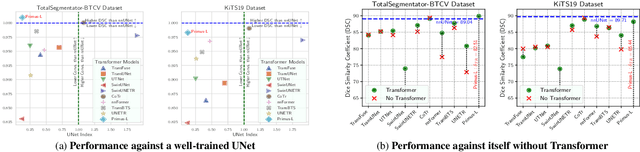
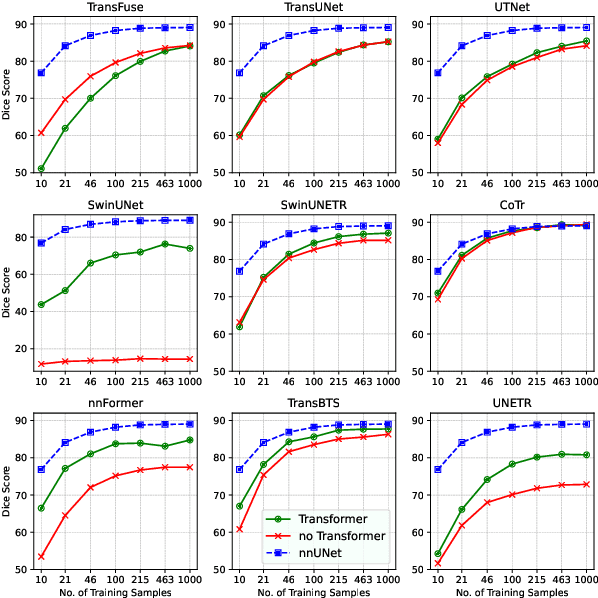
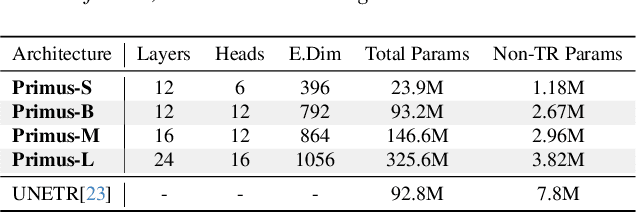
Abstract:Transformers have achieved remarkable success across multiple fields, yet their impact on 3D medical image segmentation remains limited with convolutional networks still dominating major benchmarks. In this work, we a) analyze current Transformer-based segmentation models and identify critical shortcomings, particularly their over-reliance on convolutional blocks. Further, we demonstrate that in some architectures, performance is unaffected by the absence of the Transformer, thereby demonstrating their limited effectiveness. To address these challenges, we move away from hybrid architectures and b) introduce a fully Transformer-based segmentation architecture, termed Primus. Primus leverages high-resolution tokens, combined with advances in positional embeddings and block design, to maximally leverage its Transformer blocks. Through these adaptations Primus surpasses current Transformer-based methods and competes with state-of-the-art convolutional models on multiple public datasets. By doing so, we create the first pure Transformer architecture and take a significant step towards making Transformers state-of-the-art for 3D medical image segmentation.
LesionLocator: Zero-Shot Universal Tumor Segmentation and Tracking in 3D Whole-Body Imaging
Feb 28, 2025Abstract:In this work, we present LesionLocator, a framework for zero-shot longitudinal lesion tracking and segmentation in 3D medical imaging, establishing the first end-to-end model capable of 4D tracking with dense spatial prompts. Our model leverages an extensive dataset of 23,262 annotated medical scans, as well as synthesized longitudinal data across diverse lesion types. The diversity and scale of our dataset significantly enhances model generalizability to real-world medical imaging challenges and addresses key limitations in longitudinal data availability. LesionLocator outperforms all existing promptable models in lesion segmentation by nearly 10 dice points, reaching human-level performance, and achieves state-of-the-art results in lesion tracking, with superior lesion retrieval and segmentation accuracy. LesionLocator not only sets a new benchmark in universal promptable lesion segmentation and automated longitudinal lesion tracking but also provides the first open-access solution of its kind, releasing our synthetic 4D dataset and model to the community, empowering future advancements in medical imaging. Code is available at: www.github.com/MIC-DKFZ/LesionLocator
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge