Alejandro Frangi
From Nano to Macro: Overview of the IEEE Bio Image and Signal Processing Technical Committee
Oct 31, 2022


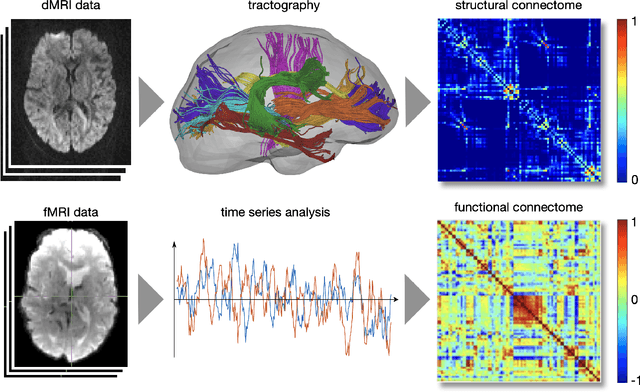
Abstract:The Bio Image and Signal Processing (BISP) Technical Committee (TC) of the IEEE Signal Processing Society (SPS) promotes activities within the broad technical field of biomedical image and signal processing. Areas of interest include medical and biological imaging, digital pathology, molecular imaging, microscopy, and associated computational imaging, image analysis, and image-guided treatment, alongside physiological signal processing, computational biology, and bioinformatics. BISP has 40 members and covers a wide range of EDICS, including CIS-MI: Medical Imaging, BIO-MIA: Medical Image Analysis, BIO-BI: Biological Imaging, BIO: Biomedical Signal Processing, BIO-BCI: Brain/Human-Computer Interfaces, and BIO-INFR: Bioinformatics. BISP plays a central role in the organization of the IEEE International Symposium on Biomedical Imaging (ISBI) and contributes to the technical sessions at the IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), and the IEEE International Conference on Image Processing (ICIP). In this paper, we provide a brief history of the TC, review the technological and methodological contributions its community delivered, and highlight promising new directions we anticipate.
The Multiscenario Multienvironment BioSecure Multimodal Database (BMDB)
Nov 17, 2021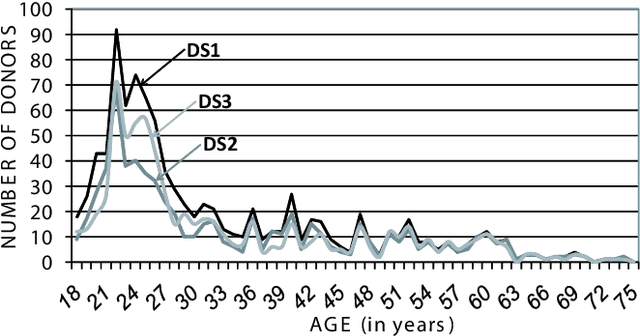
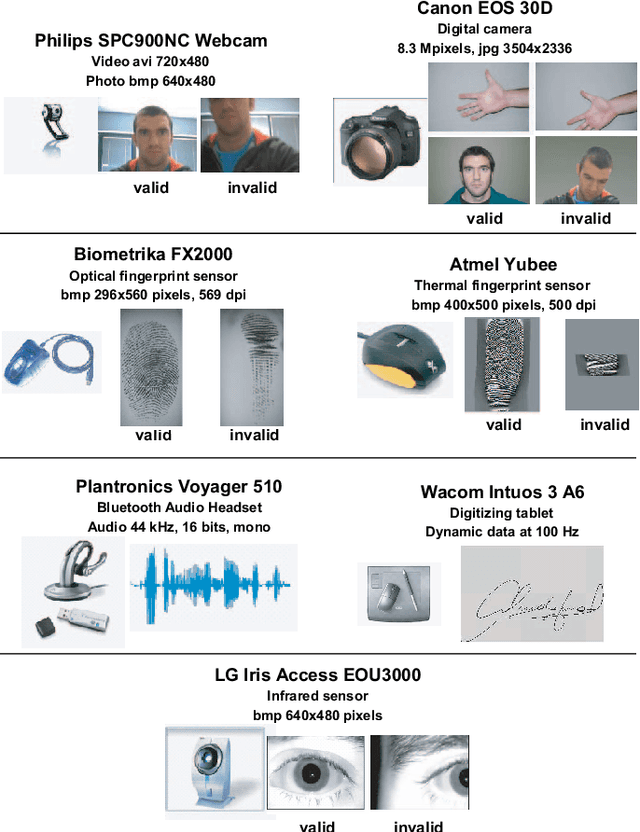

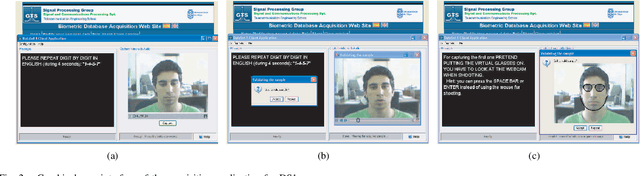

Abstract:A new multimodal biometric database designed and acquired within the framework of the European BioSecure Network of Excellence is presented. It is comprised of more than 600 individuals acquired simultaneously in three scenarios: 1) over the Internet, 2) in an office environment with desktop PC, and 3) in indoor/outdoor environments with mobile portable hardware. The three scenarios include a common part of audio/video data. Also, signature and fingerprint data have been acquired both with desktop PC and mobile portable hardware. Additionally, hand and iris data were acquired in the second scenario using desktop PC. Acquisition has been conducted by 11 European institutions. Additional features of the BioSecure Multimodal Database (BMDB) are: two acquisition sessions, several sensors in certain modalities, balanced gender and age distributions, multimodal realistic scenarios with simple and quick tasks per modality, cross-European diversity, availability of demographic data, and compatibility with other multimodal databases. The novel acquisition conditions of the BMDB allow us to perform new challenging research and evaluation of either monomodal or multimodal biometric systems, as in the recent BioSecure Multimodal Evaluation campaign. A description of this campaign including baseline results of individual modalities from the new database is also given. The database is expected to be available for research purposes through the BioSecure Association during 2008
Statistical Dependency Guided Contrastive Learning for Multiple Labeling in Prenatal Ultrasound
Aug 11, 2021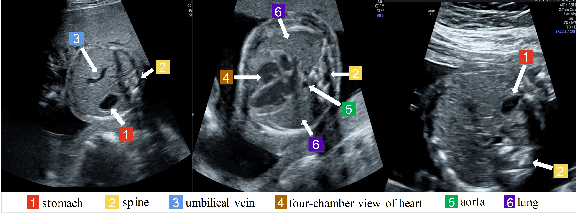
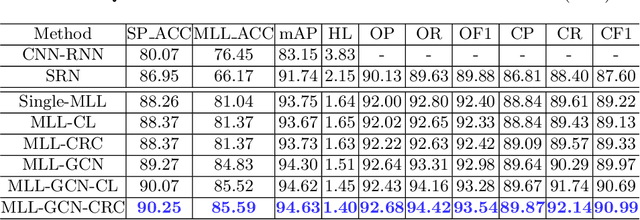
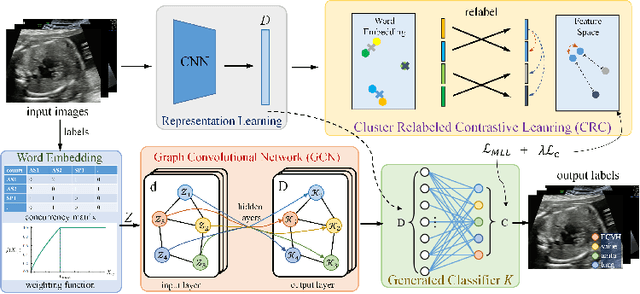
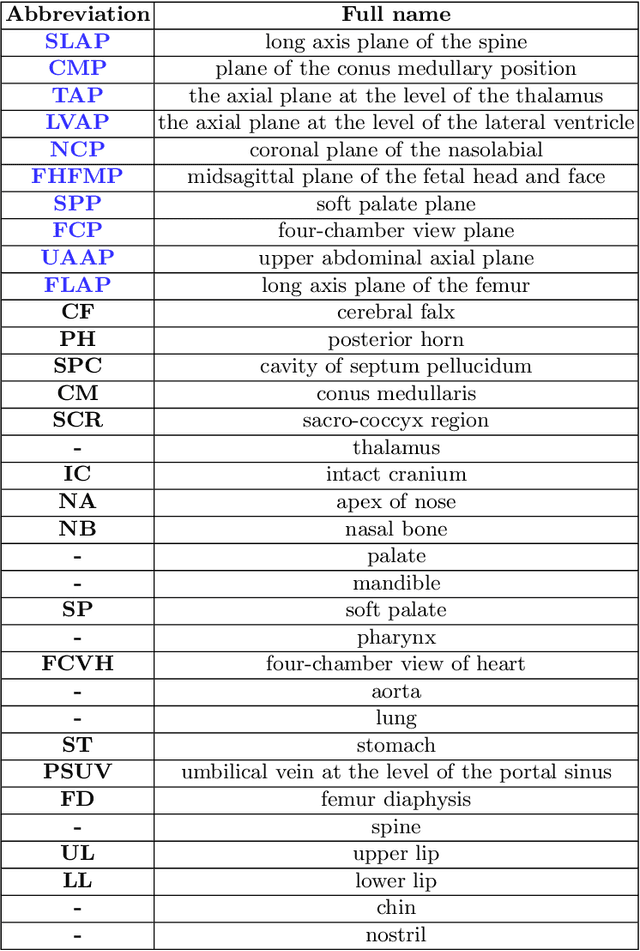
Abstract:Standard plane recognition plays an important role in prenatal ultrasound (US) screening. Automatically recognizing the standard plane along with the corresponding anatomical structures in US image can not only facilitate US image interpretation but also improve diagnostic efficiency. In this study, we build a novel multi-label learning (MLL) scheme to identify multiple standard planes and corresponding anatomical structures of fetus simultaneously. Our contribution is three-fold. First, we represent the class correlation by word embeddings to capture the fine-grained semantic and latent statistical concurrency. Second, we equip the MLL with a graph convolutional network to explore the inner and outer relationship among categories. Third, we propose a novel cluster relabel-based contrastive learning algorithm to encourage the divergence among ambiguous classes. Extensive validation was performed on our large in-house dataset. Our approach reports the highest accuracy as 90.25% for standard planes labeling, 85.59% for planes and structures labeling and mAP as 94.63%. The proposed MLL scheme provides a novel perspective for standard plane recognition and can be easily extended to other medical image classification tasks.
Contrastive Rendering for Ultrasound Image Segmentation
Oct 10, 2020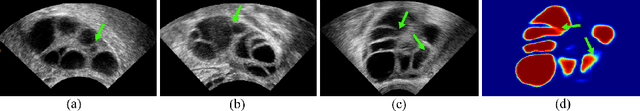
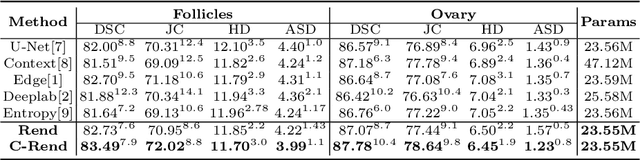
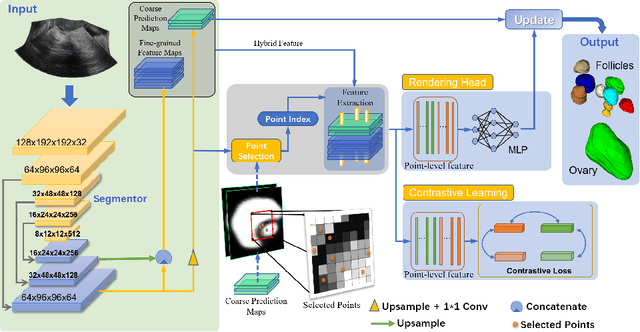
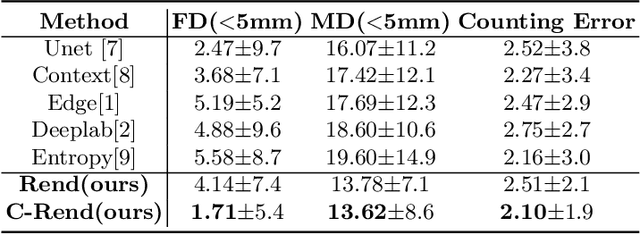
Abstract:Ultrasound (US) image segmentation embraced its significant improvement in deep learning era. However, the lack of sharp boundaries in US images still remains an inherent challenge for segmentation. Previous methods often resort to global context, multi-scale cues or auxiliary guidance to estimate the boundaries. It is hard for these methods to approach pixel-level learning for fine-grained boundary generating. In this paper, we propose a novel and effective framework to improve boundary estimation in US images. Our work has three highlights. First, we propose to formulate the boundary estimation as a rendering task, which can recognize ambiguous points (pixels/voxels) and calibrate the boundary prediction via enriched feature representation learning. Second, we introduce point-wise contrastive learning to enhance the similarity of points from the same class and contrastively decrease the similarity of points from different classes. Boundary ambiguities are therefore further addressed. Third, both rendering and contrastive learning tasks contribute to consistent improvement while reducing network parameters. As a proof-of-concept, we performed validation experiments on a challenging dataset of 86 ovarian US volumes. Results show that our proposed method outperforms state-of-the-art methods and has the potential to be used in clinical practice.
Searching Collaborative Agents for Multi-plane Localization in 3D Ultrasound
Jul 30, 2020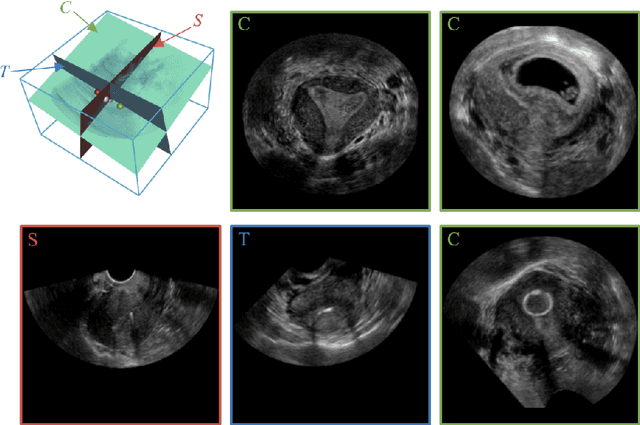
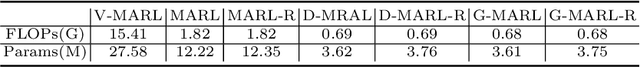
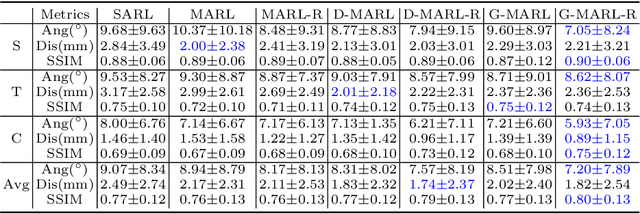
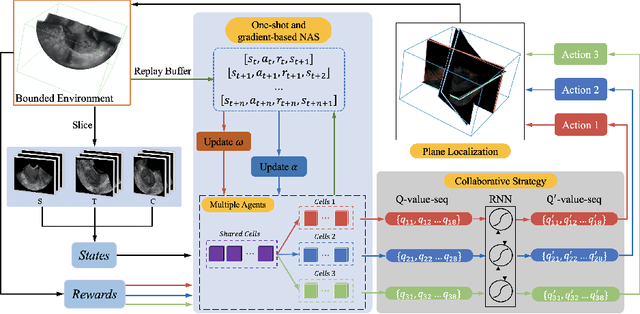

Abstract:3D ultrasound (US) is widely used due to its rich diagnostic information, portability and low cost. Automated standard plane (SP) localization in US volume not only improves efficiency and reduces user-dependence, but also boosts 3D US interpretation. In this study, we propose a novel Multi-Agent Reinforcement Learning (MARL) framework to localize multiple uterine SPs in 3D US simultaneously. Our contribution is two-fold. First, we equip the MARL with a one-shot neural architecture search (NAS) module to obtain the optimal agent for each plane. Specifically, Gradient-based search using Differentiable Architecture Sampler (GDAS) is employed to accelerate and stabilize the training process. Second, we propose a novel collaborative strategy to strengthen agents' communication. Our strategy uses recurrent neural network (RNN) to learn the spatial relationship among SPs effectively. Extensively validated on a large dataset, our approach achieves the accuracy of 7.05 degree/2.21mm, 8.62 degree/2.36mm and 5.93 degree/0.89mm for the mid-sagittal, transverse and coronal plane localization, respectively. The proposed MARL framework can significantly increase the plane localization accuracy and reduce the computational cost and model size.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge