Robin Peretzke
CRONOS: Continuous Time Reconstruction for 4D Medical Longitudinal Series
Dec 18, 2025Abstract:Forecasting how 3D medical scans evolve over time is important for disease progression, treatment planning, and developmental assessment. Yet existing models either rely on a single prior scan, fixed grid times, or target global labels, which limits voxel-level forecasting under irregular sampling. We present CRONOS, a unified framework for many-to-one prediction from multiple past scans that supports both discrete (grid-based) and continuous (real-valued) timestamps in one model, to the best of our knowledge the first to achieve continuous sequence-to-image forecasting for 3D medical data. CRONOS learns a spatio-temporal velocity field that transports context volumes toward a target volume at an arbitrary time, while operating directly in 3D voxel space. Across three public datasets spanning Cine-MRI, perfusion CT, and longitudinal MRI, CRONOS outperforms other baselines, while remaining computationally competitive. We will release code and evaluation protocols to enable reproducible, multi-dataset benchmarking of multi-context, continuous-time forecasting.
Temporal Flow Matching for Learning Spatio-Temporal Trajectories in 4D Longitudinal Medical Imaging
Aug 29, 2025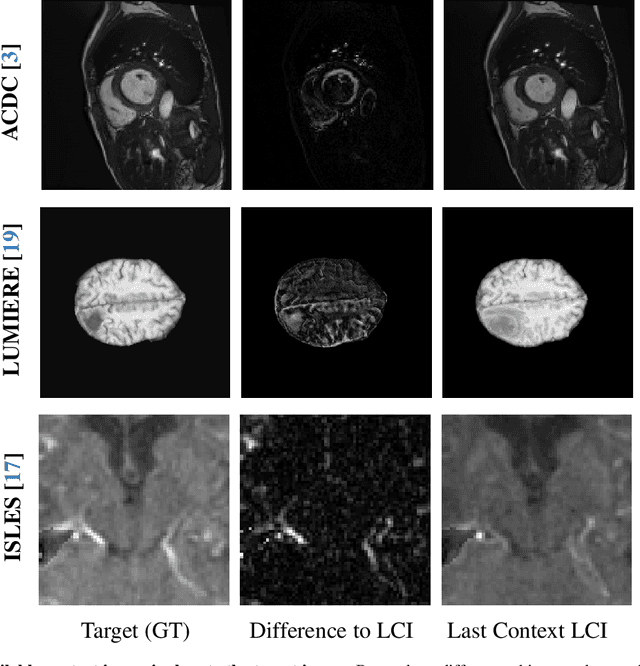
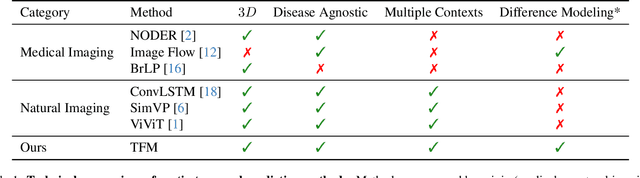

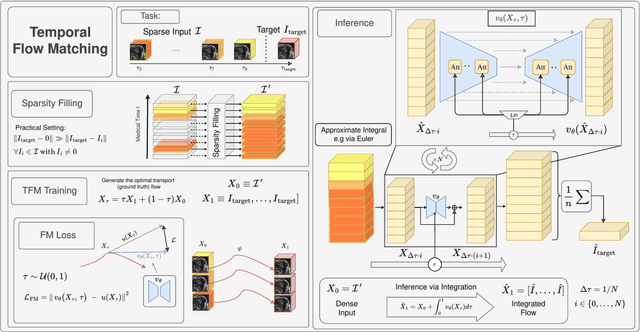
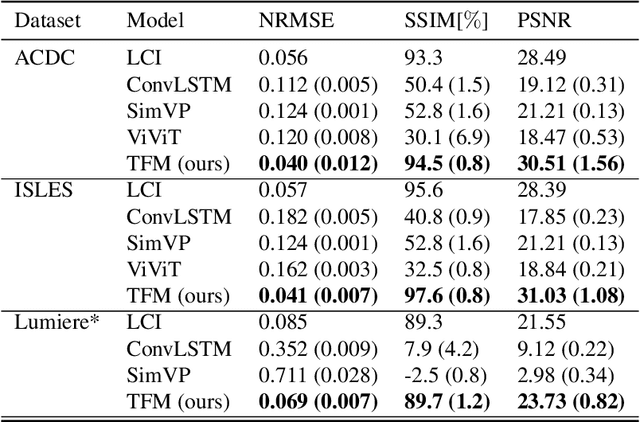
Abstract:Understanding temporal dynamics in medical imaging is crucial for applications such as disease progression modeling, treatment planning and anatomical development tracking. However, most deep learning methods either consider only single temporal contexts, or focus on tasks like classification or regression, limiting their ability for fine-grained spatial predictions. While some approaches have been explored, they are often limited to single timepoints, specific diseases or have other technical restrictions. To address this fundamental gap, we introduce Temporal Flow Matching (TFM), a unified generative trajectory method that (i) aims to learn the underlying temporal distribution, (ii) by design can fall back to a nearest image predictor, i.e. predicting the last context image (LCI), as a special case, and (iii) supports $3D$ volumes, multiple prior scans, and irregular sampling. Extensive benchmarks on three public longitudinal datasets show that TFM consistently surpasses spatio-temporal methods from natural imaging, establishing a new state-of-the-art and robust baseline for $4D$ medical image prediction.
nnLandmark: A Self-Configuring Method for 3D Medical Landmark Detection
Apr 10, 2025Abstract:Landmark detection plays a crucial role in medical imaging tasks that rely on precise spatial localization, including specific applications in diagnosis, treatment planning, image registration, and surgical navigation. However, manual annotation is labor-intensive and requires expert knowledge. While deep learning shows promise in automating this task, progress is hindered by limited public datasets, inconsistent benchmarks, and non-standardized baselines, restricting reproducibility, fair comparisons, and model generalizability. This work introduces nnLandmark, a self-configuring deep learning framework for 3D medical landmark detection, adapting nnU-Net to perform heatmap-based regression. By leveraging nnU-Net's automated configuration, nnLandmark eliminates the need for manual parameter tuning, offering out-of-the-box usability. It achieves state-of-the-art accuracy across two public datasets, with a mean radial error (MRE) of 1.5 mm on the Mandibular Molar Landmark (MML) dental CT dataset and 1.2 mm for anatomical fiducials on a brain MRI dataset (AFIDs), where nnLandmark aligns with the inter-rater variability of 1.5 mm. With its strong generalization, reproducibility, and ease of deployment, nnLandmark establishes a reliable baseline for 3D landmark detection, supporting research in anatomical localization and clinical workflows that depend on precise landmark identification. The code will be available soon.
An OpenMind for 3D medical vision self-supervised learning
Dec 22, 2024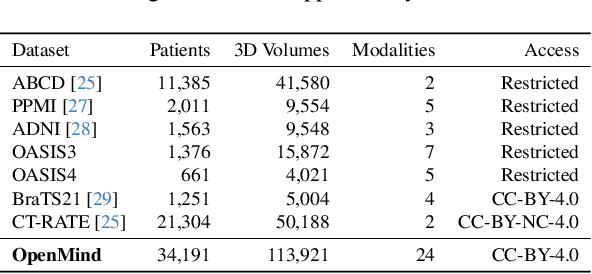
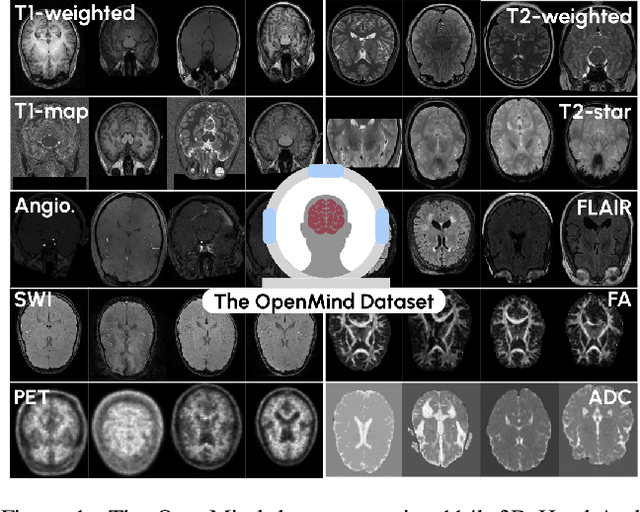
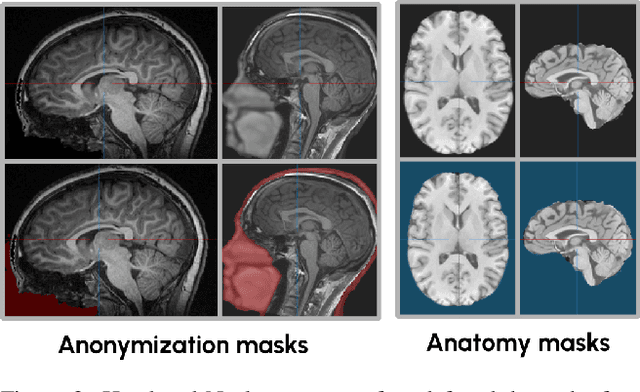
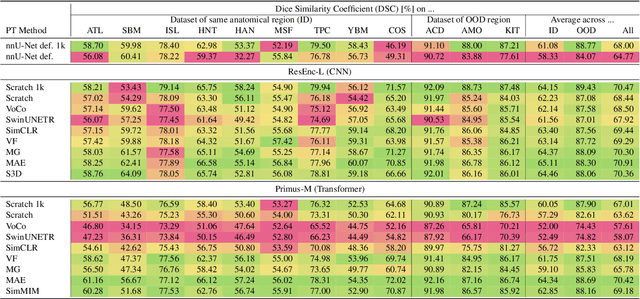
Abstract:The field of 3D medical vision self-supervised learning lacks consistency and standardization. While many methods have been developed it is impossible to identify the current state-of-the-art, due to i) varying and small pre-training datasets, ii) varying architectures, and iii) being evaluated on differing downstream datasets. In this paper we bring clarity to this field and lay the foundation for further method advancements: We a) publish the largest publicly available pre-training dataset comprising 114k 3D brain MRI volumes and b) benchmark existing SSL methods under common architectures and c) provide the code of our framework publicly to facilitate rapid adoption and reproduction. This pre-print \textit{only describes} the dataset contribution (a); Data, benchmark, and codebase will be made available shortly.
Precision ICU Resource Planning: A Multimodal Model for Brain Surgery Outcomes
Dec 20, 2024

Abstract:Although advances in brain surgery techniques have led to fewer postoperative complications requiring Intensive Care Unit (ICU) monitoring, the routine transfer of patients to the ICU remains the clinical standard, despite its high cost. Predictive Gradient Boosted Trees based on clinical data have attempted to optimize ICU admission by identifying key risk factors pre-operatively; however, these approaches overlook valuable imaging data that could enhance prediction accuracy. In this work, we show that multimodal approaches that combine clinical data with imaging data outperform the current clinical data only baseline from 0.29 [F1] to 0.30 [F1], when only pre-operative clinical data is used and from 0.37 [F1] to 0.41 [F1], for pre- and post-operative data. This study demonstrates that effective ICU admission prediction benefits from multimodal data fusion, especially in contexts of severe class imbalance.
Unlocking the Potential of Digital Pathology: Novel Baselines for Compression
Dec 17, 2024



Abstract:Digital pathology offers a groundbreaking opportunity to transform clinical practice in histopathological image analysis, yet faces a significant hurdle: the substantial file sizes of pathological Whole Slide Images (WSI). While current digital pathology solutions rely on lossy JPEG compression to address this issue, lossy compression can introduce color and texture disparities, potentially impacting clinical decision-making. While prior research addresses perceptual image quality and downstream performance independently of each other, we jointly evaluate compression schemes for perceptual and downstream task quality on four different datasets. In addition, we collect an initially uncompressed dataset for an unbiased perceptual evaluation of compression schemes. Our results show that deep learning models fine-tuned for perceptual quality outperform conventional compression schemes like JPEG-XL or WebP for further compression of WSI. However, they exhibit a significant bias towards the compression artifacts present in the training data and struggle to generalize across various compression schemes. We introduce a novel evaluation metric based on feature similarity between original files and compressed files that aligns very well with the actual downstream performance on the compressed WSI. Our metric allows for a general and standardized evaluation of lossy compression schemes and mitigates the requirement to independently assess different downstream tasks. Our study provides novel insights for the assessment of lossy compression schemes for WSI and encourages a unified evaluation of lossy compression schemes to accelerate the clinical uptake of digital pathology.
atTRACTive: Semi-automatic white matter tract segmentation using active learning
May 31, 2023



Abstract:Accurately identifying white matter tracts in medical images is essential for various applications, including surgery planning and tract-specific analysis. Supervised machine learning models have reached state-of-the-art solving this task automatically. However, these models are primarily trained on healthy subjects and struggle with strong anatomical aberrations, e.g. caused by brain tumors. This limitation makes them unsuitable for tasks such as preoperative planning, wherefore time-consuming and challenging manual delineation of the target tract is typically employed. We propose semi-automatic entropy-based active learning for quick and intuitive segmentation of white matter tracts from whole-brain tractography consisting of millions of streamlines. The method is evaluated on 21 openly available healthy subjects from the Human Connectome Project and an internal dataset of ten neurosurgical cases. With only a few annotations, the proposed approach enables segmenting tracts on tumor cases comparable to healthy subjects (dice=0.71), while the performance of automatic methods, like TractSeg dropped substantially (dice=0.34) in comparison to healthy subjects. The method is implemented as a prototype named atTRACTive in the freely available software MITK Diffusion. Manual experiments on tumor data showed higher efficiency due to lower segmentation times compared to traditional ROI-based segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge