Maximilian Fischer
Multimodal classification of Radiation-Induced Contrast Enhancements and tumor recurrence using deep learning
Mar 12, 2026Abstract:The differentiation between tumor recurrence and radiation-induced contrast enhancements in post-treatment glioblastoma patients remains a major clinical challenge. Existing approaches rely on clinically sparsely available diffusion MRI or do not consider radiation maps, which are gaining increasing interest in the tumor board for this differentiation. We introduce RICE-NET, a multimodal 3D deep learning model that integrates longitudinal MRI data with radiotherapy dose distributions for automated lesion classification using conventional T1-weighted MRI data. Using a cohort of 92 patients, the model achieved an F1 score of 0.92 on an independent test set. During extensive ablation experiments, we quantified the contribution of each timepoint and modality and showed that reliable classification largely depends on the radiation map. Occlusion-based interpretability analyses further confirmed the model's focus on clinically relevant regions. These findings highlight the potential of multimodal deep learning to enhance diagnostic accuracy and support clinical decision-making in neuro-oncology.
Enhanced Diagnostic Fidelity in Pathology Whole Slide Image Compression via Deep Learning
Mar 14, 2025Abstract:Accurate diagnosis of disease often depends on the exhaustive examination of Whole Slide Images (WSI) at microscopic resolution. Efficient handling of these data-intensive images requires lossy compression techniques. This paper investigates the limitations of the widely-used JPEG algorithm, the current clinical standard, and reveals severe image artifacts impacting diagnostic fidelity. To overcome these challenges, we introduce a novel deep-learning (DL)-based compression method tailored for pathology images. By enforcing feature similarity of deep features between the original and compressed images, our approach achieves superior Peak Signal-to-Noise Ratio (PSNR), Multi-Scale Structural Similarity Index (MS-SSIM), and Learned Perceptual Image Patch Similarity (LPIPS) scores compared to JPEG-XL, Webp, and other DL compression methods.
KPIs 2024 Challenge: Advancing Glomerular Segmentation from Patch- to Slide-Level
Feb 11, 2025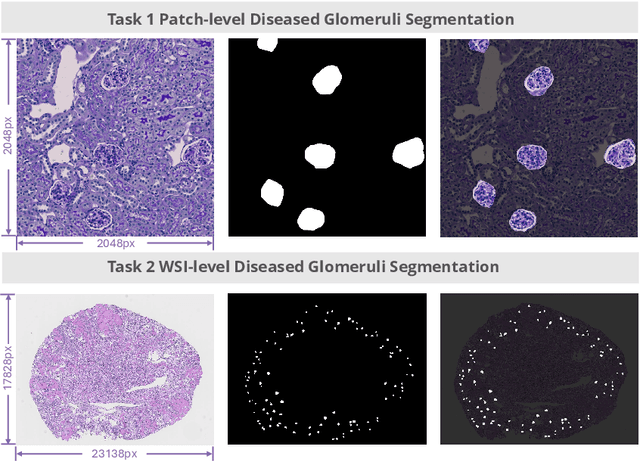
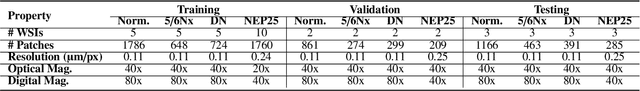
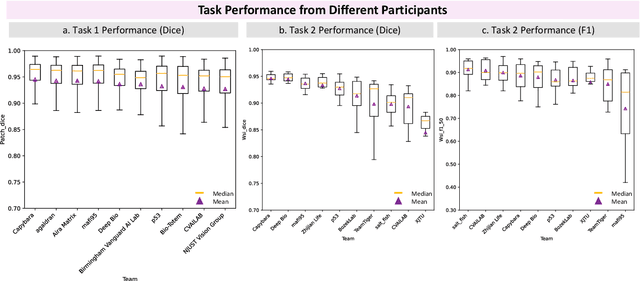

Abstract:Chronic kidney disease (CKD) is a major global health issue, affecting over 10% of the population and causing significant mortality. While kidney biopsy remains the gold standard for CKD diagnosis and treatment, the lack of comprehensive benchmarks for kidney pathology segmentation hinders progress in the field. To address this, we organized the Kidney Pathology Image Segmentation (KPIs) Challenge, introducing a dataset that incorporates preclinical rodent models of CKD with over 10,000 annotated glomeruli from 60+ Periodic Acid Schiff (PAS)-stained whole slide images. The challenge includes two tasks, patch-level segmentation and whole slide image segmentation and detection, evaluated using the Dice Similarity Coefficient (DSC) and F1-score. By encouraging innovative segmentation methods that adapt to diverse CKD models and tissue conditions, the KPIs Challenge aims to advance kidney pathology analysis, establish new benchmarks, and enable precise, large-scale quantification for disease research and diagnosis.
Precision ICU Resource Planning: A Multimodal Model for Brain Surgery Outcomes
Dec 20, 2024

Abstract:Although advances in brain surgery techniques have led to fewer postoperative complications requiring Intensive Care Unit (ICU) monitoring, the routine transfer of patients to the ICU remains the clinical standard, despite its high cost. Predictive Gradient Boosted Trees based on clinical data have attempted to optimize ICU admission by identifying key risk factors pre-operatively; however, these approaches overlook valuable imaging data that could enhance prediction accuracy. In this work, we show that multimodal approaches that combine clinical data with imaging data outperform the current clinical data only baseline from 0.29 [F1] to 0.30 [F1], when only pre-operative clinical data is used and from 0.37 [F1] to 0.41 [F1], for pre- and post-operative data. This study demonstrates that effective ICU admission prediction benefits from multimodal data fusion, especially in contexts of severe class imbalance.
Unlocking the Potential of Digital Pathology: Novel Baselines for Compression
Dec 17, 2024



Abstract:Digital pathology offers a groundbreaking opportunity to transform clinical practice in histopathological image analysis, yet faces a significant hurdle: the substantial file sizes of pathological Whole Slide Images (WSI). While current digital pathology solutions rely on lossy JPEG compression to address this issue, lossy compression can introduce color and texture disparities, potentially impacting clinical decision-making. While prior research addresses perceptual image quality and downstream performance independently of each other, we jointly evaluate compression schemes for perceptual and downstream task quality on four different datasets. In addition, we collect an initially uncompressed dataset for an unbiased perceptual evaluation of compression schemes. Our results show that deep learning models fine-tuned for perceptual quality outperform conventional compression schemes like JPEG-XL or WebP for further compression of WSI. However, they exhibit a significant bias towards the compression artifacts present in the training data and struggle to generalize across various compression schemes. We introduce a novel evaluation metric based on feature similarity between original files and compressed files that aligns very well with the actual downstream performance on the compressed WSI. Our metric allows for a general and standardized evaluation of lossy compression schemes and mitigates the requirement to independently assess different downstream tasks. Our study provides novel insights for the assessment of lossy compression schemes for WSI and encourages a unified evaluation of lossy compression schemes to accelerate the clinical uptake of digital pathology.
Learned Image Compression for HE-stained Histopathological Images via Stain Deconvolution
Jun 18, 2024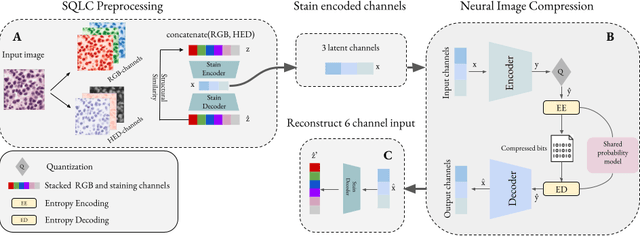
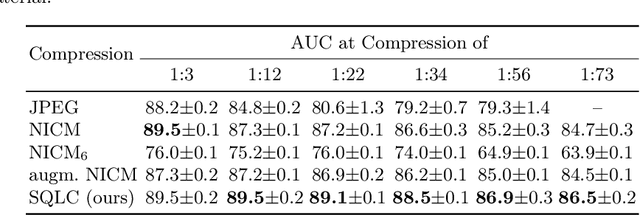
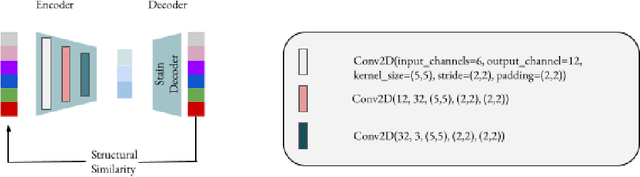
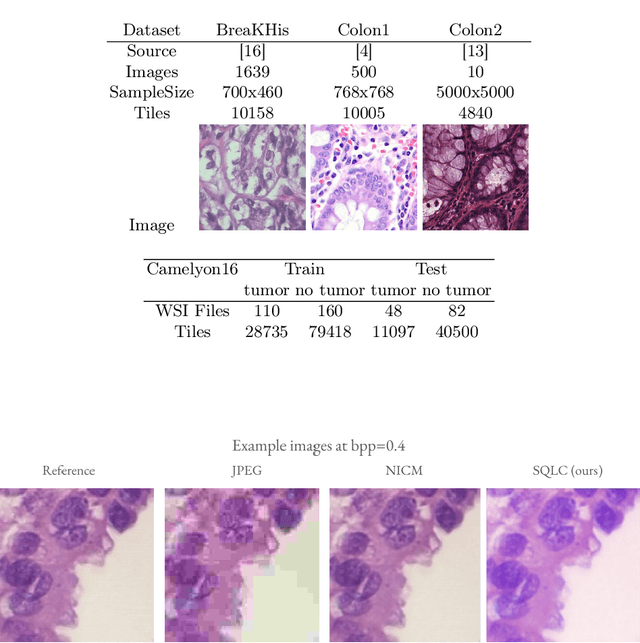
Abstract:Processing histopathological Whole Slide Images (WSI) leads to massive storage requirements for clinics worldwide. Even after lossy image compression during image acquisition, additional lossy compression is frequently possible without substantially affecting the performance of deep learning-based (DL) downstream tasks. In this paper, we show that the commonly used JPEG algorithm is not best suited for further compression and we propose Stain Quantized Latent Compression (SQLC ), a novel DL based histopathology data compression approach. SQLC compresses staining and RGB channels before passing it through a compression autoencoder (CAE ) in order to obtain quantized latent representations for maximizing the compression. We show that our approach yields superior performance in a classification downstream task, compared to traditional approaches like JPEG, while image quality metrics like the Multi-Scale Structural Similarity Index (MS-SSIM) is largely preserved. Our method is online available.
Mitigating False Predictions In Unreasonable Body Regions
Apr 24, 2024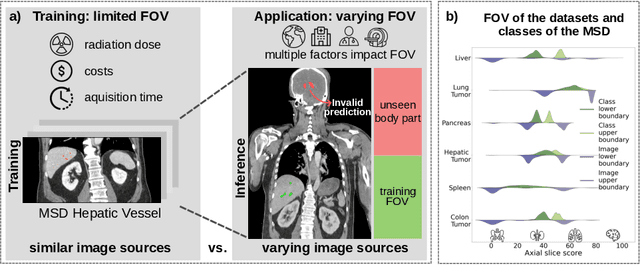
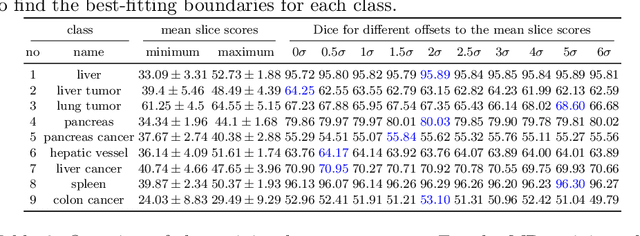
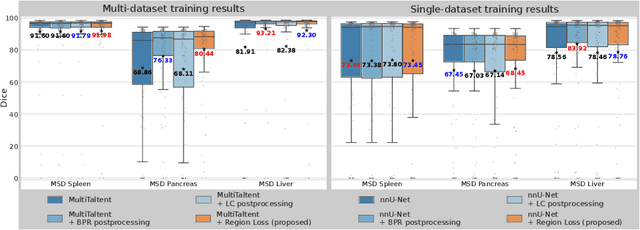

Abstract:Despite considerable strides in developing deep learning models for 3D medical image segmentation, the challenge of effectively generalizing across diverse image distributions persists. While domain generalization is acknowledged as vital for robust application in clinical settings, the challenges stemming from training with a limited Field of View (FOV) remain unaddressed. This limitation leads to false predictions when applied to body regions beyond the FOV of the training data. In response to this problem, we propose a novel loss function that penalizes predictions in implausible body regions, applicable in both single-dataset and multi-dataset training schemes. It is realized with a Body Part Regression model that generates axial slice positional scores. Through comprehensive evaluation using a test set featuring varying FOVs, our approach demonstrates remarkable improvements in generalization capabilities. It effectively mitigates false positive tumor predictions up to 85% and significantly enhances overall segmentation performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge